

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135318.3 + phase: 1 /pseudo
(726 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
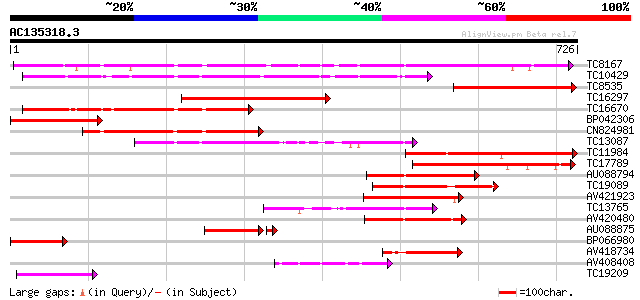
Score E
Sequences producing significant alignments: (bits) Value
TC8167 similar to PIR|JC7519|JC7519 subtilisin-like serine prote... 489 e-139
TC10429 similar to UP|Q9ZR46 (Q9ZR46) P69C protein, partial (52%) 326 9e-90
TC8535 weakly similar to UP|Q9FJF3 (Q9FJF3) Serine protease-like... 266 1e-71
TC16297 weakly similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type pro... 258 2e-69
TC16670 similar to PIR|JC7519|JC7519 subtilisin-like serine prot... 242 2e-64
BP042306 219 1e-57
CN824981 218 2e-57
TC13087 similar to UP|Q9FIF8 (Q9FIF8) Serine protease-like prote... 210 6e-55
TC11984 similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease, ... 173 8e-44
TC17789 similar to PIR|T05768|T05768 subtilisin-like proteinase ... 167 4e-42
AU088794 164 6e-41
TC19089 similar to UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C... 148 3e-36
AV421923 139 1e-33
TC13765 similar to UP|Q9LLL8 (Q9LLL8) Subtilisin-type serine end... 134 5e-32
AV420480 133 9e-32
AU088875 120 1e-28
BP066980 120 1e-27
AV418734 102 2e-22
AV408408 96 2e-20
TC19209 similar to GB|BAB11244.1|10177874|AB010074 serine protea... 90 1e-18
>TC8167 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (65%)
Length = 2760
Score = 489 bits (1258), Expect = e-139
Identities = 295/743 (39%), Positives = 416/743 (55%), Gaps = 25/743 (3%)
Frame = +3
Query: 5 SILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWE 64
S+ E + + ++Y+Y+ NGFAA L+E++A + + V+ ++ + LHTTR+ +
Sbjct: 234 SLTTDSETSDDPLLYAYDTAYNGFAASLDEQQAQTLLGSDSVLGLYEDTLYHLHTTRTPQ 413
Query: 65 FLGLRGNDINSAWQKGRFGE------NTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGN 118
FLGL + W+ R E + IIG +DTGVWPES SF+D G+ IP++WRG
Sbjct: 414 FLGL--DTQTGLWEGHRTLELDQASRDVIIGVLDTGVWPESPSFNDAGMPEIPSRWRG-- 581
Query: 119 ICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGK-----LPRSQQTARDFVGHGTHTLS 173
+ + CNRKLIGAR F++ + G R + RD GHGTHT S
Sbjct: 582 --ECENATDFSSSLCNRKLIGARSFSRGFHMAAGNDGGFGKEREPPSPRDSDGHGTHTAS 755
Query: 174 TAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDG 233
TA G+ V AS+ +GT +G +P+ARVATYKVCWS CF +D+L+ +D+AI DG
Sbjct: 756 TAAGSHVGNASLLGYASGTARGMAPQARVATYKVCWS----DGCFASDILAGMDRAIRDG 923
Query: 234 VDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVF 293
VD++S+S GG S F D I+IGAF A+ R I + SAGN GP+ S+ NVAPW+
Sbjct: 924 VDVLSLSLGGGSGP----YFRDTIAIGAFAAMERGIFVSCSAGNSGPSKASLANVAPWLM 1091
Query: 294 TVAASTLDRDFSSVMTIGNKT-LTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCR 352
