

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135317.6 + phase: 0
(432 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
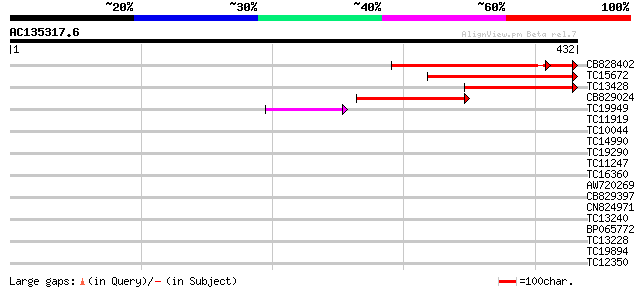
Score E
Sequences producing significant alignments: (bits) Value
CB828402 174 5e-45
TC15672 similar to UP|Q8S298 (Q8S298) Rac-GTP binding protein-li... 99 1e-21
TC13428 weakly similar to UP|Q8S298 (Q8S298) Rac-GTP binding pro... 91 5e-19
CB829024 89 1e-18
TC19949 UP|Q8H0D4 (Q8H0D4) Rac GTPase, partial (33%) 40 7e-04
TC11919 UP|RAC2_LOTJA (Q40220) RAC-like GTP binding protein RAC2... 38 0.003
TC10044 UP|O04369 (O04369) RAC1, complete 37 0.008
TC14990 UP|Q8S2V3 (Q8S2V3) Small G-protein ROP9, complete 36 0.014
TC19290 homologue to UP|Q9SXT7 (Q9SXT7) Rac-type small GTP-bindi... 35 0.031
TC11247 homologue to UP|O82482 (O82482) RAC GTP binding protein ... 34 0.054
TC16360 homologue to UP|Q9M628 (Q9M628) Small GTP binding protei... 34 0.054
AW720269 28 2.2
CB829397 28 2.9
CN824971 28 2.9
TC13240 28 3.8
BP065772 28 3.8
TC13228 homologue to UP|Q9LNN8 (Q9LNN8) F8L10.1 protein, partial... 27 5.0
TC19894 27 5.0
TC12350 weakly similar to UP|Q9FL21 (Q9FL21) Similarity to, part... 27 6.5
>CB828402
Length = 534
Score = 174 bits (441), Expect(2) = 5e-45
Identities = 88/121 (72%), Positives = 101/121 (82%)
Frame = +1
Query: 292 VAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVA 351
VAAFV+DSSD YSWK+SID LEKVV QGE+TG+R P LLIAAKDDLTPFPRA+LDS KV
Sbjct: 1 VAAFVYDSSDEYSWKKSIDFLEKVVRQGEITGYRVPCLLIAAKDDLTPFPRALLDSAKVT 180
Query: 352 QELKIDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALA 411
QEL I+API +S+K DSN+VY+KI+NAAEHPHLSIPETE RKR Q QQ+L I+ L
Sbjct: 181 QELGIEAPIHVSMKLGDSNSVYNKIVNAAEHPHLSIPETEAARKRNQFQQIL---IYGLV 351
Query: 412 G 412
G
Sbjct: 352 G 354
Score = 23.9 bits (50), Expect(2) = 5e-45
Identities = 11/23 (47%), Positives = 16/23 (68%)
Frame = +3
Query: 410 LAGAAMALAGFTARRARANRNSS 432
L+ AMA+ G TA RARA + ++
Sbjct: 345 LSWVAMAIVGVTASRARAVKKNT 413
>TC15672 similar to UP|Q8S298 (Q8S298) Rac-GTP binding protein-like, partial
(18%)
Length = 664
Score = 99.0 bits (245), Expect = 1e-21
Identities = 48/115 (41%), Positives = 78/115 (67%), Gaps = 1/115 (0%)
Frame = +2
Query: 319 GELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQELKIDAPIRLSIKSDDSNNVYSKIIN 378
GE TG P L++A KDDL F A+ ++ +V+Q++ ++API +S+K D N+++ +I+
Sbjct: 2 GEDTGFEVPCLIVATKDDLDSFTLAIEETTRVSQDMGVEAPIPISVKLGDFNSLFRRIVT 181
Query: 379 AAEHPHLSIPETEFVRKRKQHQQLLHTFIFALA-GAAMALAGFTARRARANRNSS 432
AAEHPHLSIPETE R RK + +L++ + ++ GAA+A+ G A R + R ++
Sbjct: 182 AAEHPHLSIPETEAGRTRKHYHRLINRSLMVVSVGAAVAVVGLAAYRVYSARKNT 346
>TC13428 weakly similar to UP|Q8S298 (Q8S298) Rac-GTP binding protein-like,
partial (9%)
Length = 560
Score = 90.5 bits (223), Expect = 5e-19
Identities = 47/87 (54%), Positives = 65/87 (74%), Gaps = 1/87 (1%)
Frame = +2
Query: 347 SVKVAQELKIDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLL-HT 405
SVKV +EL I+ PI +S+KS DS+NVY+ II+AAEHPHLS+PETE RKRK +++LL ++
