

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135463.3 + phase: 0 /pseudo
(924 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
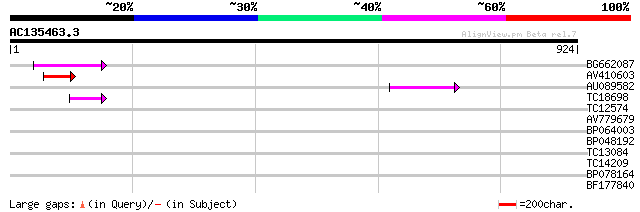
Score E
Sequences producing significant alignments: (bits) Value
BG662087 89 3e-18
AV410603 70 1e-12
AU089582 62 5e-10
TC18698 42 3e-04
TC12574 35 0.041
AV779679 35 0.041
BP064003 33 0.21
BP048192 32 0.60
TC13084 30 2.3
TC14209 similar to UP|AAQ72788 (AAQ72788) Hypersensitive-induced... 29 3.0
BP078164 29 3.9
BF177840 28 6.6
>BG662087
Length = 373
Score = 89.0 bits (219), Expect = 3e-18
Identities = 44/119 (36%), Positives = 67/119 (55%)
Frame = +1
Query: 40 GTMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQK 99
G R+ +DY LNK K+ YPLP ID L+D + S +D SGYHQIK+ D K
Sbjct: 16 GKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDEDK 195
Query: 100 TASRTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSRTEEEH 158
TA T +Y Y+ +PFG+ NA + M+R+F + + + V++D++++ S H
Sbjct: 196 TAFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRANH 372
>AV410603
Length = 162
Score = 70.5 bits (171), Expect = 1e-12
Identities = 30/53 (56%), Positives = 41/53 (76%)
Frame = +1
Query: 55 TIKNRYPLPRIDDLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYG 107
T+K+ +P+P +D+L+D+L G+ FSK+DLRSGYHQI VK ED KT RT +G
Sbjct: 4 TVKDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTHHG 162
>AU089582
Length = 383
Score = 61.6 bits (148), Expect = 5e-10
Identities = 42/115 (36%), Positives = 55/115 (47%), Gaps = 2/115 (1%)
Frame = +1
Query: 620 DLYS*HCEATWCTVEYCVG*GSEVHL*ILEEFTRCFGFEVEVEFGLSSTDRWSVGED-DP 678
DL * C WCT Y S +H+ LE F+ CFG ++ E+ SS++ WSV ED
Sbjct: 28 DLLG*DCFLAWCTCVYNFRSRSSIHITFLEVFSNCFGNSIKNEYRFSSSN*WSVREDYSD 207
Query: 679 IIGN*ECVCLN-KVELGIVTYR**SSHTTTVTIRVSVWHRSKLCMVGGP*LLYVG 732
+ G C+C+ + GI * +SH WH K CMVG L VG
Sbjct: 208 LRGYASCLCVXPERXAGISICL*WNSHIIIAIGLAFRWHHLKPCMVGDAGHL*VG 372
>TC18698
Length = 808
Score = 42.4 bits (98), Expect = 3e-04
Identities = 20/61 (32%), Positives = 35/61 (56%)
Frame = -2
Query: 98 QKTASRTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSRTEEE 157
+KT + +Y Y+VMP G+ N + M++IFH + K V V+++D+++ S E
Sbjct: 804 KKTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSSQE*F 625
Query: 158 H 158
H
Sbjct: 624 H 622
>TC12574
Length = 325
Score = 35.4 bits (80), Expect = 0.041
Identities = 14/33 (42%), Positives = 23/33 (69%)
Frame = +2
Query: 125 FMEYMNRIFHAYLDKFVVVFIDDILIYSRTEEE 157
F +N IF ++ + F++VFI+DIL Y+ +EE
Sbjct: 2 FKNSVNHIFESFFEHFMIVFINDILSYTEDKEE 100
>AV779679
Length = 440
Score = 35.4 bits (80), Expect = 0.041
Identities = 16/30 (53%), Positives = 21/30 (69%)
