

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
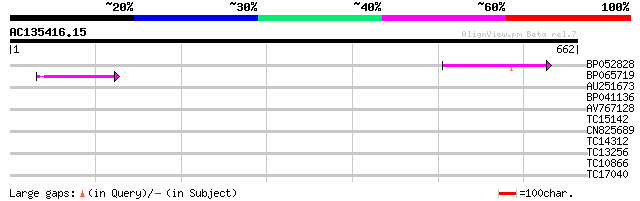
Query= AC135416.15 - phase: 0 /pseudo
(662 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP052828 93 1e-19
BP065719 41 5e-04
AU251673 33 0.19
BP041136 30 1.6
AV767128 30 1.6
TC15142 similar to UP|Q945R8 (Q945R8) Cysteine proteinase (Fragm... 30 1.6
CN825689 29 2.1
TC14312 similar to UP|Q8J174 (Q8J174) RfeF, partial (7%) 28 4.7
TC13256 similar to UP|Q9XI49 (Q9XI49) F9L1.14 protein, partial (... 28 6.2
TC10866 similar to UP|Q9FKP4 (Q9FKP4) Similarity to kinesin heav... 27 8.0
TC17040 27 8.0
>BP052828
Length = 523
Score = 93.2 bits (230), Expect = 1e-19
Identities = 47/130 (36%), Positives = 77/130 (59%), Gaps = 3/130 (2%)
Frame = +2
Query: 506 PCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGELKLTETTLKLADRSDIQPVGNVEDI 565
PC IG + DLGA ++++P ++ L G L+ T ++LA+RS+ +P G VED+
Sbjct: 71 PCTIGDNKYENCMLDLGAGINVMPTSIYNILHPGPLQHTGLIVQLANRSNARPKGVVEDV 250
Query: 566 PVKIEGIDIPTDFMVLDID---EDNECPIILGRPFLATAGAIVDVQNGRIIFQVSDELIG 622
V++ + P DF +LD++ + + PIILGRPF+ TA +DV NG + + D +
Sbjct: 251 LVQVNELIFPADFYILDMEGETKSSRAPIILGRPFMKTAKTKIDVDNGTMSMEFGDIIAK 430
Query: 623 FELENYMKGP 632
F +++ MK P
Sbjct: 431 FNIDDAMKHP 460
>BP065719
Length = 567
Score = 41.2 bits (95), Expect = 5e-04
Identities = 28/97 (28%), Positives = 44/97 (44%)
Frame = +3
Query: 32 AIKENQEAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTAKLRDQ 91
AI EN +++ FP SL A WF +L S+ +W + R F +FF +D
Sbjct: 267 AINEN---LKMKYFPSSLTKNAFTWFTTLAPRSVHTWAQLERIFHEQFFRGECKVSXKD- 434
Query: 92 ITRFNQKDGEYLYEAWERFKEMLRLCPHHGLEKWLIV 128
+ +K E + + RF+ + C H E L+V
Sbjct: 435 LASVKRKPAESIDDYLNRFRMLKSRCFTHVSEHELVV 545
>AU251673
Length = 413
Score = 32.7 bits (73), Expect = 0.19
Identities = 26/88 (29%), Positives = 36/88 (40%), Gaps = 5/88 (5%)
Frame = +2
Query: 42 LHLFPFSLRDRARAWFHSLE----VGSIT-SWDDMRRAFLARFFPPSKTAKLRDQITRFN 96
+ L F L AR W++ L VGS +W D F+ RF P S R
Sbjct: 23 VELASFQLEGVARDWYNVLTRAKPVGSPPWTWADFSAEFMNRFLPQSVRDGFVRDFERLE 202
Query: 97 QKDGEYLYEAWERFKEMLRLCPHHGLEK 124
Q +G + E F + R P+ LE+
Sbjct: 203 QAEGMTVSEYSAHFTHLSRYVPYPLLEE 286
>BP041136
Length = 564
Score = 29.6 bits (65), Expect = 1.6
Identities = 17/52 (32%), Positives = 27/52 (51%), Gaps = 1/52 (1%)
Frame = +2
Query: 285 NNQRVAKKSSLEILLENCIANQ-NKNLQELKNQTGFLNGSLSKLTTKVDSIA 335
N Q + +K SL LL +C+ + + NL N+TGF+ + L +D A
Sbjct: 347 NTQMLTRKPSLTSLLPHCLVSAVSVNLTPSLNETGFILRVCTNLVISIDMSA 502
>AV767128
Length = 267
Score = 29.6 bits (65), Expect = 1.6
Identities = 13/22 (59%), Positives = 16/22 (72%)
Frame = -3
Query: 573 DIPTDFMVLDIDEDNECPIILG 594
DIP D MV +ID+DN+ PI G
Sbjct: 247 DIPIDDMVKEIDQDNDGPIDFG 182
>TC15142 similar to UP|Q945R8 (Q945R8) Cysteine proteinase (Fragment),
partial (36%)
