

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135320.7 + phase: 0
(190 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
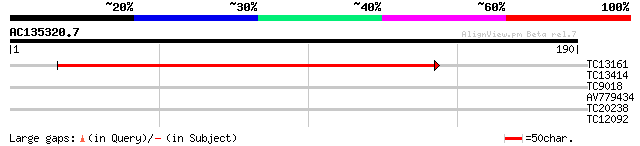
Score E
Sequences producing significant alignments: (bits) Value
TC13161 204 6e-54
TC13414 homologue to UP|Q84TL5 (Q84TL5) At1g75760, partial (39%) 26 5.0
TC9018 weakly similar to AAQ65175 (AAQ65175) At5g62590, partial ... 25 6.5
AV779434 25 6.5
TC20238 homologue to UP|O22427 (O22427) Reversibly glycosylated ... 25 6.5
TC12092 25 8.5
>TC13161
Length = 428
Score = 204 bits (520), Expect = 6e-54
Identities = 99/128 (77%), Positives = 111/128 (86%)
Frame = +2
Query: 17 IDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWCKAI 76
I TQI+LRPL+LSDLDD+M+WTTDEKV KFC+WE YTSK+DGINFIENIATKFLWCKAI
Sbjct: 44 ISTTQISLRPLHLSDLDDVMVWTTDEKVPKFCTWETYTSKEDGINFIENIATKFLWCKAI 223
Query: 77 CINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYL 136
C++DRA+G V LSS + DK R K AELGYVLGSKYWGKG+AT VVKQVVK F EL +L
Sbjct: 224 CLDDRAVGFVHLSSYAEHDKCRIKSAELGYVLGSKYWGKGIATYVVKQVVKDVFSELPHL 403
Query: 137 ERLEALVD 144
ERLEALVD
Sbjct: 404 ERLEALVD 427
>TC13414 homologue to UP|Q84TL5 (Q84TL5) At1g75760, partial (39%)
Length = 544
Score = 25.8 bits (55), Expect = 5.0
Identities = 11/44 (25%), Positives = 23/44 (52%), Gaps = 2/44 (4%)
Frame = +2
Query: 32 LDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFL--WC 73
+D ++W +K++K SW ++ + + +N+ FL WC
Sbjct: 194 MDCGLLWYFYQKLSKLSSWLIFATTTSRASLADNLL*GFLQEWC 325
>TC9018 weakly similar to AAQ65175 (AAQ65175) At5g62590, partial (13%)
Length = 771
Score = 25.4 bits (54), Expect = 6.5
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = +3
Query: 76 ICINDRAIGCVSLSSSSPGDKSRN 99
IC + RA GC+ SS G+ SR+
Sbjct: 303 ICSH*RACGCLPAQQSSSGNSSRS 374
>AV779434
Length = 544
Score = 25.4 bits (54), Expect = 6.5
Identities = 16/47 (34%), Positives = 25/47 (53%)
Frame = -2
Query: 108 LGSKYWGKGVATCVVKQVVKVAFCELSYLERLEALVDVENAGSQRVL 154
L +KY G +AT ++KQ++ S LE LE + + G RV+
Sbjct: 327 LRAKYQGFQIATGIMKQIISHLPRIQSSLEFLEVAIGLHANGIIRVM 187
>TC20238 homologue to UP|O22427 (O22427) Reversibly glycosylated
polypeptide-1, partial (43%)
Length = 466
Score = 25.4 bits (54), Expect = 6.5
Identities = 14/61 (22%), Positives = 25/61 (40%), Gaps = 16/61 (26%)
Frame = -2
Query: 35 LMIWTTDEKVAKFCSWELYTSKDD---------GINFIENI-------ATKFLWCKAICI 78
+++W + + + CSW +T + I FI+ + A K L+ CI
Sbjct: 384 IIVWDVEPQTMRSCSWHTFTETEGVPANKICTFAIRFIQGVEKVRCGWAEKVLYMLLQCI 205
Query: 79 N 79
N
Sbjct: 204 N 202
>TC12092
Length = 286
Score = 25.0 bits (53), Expect = 8.5
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = +2
Query: 73 CKAICINDRAIGCVSLSS 90
C+ ICI AIGCVS+ +
Sbjct: 104 CRNICIFFTAIGCVSIGN 157
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,211,224
Number of Sequences: 28460
Number of extensions: 43230
Number of successful extensions: 166
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 166
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 166
length of query: 190
length of database: 4,897,600
effective HSP length: 85
effective length of query: 105
effective length of database: 2,478,500
effective search space: 260242500
effective search space used: 260242500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC135320.7