TV A TLDRDF + +GNK G SL+ + + +N+ C
Sbjct: 1092TVGAGTLDRDFPASALLGNKKRFAGVSLYSGKGMGAE-----PVGLVYSKGSNQSGILCL 1256
Query: 353 PRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLS 412
P +LDP+ V GK+V CDR G V +G+ AG G+IL N NG+ L+++ H+L
Sbjct: 1257PGSLDPAVVRGKVVLCDR-GLNARVEKGKVVKEAGGIGMILTNTAA-NGEELVADSHLLP 1430
Query: 413 TISYPGNHSRTTGRSL-DIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQP 471
++ R G + + + SD L S T+ +P+PV+A++SSRGPN +
Sbjct: 1431AVAV----GRIVGDQIREYVTSDPNPTAVLSFSG--TVLNVRPSPVVAAFSSRGPNMITK 1592
Query: 472 SILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTL 531
ILKPDV PGVNILA +S S L D+R+ FN+M GTSMSCPH++G L+K
Sbjct: 1593QILKPDVIGPGVNILAGWSEAIGPSGLPQDSRKS-QFNIMSGTSMSCPHISGLGALLKAA 1769
Query: 532 HPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDL 591
HP+WSP+AIKSA+MTTA DNTN P+ DA + P+A+G+GH+ P A+ PGLVYD
Sbjct: 1770HPDWSPSAIKSALMTTAYVHDNTNSPLRDAAGGEFSTPWAHGAGHVNPQKALSPGLVYDA 1949
Query: 592 GIKDYLNFLCASGYNQQLISALNFNMTFTCS-GTSSIDDLNYPSITLPNLGL-----NSV 645
+DY+ FLC+ Y+ + + CS S LNYPS ++ + V
Sbjct: 1950KARDYIAFLCSLDYSPDHLQLIVKRAGVNCSRKLSDPGQLNYPSFSVVFDSVVFDAKKVV 2129
Query: 646 TVTRTVTNVGPPSTYFAKV----QLAGYKIAVVPSSLNFKKIGEKKTFQV--IVQATSVT 699
TRTVTNVG + + V + G I V P+ L F K+GE+K + V + + +
Sbjct: 2130RYTRTVTNVGEAGSVYDVVVDGPSMVG--ITVYPTKLEFGKVGERKRYTVTFVSKKGASD 2303
Query: 700 PRRKYQFGELRWTNGKHIVRSPV 722
+ FG + W N +H VRSPV
Sbjct: 2304DLVRSAFGSITWKNEQHQVRSPV 2372
>TC10429 similar to UP|Q9ZR46 (Q9ZR46) P69C protein, partial (52%)
Length = 1742
Score = 326 bits (835), Expect = 9e-90
Identities = 206/527 (39%), Positives = 283/527 (53%), Gaps = 2/527 (0%)
Frame = +3
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSA 76
+IYSY + GFAA L +EE + + + + + L + + +FLGL+ +
Sbjct: 270 VIYSYKNVLRGFAASLTQEELSAVERKMALFQLILRGCYIVKPLIPPKFLGLQQD--TGV 443
Query: 77 WQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRK 136
W++ FG+ IIG +D+G+ P SFSD GI P P KW+G C L+ CN K
Sbjct: 444 WKESNFGKGVIIGVLDSGITPGHPSFSDVGIPPPPPKWKGR--CDLNV------TACNNK 599
Query: 137 LIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGG 196
LIGAR FN A + NGK + D GHGTHT STA G FV A + GT G
Sbjct: 600 LIGARAFNLAAEAMNGK---KAEAPIDEDGHGTHTASTAAGAFVNYAEVLGNAKGTAAGM 770
Query: 197 SPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDE 256
+P A +A YKVC+ C +D+L+A+D A++DGVD+IS+S G + F D
Sbjct: 771 APHAHLAIYKVCFG----EDCPESDILAALDAAVEDGVDVISISLG---LSEPPPFFNDS 929
Query: 257 ISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-KTL 315
+IGAF A+ + I + +AGN GP S+VN APW+ TV AST+DR + +GN +
Sbjct: 930 TAIGAFAAMQKGIFVSCAAGNSGPFNSSIVNAAPWILTVGASTIDRRIVATAKLGNGQEF 1109