Length = 646
Score = 29.6 bits (65), Expect = 1.6
Identities = 24/70 (34%), Positives = 32/70 (45%)
Frame = -3
Query: 494 PTKLRDPGSFAIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGELKLTETTLKLADR 553
PT+ R P S +P M G ET +C A + + P F R TETTLKL
Sbjct: 212 PTR-RTP*SKCLPQMYGHETPPMYVCIKTALIPIARGPCFTRFAAI*SSSTETTLKLDAE 36
Query: 554 SDIQPVGNVE 563
+ I N++
Sbjct: 35 AAILLFSNLQ 6
>CN825689
Length = 639
Score = 29.3 bits (64), Expect = 2.1
Identities = 15/37 (40%), Positives = 20/37 (53%)
Frame = +1
Query: 325 SKLTTKVDSIATHTKMLKTQISQVAQQVATSSQTPGV 361
SKL T + + T +KT + QV + A SQT GV
Sbjct: 127 SKLATFLQLLEKGTSSIKTSLCQVIESTAAISQTDGV 237
>TC14312 similar to UP|Q8J174 (Q8J174) RfeF, partial (7%)
Length = 622
Score = 28.1 bits (61), Expect = 4.7
Identities = 13/19 (68%), Positives = 14/19 (73%)
Frame = +1
Query: 268 YKNPQGSYGQVAPPGYTNN 286
Y PQGSYGQ APP Y N+
Sbjct: 199 YGAPQGSYGQ-APPSYGNS 252
>TC13256 similar to UP|Q9XI49 (Q9XI49) F9L1.14 protein, partial (22%)
Length = 552
Score = 27.7 bits (60), Expect = 6.2
Identities = 25/82 (30%), Positives = 38/82 (45%), Gaps = 3/82 (3%)
Frame = -1
Query: 9 ISFLDHPLRIQISIFLPSLRLSGAIKENQEA---VRLHLFPFSLRDRARAWFHSLEVGSI 65
+SFLDHPL +++ FL SL + A+ A + LH+ P R L++
Sbjct: 438 LSFLDHPLWLELKAFL-SL*PTAAVPFTASAA*ILCLHIIPCISIFRQ---LPILQLDDS 271
Query: 66 TSWDDMRRAFLARFFPPSKTAK 87
+RR + R P S+T K
Sbjct: 270 R*KSTLRRRLIIRIIPVSRTMK 205
>TC10866 similar to UP|Q9FKP4 (Q9FKP4) Similarity to kinesin heavy chain,
partial (8%)
Length = 537
Score = 27.3 bits (59), Expect = 8.0
Identities = 24/81 (29%), Positives = 43/81 (52%)
Frame = -2
Query: 449 PFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRENSAIIKKPPTKLRDPGSFAIPCM 508
PF+ +L+ K L++ K + + E +A R SA++ + +KLR GS A C
Sbjct: 302 PFSSSLT------KSLRDGTLVAKVLYNLEYRA--RVPSAVLARA-SKLRASGSSANTCF 150
Query: 509 IGSETLNQALCDLGASVSLLP 529
I S+ + + LGA++++ P
Sbjct: 149 ISSKLMFSKMEGLGATLTISP 87
>TC17040
Length = 523
Score = 27.3 bits (59), Expect = 8.0
Identities = 13/48 (27%), Positives = 25/48 (52%), Gaps = 6/48 (12%)
Frame = +2
Query: 587 NECPI------ILGRPFLATAGAIVDVQNGRIIFQVSDELIGFELENY 628
N+C + + G+ L +++ + GRI+ +V D L+G EN+
Sbjct: 35 NQCSLWKIRYRLFGKRILLWTRSLLQLLRGRILVEVGDGLVGVRNENH 178
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,866,490
Number of Sequences: 28460
Number of extensions: 139397
Number of successful extensions: 697
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 692
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 696
length of query: 662
length of database: 4,897,600
effective HSP length: 96
effective length of query: 566
effective length of database: 2,165,440
effective search space: 1225639040
effective search space used: 1225639040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC135416.15


