

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135463.1 - phase: 0 /pseudo
(488 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
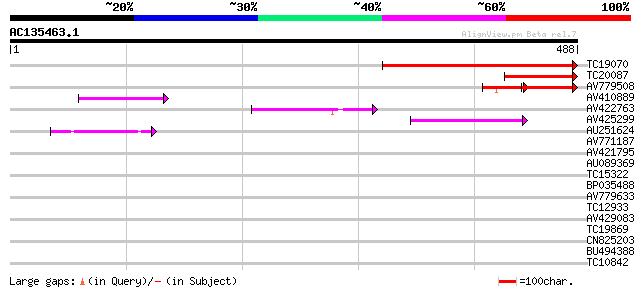
TC19070 weakly similar to UP|Q9LRP6 (Q9LRP6) DNA-binding protein... 262 1e-70
TC20087 weakly similar to UP|Q9MQV2 (Q9MQV2) DNA-binding protein... 91 6e-19
AV779508 65 3e-15
AV410889 54 7e-08
AV422763 47 7e-06
AV425299 44 8e-05
AU251624 42 2e-04
AV771187 35 0.021
AV421795 35 0.021
AU089369 31 0.40
TC15322 weakly similar to UP|Q9LG23 (Q9LG23) F14J16.14 (At1g5589... 30 0.68
BP035488 30 1.2
AV779633 29 2.0
TC12933 homologue to UP|Q39821 (Q39821) SDL12A, partial (33%) 28 3.4
AV429083 28 4.4
TC19869 27 7.5
CN825203 27 9.8
BU494388 27 9.8
TC10842 27 9.8
>TC19070 weakly similar to UP|Q9LRP6 (Q9LRP6) DNA-binding protein
(AT3g15590/MQD17_5), partial (20%)
Length = 744
Score = 262 bits (669), Expect = 1e-70
Identities = 120/167 (71%), Positives = 145/167 (85%)
Frame = +3
Query: 322 LAAIEAWGRLKKIEEAEAVFEMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDS 381
LAA+EAWGRL+KIEEAEA+FEMM+NKWKL+++NY LLK+Y HKML KGKDLI+ MG++
Sbjct: 15 LAAVEAWGRLQKIEEAEAIFEMMANKWKLSSKNYSVLLKVYANHKMLKKGKDLIQRMGEN 194
Query: 382 GCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHN 441
GC IGP TWD+LV LY QAGEVEKAD VLQK ++Q++M PMF T+M +MEQYAKRGDVHN
Sbjct: 195 GCKIGPLTWDSLVKLYTQAGEVEKADAVLQKVMEQSRMTPMFNTYMAVMEQYAKRGDVHN 374
Query: 442 AEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADNLF 488
+EKIF+R+RQA Y SRI+PF L QAY AK+PAYGIR+RMKADN+F
Sbjct: 375 SEKIFHRMRQAGYTSRITPFQVLVQAYIKAKVPAYGIRDRMKADNVF 515
>TC20087 weakly similar to UP|Q9MQV2 (Q9MQV2) DNA-binding protein, partial
(13%)
Length = 492
Score = 90.5 bits (223), Expect = 6e-19
Identities = 39/62 (62%), Positives = 51/62 (81%)
Frame = +1
Query: 427 MTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADN 486
+ I+EQY+KRGD+HN+EKIFYR++QA Y SRI + L QAY AKLPAYGIR+R+K DN
Sbjct: 1 IAILEQYSKRGDIHNSEKIFYRMKQAGYTSRIRQYQVLLQAYIKAKLPAYGIRDRLKGDN 180
Query: 487 LF 488
++
Sbjct: 181 IY 186
>AV779508
Length = 431
Score = 64.7 bits (156), Expect(2) = 3e-15
Identities = 29/48 (60%), Positives = 36/48 (74%)
Frame = -3
Query: 441 NAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADNLF 488
N EKIFYR++ A Y SRI + L QAY AKLPAYGIR+R+K DN++
Sbjct: 321 NPEKIFYRMKPAGYTSRIRQYPVLVQAYIKAKLPAYGIRDRLKGDNIY 178
Score = 33.5 bits (75), Expect(2) = 3e-15
Identities = 14/42 (33%), Positives = 28/42 (66%), Gaps = 3/42 (7%)
Frame = -2
Query: 408 TVLQKALQQN---KMKPMFTTFMTIMEQYAKRGDVHNAEKIF 446
++ QKA Q + ++ P+F +++ I++ +KRGD+H + K F
Sbjct: 430 SIFQKATQPSHAHQVNPLFFSYIAILDPDSKRGDIHKSRKDF 305
>AV410889
Length = 436
Score = 53.5 bits (127), Expect = 7e-08