Query: 316 TGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIK 375
G S+F FT A ++ FC +LD S GK+V C+R G I
Sbjct: 1110DGESVF----QPSSFTPTLLPLAYAGKNGKEESAFCANGSLDDSAFRGKVVLCERGGGIA 1277
Query: 376 SVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNH-SRTTGRSLDIIPSD 434
+A+G+E AG +IL N E N +L ++ H L P H S G + +
Sbjct: 1278RIAKGEEVKRAGGAAMILMND-ETNAFSLSADVHAL-----PATHVSYAAGIEIKAYINS 1439
Query: 435 IKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFAS 494
+ T + T+ AP +AS+SSRGPN P ILKPD+ PGVNILAA+ S
Sbjct: 1440TATPTATILFKG-TVIGNSLAPAVASFSSRGPNLPSPGILKPDIIGPGVNILAAWPFPLS 1616
Query: 495 ASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIK 541
S TD++ FN+ GTSMSCPH++G A L+K+ HP+WSPAAIK
Sbjct: 1617NS---TDSK--LTFNIESGTSMSCPHLSGIAALLKSSHPHWSPAAIK 1742
>TC8535 weakly similar to UP|Q9FJF3 (Q9FJF3) Serine protease-like protein,
partial (10%)
Length = 780
Score = 266 bits (679), Expect = 1e-71
Identities = 128/157 (81%), Positives = 141/157 (89%)
Frame = +2
Query: 569 PFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGTSSID 628
PFAYGSGH++PNSA+DPGLVYDL I DYLNFLCASGY+QQLISALNFN TFTCSG+ SI
Sbjct: 5 PFAYGSGHVQPNSAIDPGLVYDLSIVDYLNFLCASGYDQQLISALNFNKTFTCSGSHSIT 184
Query: 629 DLNYPSITLPNLGLNSVTVTRTVTNVGPPSTYFAKVQLAGYKIAVVPSSLNFKKIGEKKT 688
DLNYPSITLPN+ N V VTRTVTNVGPPS YFA QL G+KI VVP+SL+FKKIGEKKT
Sbjct: 185 DLNYPSITLPNIRSNVVNVTRTVTNVGPPSKYFANAQLPGHKIVVVPNSLSFKKIGEKKT 364
Query: 689 FQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPVTVR 725
FQVIVQATSVT RRKY FG+L+WTNG+HIVRSP+TVR
Sbjct: 365 FQVIVQATSVTQRRKYLFGKLQWTNGRHIVRSPITVR 475
>TC16297 weakly similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease,
partial (10%)
Length = 574
Score = 258 bits (659), Expect = 2e-69
Identities = 132/191 (69%), Positives = 158/191 (82%), Gaps = 1/191 (0%)
Frame = +2
Query: 221 DVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGP 280
DVL+AIDQAI DGVD+ SVSAGG S ++EEIFTDE+SIG+ HALA+NIL+VASAGN+GP
Sbjct: 2 DVLAAIDQAITDGVDVNSVSAGGRSIISAEEIFTDEVSIGSVHALAKNILVVASAGNDGP 181
Query: 281 TPGSVVNVAPWVFTVAASTLDRDFSSVMTIG-NKTLTGASLFVNLPPNQDFTIVTSTDAK 339
G+V+NVAPWVFTVAAST+DR+FSS +T+G N+ +TGASLFVNLPPNQ F+++ STDAK
Sbjct: 182 ALGTVLNVAPWVFTVAASTIDREFSSTITLGNNQQITGASLFVNLPPNQLFSLILSTDAK 361
Query: 340 LANATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEI 399
LANATNRDA+ CRP TLDP+KV GK+V C R GKIKSVAEG EALS+GAKG++L
Sbjct: 362 LANATNRDAQLCRPGTLDPAKVKGKVVVCTRGGKIKSVAEGSEALSSGAKGMVLE*SKAK 541
Query: 400 NGKTLLSEPHV 410
EPHV
Sbjct: 542 WKTHFFLEPHV 574
>TC16670 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (42%)
Length = 1183
Score = 242 bits (617), Expect = 2e-64