Sbjct: 2 SVKVTEELGIEGPIHVSMKSGDSSNVYNTIISAAEHPHLSLPETEIGRKRKPYRRLLQNS 181
Query: 406 FIFALAGAAMALAGFTARRARANRNSS 432
I+A G A+A+ G A R A + +S
Sbjct: 182 LIYASVGTAVAVVGVAAFRVYAMKKNS 262
>CB829024
Length = 473
Score = 89.4 bits (220), Expect = 1e-18
Identities = 41/86 (47%), Positives = 58/86 (66%)
Frame = +3
Query: 265 KTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGH 324
K L+++EIPE EV L+NK+ LA+CD+A FVHD SD SWK + +LL ++ GE TG
Sbjct: 3 KFLVMKEIPEGEVRKLLANKESLASCDIAVFVHDRSDESSWKAATELLVEIAGHGEDTGF 182
Query: 325 RFPSLLIAAKDDLTPFPRAVLDSVKV 350
P L++A KDDL F A+ ++ +V
Sbjct: 183 EVPCLIVATKDDLDSFTLAIEETTRV 260
>TC19949 UP|Q8H0D4 (Q8H0D4) Rac GTPase, partial (33%)
Length = 549
Score = 40.0 bits (92), Expect = 7e-04
Identities = 16/62 (25%), Positives = 30/62 (47%)
Frame = +1
Query: 196 LRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAAN 255
LR ++ + +K + + +C G GK+ LL + F DY PT + ++AN
Sbjct: 313 LREKKKKKKKKKKKMSASRFIKCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSAN 492
Query: 256 II 257
++
Sbjct: 493 VV 498
>TC11919 UP|RAC2_LOTJA (Q40220) RAC-like GTP binding protein RAC2, complete
Length = 974
Score = 38.1 bits (87), Expect = 0.003
Identities = 53/228 (23%), Positives = 91/228 (39%), Gaps = 9/228 (3%)
Frame = +2
Query: 178 YSLANLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLG 237
+SL + R+ S A+R + R T R + +C G GK+ +L +
Sbjct: 20 FSLVLFLVSSERNMESFAVR*GKSVQKRRLKMSTARFI-KCVTVGDGAVGKTCMLISYTS 196
Query: 238 RPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVH 297
F DY PT + ++AN++ + G+ L L + E N L A DV
Sbjct: 197 NTFPTDYVPTVFDNFSANVV--VDGSTVNLGLWDTAGQEDYNRLRPLSYRGA-DVFLLAF 367
Query: 298 DSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLD-------SVKV 350
S++ ++ +K + + P +L+ K DL + ++D +
Sbjct: 368 SLLSRASYE---NISKKWIPELRHYAPTVPIVLVGTKLDLREDRQYLIDHPGATPITTAQ 538
Query: 351 AQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKR 396
+ELK I A + L S NV + +AA L P+ + RK+
Sbjct: 539 GEELKKAIGAAVYLECSSKTQQNV-KAVFDAAIKVVLQPPKPKKKRKK 679
>TC10044 UP|O04369 (O04369) RAC1, complete
Length = 1292
Score = 36.6 bits (83), Expect = 0.008
Identities = 48/195 (24%), Positives = 77/195 (38%), Gaps = 10/195 (5%)
Frame = +3
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G+ L L + E
Sbjct: 234 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VDGSTVNLGLWDTAGQE 407
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + P +L+ K
Sbjct: 408 DYNRLRPLSYRGADVFILAFSLISKASYE-----NIAKKWIPELRHYAPGVPIILVGTKL 572
Query: 336 D-------LTPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
D L P AV + +EL+ I AP + S NV + +AA L
Sbjct: 573 DLRDDKHFLADHPGAVPITTAQGEELRKLIGAPAYIECSSKTQQNV-KAVFDAAIKVVLQ 749
Query: 387 IPETEFVRKRKQHQQ 401
P+ +K+K+ Q
Sbjct: 750 PPKQ---KKKKREAQ 785
>TC14990 UP|Q8S2V3 (Q8S2V3) Small G-protein ROP9, complete
Length = 1029
Score = 35.8 bits (81), Expect = 0.014
Identities = 46/192 (23%), Positives = 77/192 (39%), Gaps = 10/192 (5%)
Frame = +3