Identities = 24/77 (31%), Positives = 45/77 (58%)
Frame = +1
Query: 60 LSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYAS 119
+ V L++WV++G+ ++ ++ + LR + + ALQ +W+ + + A
Sbjct: 205 IPVTPLLNQWVQEGRPINHGDLQFFIKQLRSFRRFKHALQISEWMSDERNHYLHSGDIAI 384
Query: 120 KLDLIAKLRGLPKAEKY 136
+LDLI+K+RGL +AEKY
Sbjct: 385 RLDLISKVRGLEEAEKY 435
>AV422763
Length = 410
Score = 47.0 bits (110), Expect = 7e-06
Identities = 33/113 (29%), Positives = 58/113 (51%), Gaps = 5/113 (4%)
Frame = +2
Query: 209 KENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEG-CELDHLTRASLARHYAAAGLT 267
KEN + T+ I ++ + D++GM +++ M+A+ +D T ++ A Y AG
Sbjct: 14 KENDLCNGATFTIRLNAYVAAKDVEGMEKLLMQMEADPMATVDWYTYSTAANGYLKAGNV 193
Query: 268 EKTEAILKE----IEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCESKP 316
EK A LK+ ++G+ L+ +L YA +G DEV R+W ++ P
Sbjct: 194 EKALAALKKSEQSVKGKKLR---LAYESLQSTYAAIGNNDEVYRLWNRIKNLP 343
>AV425299
Length = 323
Score = 43.5 bits (101), Expect = 8e-05
Identities = 22/100 (22%), Positives = 52/100 (52%)
Frame = +3
Query: 346 NKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEK 405
N+ L A +L+ +Y + L + K+ + + G +++++ YV AG+ +
Sbjct: 9 NQIGLDASTASALVHLYCKVGNLERAKEAFEVLTRKGFQPDIKVYNSMIMGYVNAGDPKA 188
Query: 406 ADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKI 445
+++++K ++ +KP + ++ Y++RGDV A +I
Sbjct: 189 GESLMKKMEERGGIKPTKEIYTALLRAYSERGDVIGAGRI 308
>AU251624
Length = 350
Score = 42.0 bits (97), Expect = 2e-04
Identities = 29/91 (31%), Positives = 48/91 (51%)
Frame = +1
Query: 36 SSDKKSFSSRRRSELFKAIVSVSGLSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYG 95
S KK +LFK S L+V L+++++ GK + + E+G L LR+RK+Y
Sbjct: 82 SCSKKYLDETLYLKLFKD--GGSDLNVRQQLNQFIKSGKRVYKWEVGDTLKKLRQRKLYV 255
Query: 96 RALQALDWLESNKKLEFTEKEYASKLDLIAK 126
AL+ + + ++ T + A LDL+AK
Sbjct: 256 PALKLSETMAKRNMIK-TVSDQAIHLDLLAK 345
>AV771187
Length = 484
Score = 35.4 bits (80), Expect = 0.021
Identities = 19/79 (24%), Positives = 36/79 (45%)
Frame = -3
Query: 364 RHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMF 423
R + + +L + M GC T+D+L+ Y E+++A VL+ + N P
Sbjct: 248 RRYRIQRALELFEDMKKKGCAPNRVTYDSLIRYYSATNEIDRAVEVLRDMQRLNHGIPSS 69
Query: 424 TTFMTIMEQYAKRGDVHNA 442
+++ I+ + G V A
Sbjct: 68 SSYTPIIHALCEAGRVAEA 12
Score = 31.2 bits (69), Expect = 0.40
Identities = 13/45 (28%), Positives = 27/45 (59%)
Frame = -3
Query: 237 QIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGEN 281
++ E MK +GC + +T SL R+Y+A ++ +L++++ N
Sbjct: 221 ELFEDMKKKGCAPNRVTYDSLIRYYSATNEIDRAVEVLRDMQRLN 87
>AV421795
Length = 267
Score = 35.4 bits (80), Expect = 0.021
Identities = 21/72 (29%), Positives = 38/72 (52%), Gaps = 3/72 (4%)
Frame = +2
Query: 41 SFSSRRRSELFKAIVSVSG---LSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRA 97