Identities = 134/299 (44%), Positives = 191/299 (63%), Gaps = 3/299 (1%)
Frame = +2
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGL-RGNDINS 75
++Y+Y+ I+GF+ L EEA + V++V ++ +LHTTR+ FLGL + D+
Sbjct: 332 MLYTYDNAIHGFSTRLTPEEARLLEAQTGVLAVLPERKFELHTTRTPLFLGLDKSADMFP 511
Query: 76 AWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNR 135
A G GE I+G +DTGVWPESKSF D G GP+P+ W+G C+ T+ CN+
Sbjct: 512 A--SGAGGE-VIVGVLDTGVWPESKSFDDTGFGPVPSTWKGE--CESGTNFTAAN--CNK 670
Query: 136 KLIGARFFNKAYQKRNGKLPRSQQTA--RDFVGHGTHTLSTAGGNFVPGASIFNIGNGTI 193
KLIGAR+F+K + G + + ++ RD GHGTHT STA G+ V GAS+F GT
Sbjct: 671 KLIGARYFSKGAEAMLGPIDETAESKSPRDDDGHGTHTASTAAGSVVSGASLFGYAAGTA 850
Query: 194 KGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIF 253
+G +P ARVA YKVCW CF +D+L+AID+AI D V+++S+S GG + + +
Sbjct: 851 RGMAPHARVAAYKVCWK----GGCFSSDILAAIDKAIADNVNVLSLSLGGGMA----DYY 1006
Query: 254 TDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN 312
D ++IGAF A+ + IL+ SAGN GP+ S+ NVAPW+ TV A TLDRDF +++ +GN
Sbjct: 1007RDSVAIGAFAAMEKGILVSCSAGNAGPSAYSLSNVAPWITTVGAGTLDRDFPALVELGN 1183
>BP042306
Length = 549
Score = 219 bits (558), Expect = 1e-57
Identities = 104/119 (87%), Positives = 112/119 (93%)
Frame = +3
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
DLLGS+LGS E AKEAI+YSYNK INGFAA+LEEEEAA+IAKNPKVVSVF+SKEHKLHTT
Sbjct: 192 DLLGSVLGSHEKAKEAILYSYNKHINGFAALLEEEEAARIAKNPKVVSVFVSKEHKLHTT 371
Query: 61 RSWEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNI 119
RSWEFLGLRGN NSAWQKGRFGENTII NIDTGVWPES+SFSD+GIG +PAKWRGGNI
Sbjct: 372 RSWEFLGLRGNGGNSAWQKGRFGENTIIANIDTGVWPESQSFSDKGIGSVPAKWRGGNI 548
>CN824981
Length = 726
Score = 218 bits (556), Expect = 2e-57
Identities = 116/235 (49%), Positives = 155/235 (65%), Gaps = 3/235 (1%)
Frame = +1
Query: 94 GVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGK 153
GVWPE KS D G+ P+P+ W+G Q + N CNRKLIGARFF+K Y+ G
Sbjct: 4 GVWPELKSLDDTGLSPVPSTWKG----QCEAGNNMNSSSCNRKLIGARFFSKGYEATLGP 171
Query: 154 LPRSQQT--ARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSL 211
+ S ++ ARD GHG+HTL+TA G+ V GAS+F + +GT +G + +ARVA YKVCW
Sbjct: 172 IDVSTESRSARDDDGHGSHTLTTAAGSAVAGASLFGLASGTARGMATQARVAAYKVCW-- 345
Query: 212 TDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILL 271
CF +D+ + ID+AI+DGV+IIS+S GG S+ + F D I+IGAF A + IL+
Sbjct: 346 --LGGCFSSDIAAGIDKAIEDGVNIISMSIGGSSA----DYFRDIIAIGAFTANSHGILV 507
Query: 272 VASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGNK-TLTGASLFVNLP 325
SAGN GP+P S+ N APW+ TV A T+DRDF + +T+GN T TGASL+ P
Sbjct: 508 STSAGNGGPSPSSLSNTAPWITTVGAGTIDRDFPAYITLGNNITHTGASLYRGKP 672
>TC13087 similar to UP|Q9FIF8 (Q9FIF8) Serine protease-like protein, partial