Frame = +3
Query: 41 TMRLCIDYRQLNKVTIKNRYPLPRIDDLMD 70
TM+LC DY QL+ VTI N+ LP +D+ D
Sbjct: 345 TMQLCDDYMQLDYVTIPNKSLLPHLDEWSD 434
>BP064003
Length = 509
Score = 33.1 bits (74), Expect = 0.21
Identities = 30/101 (29%), Positives = 47/101 (45%), Gaps = 6/101 (5%)
Frame = +1
Query: 347 GSRFEVFSDHKSLKYLFDQKELNMRQRRWLELLKDYDFGLNYHP------GKANVVADAL 400
GSRF + +D+ + QK R R W + L G+ + P ++N VADAL
Sbjct: 115 GSRFVIKTDNVATSSFLTQKRAPTRAR-WQDFLSG---GVRHGPRVQAREDQSNKVADAL 282
Query: 401 SRKTLHMSALMVKEFDLLEQFRDLSLVCELTPQSVQLGMLK 441
SRK S + LE+ D+S + P+ ++ G+ K
Sbjct: 283 SRKAELAS-------NKLEEIADMSQLKGAIPERIREGLEK 384
>BP048192
Length = 551
Score = 31.6 bits (70), Expect = 0.60
Identities = 13/42 (30%), Positives = 25/42 (58%), Gaps = 3/42 (7%)
Frame = +3
Query: 348 SRFEVFSDHKSLKYLFDQ---KELNMRQRRWLELLKDYDFGL 386
+ F ++D+ ++ Y Q KE+++ + RW +KDYD G+
Sbjct: 192 THFLTYADNNAVGYFIKQGFTKEIHLEKERWQGYIKDYDGGI 317
>TC13084
Length = 508
Score = 29.6 bits (65), Expect = 2.3
Identities = 12/24 (50%), Positives = 17/24 (70%)
Frame = +1
Query: 468 NGTKDSDFKVDDQGVLRFRGRICI 491
N T+DSD+ +++ RFRGR CI
Sbjct: 160 NMTEDSDYDEEEEDYPRFRGRYCI 231
>TC14209 similar to UP|AAQ72788 (AAQ72788) Hypersensitive-induced response
protein, complete
Length = 1298
Score = 29.3 bits (64), Expect = 3.0
Identities = 14/54 (25%), Positives = 24/54 (43%)
Frame = -1
Query: 319 IHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQKELNMRQ 372
+ NY THD+EL + H + G++ + F + YL N+R+
Sbjct: 1124 VQNSNYSTHDIELG-------VQLHSIIGAKMQSFKQQRYFCYLLSTIHQNLRK 984
>BP078164
Length = 349
Score = 28.9 bits (63), Expect = 3.9
Identities = 9/19 (47%), Positives = 13/19 (68%)
Frame = -1
Query: 823 HIELVCHHIFRICMMYFMC 841
H EL+ HH +RIC++ C
Sbjct: 109 HKELIIHHFYRICILQLTC 53
>BF177840
Length = 410
Score = 28.1 bits (61), Expect = 6.6
Identities = 15/39 (38%), Positives = 22/39 (55%)
Frame = +1
Query: 634 EYCVG*GSEVHL*ILEEFTRCFGFEVEVEFGLSSTDRWS 672
E+ VG G ++H LE G +V V + LSST+ W+
Sbjct: 13 EHSVGSGHQIHKPFLENIMG*GGNKVVVLYNLSSTNGWA 129
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.341 0.151 0.492
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,989,605
Number of Sequences: 28460
Number of extensions: 222187
Number of successful extensions: 1567
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 1554
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1565
length of query: 924
length of database: 4,897,600
effective HSP length: 99
effective length of query: 825
effective length of database: 2,080,060
effective search space: 1716049500
effective search space used: 1716049500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC135463.3