S SS LF I S SG + + L++W+++GK ++ E+ + LR + + A
Sbjct: 41 SSSSSPTDSLFLRI-SRSGDRAIPMTPLLNQWLQEGKLINYSELQFFIKQLRASRRFNHA 217
Query: 98 LQALDWLESNKK 109
LQ +W+ + +K
Sbjct: 218 LQISEWMSNERK 253
>AU089369
Length = 728
Score = 31.2 bits (69), Expect = 0.40
Identities = 35/138 (25%), Positives = 58/138 (41%), Gaps = 7/138 (5%)
Frame = +3
Query: 153 LLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDKKKIA----DVLLMME 208
L+ A L+ L K EE + R G A L + + K+A D + +
Sbjct: 45 LIGTYAKLDMLDKAEELKERARRRGAKPNAKTWEIFLDYHLRKGDFKLAVDSLDKAISIG 224
Query: 209 KENVK---PSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAG 265
+ N PSS + ++ D+DG + +E +K L L R YAAAG
Sbjct: 225 RGNGDKWVPSSESVGTIMKHFEQEKDVDGAXEFLEILKKSTDSLXADVFEPLIRTYAAAG 404
Query: 266 LTEKTEAILKEIEGENLK 283
T + A+ + ++ EN++
Sbjct: 405 RT--SSALQRRLKMENVE 452
>TC15322 weakly similar to UP|Q9LG23 (Q9LG23) F14J16.14
(At1g55890/F14J16_4), partial (33%)
Length = 588
Score = 30.4 bits (67), Expect = 0.68
Identities = 12/76 (15%), Positives = 39/76 (50%)
Frame = +2
Query: 358 LLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQN 417
L+ +Y + M + + + M + CT +++AL++ Y+ + + + + ++ +
Sbjct: 359 LITLYGKSGMPKRARQMFDEMPERNCTRTVLSFNALLAAYLHSKQYRVVERIFKEVPGEI 538
Query: 418 KMKPMFTTFMTIMEQY 433
+KP ++ T+++ +
Sbjct: 539 SVKPDLVSYNTVIKAF 586
>BP035488
Length = 534
Score = 29.6 bits (65), Expect = 1.2
Identities = 31/147 (21%), Positives = 52/147 (35%), Gaps = 43/147 (29%)
Frame = +3
Query: 292 LLRLYAILGRADEVERIWK----------------------------------------V 311
+L + +LG A E +R+WK
Sbjct: 87 ILNGWCVLGNAHEAKRVWKDIMASKCRPDLFTYATFIKALTKKGKLGTALKLFRGMWNEG 266
Query: 312 CESKPRVEDCLAAIEAWGRLKKIEEAEAVFEMMSNK-WKLTARNYESLLKIYIRHKMLNK 370
C KP V C I+A K++ EA VF+ M + + Y SL+K + + + K
Sbjct: 267 CNCKPDVVICNCIIDALCFKKRVPEALEVFQDMKERGCEPNVATYNSLIKHLCKIRRMEK 446
Query: 371 GKDLIKTM--GDSGCTIGPTTWDALVS 395
+L++ M C T+ L++
Sbjct: 447 VYELVEDMERKKGSCMPNAVTYSCLLN 527
Score = 29.3 bits (64), Expect = 1.5
Identities = 15/58 (25%), Positives = 30/58 (50%)
Frame = +3
Query: 389 TWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKIF 446
TW+ +++ + G +A V K + +K +P T+ T ++ K+G + A K+F
Sbjct: 75 TWNVILNGWCVLGNAHEAKRVW-KDIMASKCRPDLFTYATFIKALTKKGKLGTALKLF 245
>AV779633
Length = 440
Score = 28.9 bits (63), Expect = 2.0
Identities = 13/56 (23%), Positives = 33/56 (58%)
Frame = -2
Query: 391 DALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKIF 446
+ L+ Y +AG + +A +++ + + +KP ++ ++++ + K GD+ AE +F
Sbjct: 403 NTLIDGYCEAGLMSQALALMENSWKTG-VKPDIVSYNSLLKGFCKAGDLVRAESLF 239
>TC12933 homologue to UP|Q39821 (Q39821) SDL12A, partial (33%)
Length = 603
Score = 28.1 bits (61), Expect = 3.4