(31%)
Length = 1036
Score = 210 bits (535), Expect = 6e-55
Identities = 142/374 (37%), Positives = 209/374 (54%), Gaps = 11/374 (2%)
Frame = +2
Query: 160 TARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFG 219
T RD GHG+HT STA GN V GAS + + GT +G P AR+A YKVC+ C
Sbjct: 20 TVRDIEGHGSHTASTAAGNTVIGASFYGLAQGTARGAVPSARIAAYKVCYP---NHGCDD 190
Query: 220 ADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEG 279
A++L+A D AI DGV+I+SVS G + ++ +D ++IG+FHA+ R IL + +AGN G
Sbjct: 191 ANILAAFDDAIADGVNILSVSLG---TNYCVDLLSDTVAIGSFHAMERGILTLQAAGNSG 361
Query: 280 PTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-KTLTGASLFVNLPPNQDFTIV--TST 336
P S+ +VAPW+ TVAAST+DR F + +GN TL G S+ + F I ++
Sbjct: 362 PASPSICSVAPWLLTVAASTIDRQFIDKVLLGNGMTLIGRSVNAVVSNGTKFPIAKWNAS 541
Query: 337 DAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQ 396
D + ++ C + L V GK+V C EG++ A A GA G I +
Sbjct: 542 DERCKGFSD----LC--QCLGSESVKGKVVLC--EGRMHDEA----AYLDGAVGSITED- 682
Query: 397 PEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSD---IKSGTKLRMSP-----AKT 448
L + + +S ++Y S+++ P D +++ T +P
Sbjct: 683 -------LTNPDNDVSFVTY--------FPSVNLNPKDYAIVQNYTNSAKNPKAEILKSE 817
Query: 449 LNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPF 508
+ + + AP + S+SSRGPN + P I+KPDV+APGV+ILAA+S A+ S +D R+ +
Sbjct: 818 IFKDRSAPKVVSFSSRGPNPLIPEIMKPDVSAPGVDILAAWSPEAAPSEYSSDKRK-VRY 994
Query: 509 NVMQGTSMSCPHVA 522
NV+ GTSM+CPHVA
Sbjct: 995 NVVSGTSMACPHVA 1036
>TC11984 similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease, partial
(30%)
Length = 966
Score = 173 bits (439), Expect = 8e-44
Identities = 96/228 (42%), Positives = 140/228 (61%), Gaps = 9/228 (3%)
Frame = +1
Query: 508 FNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLA 567
+N++ GTSMSCPHV+G AG IK+ H WS +AIKSAIMT+AT +N P++ ++A
Sbjct: 58 YNIISGTSMSCPHVSGLAGSIKSRHTTWSASAIKSAIMTSATQNNNLKAPMT-TDSGSVA 234
Query: 568 NPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNM--TFTCSGTS 625
P+ YG+G + +++ PGLVY+ DYLN+LC G N I ++ + +FTC S
Sbjct: 235 TPYDYGAGEMTTSASFQPGLVYETSTTDYLNYLCYIGLNVTTIKVISRTVPDSFTCPQDS 414
Query: 626 SID---DLNYPSITLPNL-GLNSVTVTRTVTNVG-PPSTYFAKVQLA--GYKIAVVPSSL 678
S D ++NYPSI + N G ++ V+RTVTNVG T ++ + A G I ++P L
Sbjct: 415 SADHVSNINYPSIAITNFNGKGAINVSRTVTNVGEEDETVYSPIVDAPSGVNIKLIPEKL 594
Query: 679 NFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPVTVRR 726
F K +K ++ VI TS T ++ FG L W+NGK+IVRSP + +
Sbjct: 595 PFTKSSQKLSYPVIFSLTSSTSLKEDLFGSLAWSNGKYIVRSPFVLAK 738
>TC17789 similar to PIR|T05768|T05768 subtilisin-like proteinase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (14%)
Length = 940