Identities = 16/47 (34%), Positives = 25/47 (53%), Gaps = 1/47 (2%)
Frame = +1
Query: 366 KMLNKGKDLIKTMGDSGCTIG-PTTWDALVSLYVQAGEVEKADTVLQ 411
+++NK + +GD G PT WDAL S+ V G+ +VL+
Sbjct: 4 QLVNKIQRACTALGDHGEESALPTLWDALPSIAVVGGQSSGKSSVLE 144
>AV429083
Length = 316
Score = 27.7 bits (60), Expect = 4.4
Identities = 14/53 (26%), Positives = 24/53 (44%)
Frame = +3
Query: 209 KENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHY 261
K N+KP T+ IL+D + ++ M +G D +T ++L Y
Sbjct: 24 KNNIKPDVSTFNILVDALCKKGKVKHAKNVLAVMIKQGVAPDLVTYSTLLDGY 182
>TC19869
Length = 560
Score = 26.9 bits (58), Expect = 7.5
Identities = 14/44 (31%), Positives = 24/44 (53%)
Frame = +2
Query: 186 NQLLLIYKKIDKKKIADVLLMMEKENVKPSSYTYKILIDVKGLS 229
++ L Y +KK A++ + E NVK + ++ L+DV LS
Sbjct: 194 SEALTFYGVPEKKPPANLAALCEHYNVKLNGAAHRALVDVDALS 325
>CN825203
Length = 644
Score = 26.6 bits (57), Expect = 9.8
Identities = 27/115 (23%), Positives = 52/115 (44%), Gaps = 11/115 (9%)
Frame = -2
Query: 261 YAAAGLTEKTEAILKEI--EGENLKENMWVCPTLLRLYAILGRADEVERIWK-----VCE 313
Y+ GL EK E +L+ + +G+ N W + + ++ + +K + E
Sbjct: 616 YSRKGLIEKAETMLRSMVDKGKTPTPNSW--SIIASGHVAKENMEKAFQCFKEALAVLAE 443
Query: 314 SK---PRVEDCLAAIEAW-GRLKKIEEAEAVFEMMSNKWKLTARNYESLLKIYIR 364
+K P+ D +++I +W + IEE E + + Y SL+K+Y+R
Sbjct: 442 NKGWRPK-SDVVSSILSWVSDNRDIEEVEDFVNSLKKVMSMNRDMYLSLIKLYVR 281
>BU494388
Length = 350
Score = 26.6 bits (57), Expect = 9.8
Identities = 19/60 (31%), Positives = 29/60 (47%), Gaps = 1/60 (1%)
Frame = +2
Query: 186 NQLLLIYKKIDKKKIA-DVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKA 244
++L+L+ K+ + V ME VKPSS + LI SNDI + E M++
Sbjct: 140 SELILLCGKLKNLPLGMRVFTSMETIGVKPSSLVFNSLISACLSSNDIVTAYSLFEIMES 319
>TC10842
Length = 597
Score = 26.6 bits (57), Expect = 9.8
Identities = 19/83 (22%), Positives = 42/83 (49%), Gaps = 4/83 (4%)
Frame = +2
Query: 280 ENLKENMWVC-PTLLRLYAILGRADEVERIW---KVCESKPRVEDCLAAIEAWGRLKKIE 335
+ + ++ W+ L+ LYA LG D++++IW ++ + K + I A+ L ++
Sbjct: 26 KRITQSQWITYDFLIILYAGLGSKDKLDQIWNSLRMTKQKMINRNYSCIISAYLMLGHVK 205
Query: 336 EAEAVFEMMSNKWKLTARNYESL 358
E V + +WK + +++ L
Sbjct: 206 EVGEVID----QWKNSTIDFDML 262
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.133 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,756,269
Number of Sequences: 28460
Number of extensions: 78275
Number of successful extensions: 371
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 367
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 370
length of query: 488
length of database: 4,897,600
effective HSP length: 94
effective length of query: 394
effective length of database: 2,222,360
effective search space: 875609840
effective search space used: 875609840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC135463.1