Score = 167 bits (424), Expect = 4e-42
Identities = 89/220 (40%), Positives = 133/220 (60%), Gaps = 11/220 (5%)
Frame = +2
Query: 516 MSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSG 575
M+CPH++G A L+K+ HP+WSPAA++SA+MTTAT DN + ++D + P+ +G+G
Sbjct: 2 MACPHISGAAALLKSAHPDWSPAAVRSAMMTTATVLDNRKQILTDEATGNSSTPYDFGAG 181
Query: 576 HIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGTSSIDDLNYPSI 635
H+ AMDPGLVYD+ DY+NFLC GY ++I + + S ++LNYPS
Sbjct: 182 HVNLGLAMDPGLVYDITGADYVNFLCGIGYGPRVIQVITRAPVSCPARKPSPENLNYPSF 361
Query: 636 T----LPNLGLNSVTVTRTVTNVGPPSTYF---AKVQLAGYKIAVVPSSLNFKKIGEKKT 688
+ + GL + T RTVTNVGP ++ + + Q+ G + V PS L F + +K++
Sbjct: 362 VAMFPVGSKGLTTKTFIRTVTNVGPANSVYRVSVESQMKGVTVTVKPSRLVFSEAVKKRS 541
Query: 689 FQVIVQATS----VTPRRKYQFGELRWTNGKHIVRSPVTV 724
+ V V A + + P FG L WT+GKH+VRSP+ V
Sbjct: 542 YVVTVAAETRFLKMGPSGAV-FGSLTWTDGKHVVRSPIVV 658
>AU088794
Length = 433
Score = 164 bits (414), Expect = 6e-41
Identities = 80/145 (55%), Positives = 101/145 (69%)
Frame = +2
Query: 457 VMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSM 516
V+AS+S+RGPN P ILKPDV PG+NILAA+ S + +D RR FN++ GTSM
Sbjct: 2 VVASFSARGPNPESPEILKPDVIXPGLNILAAWPDXXGPSGVPSDVRRT-EFNILSGTSM 178
Query: 517 SCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGH 576
+CPHV+G A L+K HP WSPAAIKSA+MTTA T DN + D + ++ F YGSGH
Sbjct: 179 ACPHVSGLAALLKAAHPXWSPAAIKSALMTTAYTVDNKGDAMLDESNGNVSLVFDYGSGH 358
Query: 577 IRPNSAMDPGLVYDLGIKDYLNFLC 601
+ P AMDPGLVYD+ DY++FLC
Sbjct: 359 VHPXKAMDPGLVYDISTYDYVDFLC 433
>TC19089 similar to UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C1, partial
(16%)
Length = 600
Score = 148 bits (374), Expect = 3e-36
Identities = 75/161 (46%), Positives = 100/161 (61%)
Frame = +1
Query: 465 GPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGT 524
GPN + P+ILKPD+ APGV +LAA+S S + D R+ P+N + GTSM+CPH A
Sbjct: 16 GPNPITPNILKPDIAAPGVTVLAAWSPLNPISEVEGDKRK-LPYNAISGTSMACPHAAAA 192
Query: 525 AGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMD 584
A +K+ HPNWSPA IKSA+MTTAT ++ P ++ FAYG+G I P A++
Sbjct: 193 AAYVKSFHPNWSPAMIKSALMTTATPMNSALNPEAE---------FAYGAGQINPVKAVN 345
Query: 585 PGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGTS 625
PGLVYD+ +DY+ FLC GY L+ L + C G S
Sbjct: 346 PGLVYDITEQDYIQFLCGXGYTDTLLQNLTQHKR-XCFGXS 465
>AV421923
Length = 463
Score = 139 bits (351), Expect = 1e-33
Identities = 74/139 (53%), Positives = 97/139 (69%), Gaps = 11/139 (7%)
Frame = +2
Query: 454 PAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQG 513
PAPV+A++SSRGP+ V ++KPDVTAPGV+ILAA+ S S L D RR FN++ G
Sbjct: 50 PAPVVAAFSSRGPSIVGLDVIKPDVTAPGVHILAAWPSKTSPSMLKNDKRRVL-FNIISG 226
Query: 514 TSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISD-AFDKTLAN---- 568
TSMSCPHV+G A LI+++H +WSPAA+KSA+MTTA T +N PISD +++N
Sbjct: 227 TSMSCPHVSGVAALIRSVHRDWSPAAVKSALMTTAYTLNNKGAPISDIGASNSISNSNSN 406
Query: 569 ------PFAYGSGHIRPNS 581
PFA+GSGH+ P S
Sbjct: 407 HSGFAVPFAFGSGHVNPES 463
>TC13765 similar to UP|Q9LLL8 (Q9LLL8) Subtilisin-type serine endopeptidase
XSP1, partial (20%)
Length = 635
Score = 134 bits (337), Expect = 5e-32
Identities = 84/229 (36%), Positives = 125/229 (53%), Gaps = 5/229 (2%)
Frame = +1
Query: 325 PPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACD-----REGKIKSVAE 379
P ++++++ DA + + DA FC +L P+KV GK+V C EG +K
Sbjct: 1 PKRKEYSLINGIDAAKDSKSKDDAGFCYEDSLQPNKVKGKLVYCKLGNWGTEGVVKKF-- 174
Query: 380 GQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDIKSGT 439
G G I+ + + + P + NH T G S + + IKS
Sbjct: 175 -------GGIGSIMESDQYPDLAQIFMAPATIL------NH--TIGES---VTNYIKSTR 300
Query: 440 KLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLI 499
KT + PAP +A++SSRGPN ++LKPD+ APG++ILA+Y+L S +
Sbjct: 301 SPSAVIYKTHEEKCPAPFVATFSSRGPNPGSHNVLKPDIAAPGIDILASYTLRKSITGSE 480
Query: 500 TDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTA 548
DT+ F+++ GTSM+CPHVAG A +K+ HPNW+PAAI+SAI+TTA
Sbjct: 481 GDTQFS-EFSLLSGTSMACPHVAGVAAYVKSFHPNWTPAAIRSAIITTA 624
>AV420480
Length = 412
Score = 133 bits (335), Expect = 9e-32
Identities = 71/130 (54%), Positives = 91/130 (69%)
Frame = +3
Query: 455 APVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGT 514
AP +A++S+RGP+ PSILKPDV APGVNI+AA+ ++L D RR F+VM GT
Sbjct: 30 APAVATFSARGPSFTNPSILKPDVVAPGVNIIAAWPQNLGPTSLPQDLRR-VNFSVMSGT 206
Query: 515 SMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGS 574
SMSCPHV+G A L+ + HP WSPAAIKSAIMTTA D+ +PI D DK A FA G+
Sbjct: 207 SMSCPHVSGIAALVHSAHPKWSPAAIKSAIMTTADVTDHMKRPILDE-DKP-AGVFAIGA 380
Query: 575 GHIRPNSAMD 584
G++ P A++
Sbjct: 381 GNVNPQRALN 410
>AU088875
Length = 322
Score = 120 bits (300), Expect(2) = 1e-28
Identities = 58/77 (75%), Positives = 71/77 (91%), Gaps = 1/77 (1%)
Frame = +3
Query: 250 EEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMT 309
EEIFTDE+SIG+FHALA+NIL+VASAGN+GP G+V+NVAPWVFTVAAST+D +FSS +T
Sbjct: 3 EEIFTDEVSIGSFHALAKNILVVASAGNDGPALGTVLNVAPWVFTVAASTIDXEFSSTIT 182
Query: 310 IG-NKTLTGASLFVNLP 325
+G N+ +TGASLFVNLP
Sbjct: 183 LGNNQQITGASLFVNLP 233
Score = 23.9 bits (50), Expect(2) = 1e-28
Identities = 10/14 (71%), Positives = 12/14 (85%)
Frame = +2
Query: 330 FTIVTSTDAKLANA 343
F+ + STDAKLANA
Sbjct: 248 FSXILSTDAKLANA 289
>BP066980
Length = 472
Score = 120 bits (300), Expect = 1e-27
Identities = 59/74 (79%), Positives = 65/74 (87%)
Frame = +3
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
DLLGS+LGS E AKEAI+YSYNK INGFAA+LEEEEAA+I KNPKVVSVF+SKEH + T
Sbjct: 249 DLLGSVLGSHEKAKEAILYSYNKHINGFAALLEEEEAARIGKNPKVVSVFVSKEH*IDTN 428
Query: 61 RSWEFLGLRGNDIN 74
RSWEFLGLRGN N
Sbjct: 429 RSWEFLGLRGNGGN 470
>AV418734
Length = 287
Score = 102 bits (255), Expect = 2e-22
Identities = 54/102 (52%), Positives = 70/102 (67%)
Frame = +1
Query: 478 VTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSP 537
VTAPG+NILAA+S +A N+ FN++ GTSM+CPHV G A L+K +HP+WSP
Sbjct: 4 VTAPGLNILAAWS--PAAGNM---------FNIVSGTSMACPHVTGIATLVKAVHPSWSP 150
Query: 538 AAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRP 579
+AIKSAIMTTAT D ++ IS ++ AN F YGSG + P
Sbjct: 151 SAIKSAIMTTATILDKHHRHISADPEQRTANAFDYGSGFVNP 276
>AV408408
Length = 429
Score = 95.9 bits (237), Expect = 2e-20
Identities = 62/151 (41%), Positives = 84/151 (55%)
Frame = +3
Query: 340 LANATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEI 399
++N TN C TL P KV GKIV CDR G V +GQ AG G++L N
Sbjct: 3 ISNTTN--GNLCMMGTLSPDKVAGKIVMCDR-GMNARVQKGQVVKGAGGLGMVLSNTAA- 170
Query: 400 NGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMA 459
NG+ L+++ H+L T + + L SD K+ K+ K +P+PV+A
Sbjct: 171 NGEELVADAHLLPTSAVGEKAGNAIKKYLF---SDPKATVKILFEGTKV--GIQPSPVVA 335
Query: 460 SYSSRGPNKVQPSILKPDVTAPGVNILAAYS 490
++SSRGPN + P ILKPD+ APGVNILA +S
Sbjct: 336 AFSSRGPNSITPQILKPDLIAPGVNILAGWS 428
>TC19209 similar to GB|BAB11244.1|10177874|AB010074 serine protease-like
protein {Arabidopsis thaliana;} , partial (12%)
Length = 553
Score = 89.7 bits (221), Expect = 1e-18
Identities = 43/104 (41%), Positives = 61/104 (58%)
Frame = +1
Query: 9 SKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGL 68
S E +E IIYSY+ +G AA L EEA ++ VV++F ++LHTTRS FLGL
Sbjct: 238 SSEEEEERIIYSYHTAFHGVAAKLTPEEAQKLESQEGVVAIFPETRYQLHTTRSPTFLGL 417
Query: 69 RGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPA 112
W + + ++G DTG+WPES+SF+D G P+P+
Sbjct: 418 EKMQSTEIWSEKLSNHDVVVGVSDTGIWPESQSFNDTGFKPVPS 549
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.132 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,368,468
Number of Sequences: 28460
Number of extensions: 147835
Number of successful extensions: 827
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 760
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 764
length of query: 726
length of database: 4,897,600
effective HSP length: 97
effective length of query: 629
effective length of database: 2,136,980
effective search space: 1344160420
effective search space used: 1344160420
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC135318.3