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G+ L L + E
Sbjct: 249 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VNGSIVNLGLWDTAGQE 422
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + + P +L+ K
Sbjct: 423 DYNRLRPLSYRGADVFILAFSLISKASYE-----NVSKKWIPELKHYAPGVPIILVGTKL 587
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +EL+ I+AP + S NV + +AA L
Sbjct: 588 DLRDDKQFCIDHPGAVPITTAQGEELRKLINAPAYIECSSKTQENV-KAVFDAAIRVVLQ 764
Query: 387 IPETEFVRKRKQ 398
P+ + + + Q
Sbjct: 765 PPKQKKKKNKAQ 800
>TC19290 homologue to UP|Q9SXT7 (Q9SXT7) Rac-type small GTP-binding protein,
partial (72%)
Length = 427
Score = 34.7 bits (78), Expect = 0.031
Identities = 13/41 (31%), Positives = 21/41 (50%)
Frame = +3
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
+C G GK+ LL + F DY PT + ++AN++
Sbjct: 15 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV 137
>TC11247 homologue to UP|O82482 (O82482) RAC GTP binding protein ARAC8
(At3g48040), partial (60%)
Length = 559
Score = 33.9 bits (76), Expect = 0.054
Identities = 13/47 (27%), Positives = 21/47 (44%)
Frame = +1
Query: 211 TERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
T +C G GK+ +L F DY PT + ++AN++
Sbjct: 193 TASRFIKCVTVGDGAVGKTCMLICYTSNKFPTDYIPTVFDNFSANVV 333
>TC16360 homologue to UP|Q9M628 (Q9M628) Small GTP binding protein RACDP,
partial (64%)
Length = 479
Score = 33.9 bits (76), Expect = 0.054
Identities = 12/41 (29%), Positives = 21/41 (50%)
Frame = +3
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
+C G GK+ +L + F DY PT + ++AN++
Sbjct: 123 KCVTVGDGAVGKTCMLISYTSNTFPTDYVPTVFDNFSANVV 245
>AW720269
Length = 524
Score = 28.5 bits (62), Expect = 2.2
Identities = 13/30 (43%), Positives = 16/30 (53%)
Frame = +3
Query: 220 VFGSKTAGKSALLRALLGRPFSNDYTPTTV 249
V G GKS LLR + GR D+ P T+
Sbjct: 102 VVGPSGTGKSTLLRIIAGRVKDRDFDPKTI 191
>CB829397
Length = 550
Score = 28.1 bits (61), Expect = 2.9
Identities = 15/50 (30%), Positives = 29/50 (58%), Gaps = 1/50 (2%)
Frame = -3
Query: 381 EHPHLSIPETEFVRKRKQHQQLLH-TFIFALAGAAMALAGFTARRARANR 429
++PH + PE E ++R+ HQQ +H ++F+ +GF+ R + +R
Sbjct: 272 DNPHQTSPEAEQFQERRAHQQDIHQMYLFSYP----LESGFSGREQQYHR 135
>CN824971
Length = 683
Score = 28.1 bits (61), Expect = 2.9
Identities = 15/50 (30%), Positives = 29/50 (58%), Gaps = 1/50 (2%)
Frame = -2
Query: 381 EHPHLSIPETEFVRKRKQHQQLLH-TFIFALAGAAMALAGFTARRARANR 429
++PH + PE E ++R+ HQQ +H ++F+ +GF+ R + +R
Sbjct: 253 DNPHQTSPEAEQFQERRAHQQDIHQMYLFSYP----LESGFSGREQQYHR 116
>TC13240
Length = 518
Score = 27.7 bits (60), Expect = 3.8
Identities = 12/39 (30%), Positives = 18/39 (45%)
Frame = +3
Query: 270 REIPEDEVSNFLSNKDCLAACDVAAFVHDSSDGYSWKRS 308
RE + E FL N A C++ F+H S Y ++
Sbjct: 261 REEAQSETDPFLINLHTSAYCNIPTFIHSKSSPYQTNKN 377
>BP065772
Length = 495
Score = 27.7 bits (60), Expect = 3.8
Identities = 18/67 (26%), Positives = 29/67 (42%)
Frame = +1
Query: 173 LLDPRYSLANLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALL 232
L DP ++ L+ + + A+ ED K + + G+ AGKSA L
Sbjct: 91 LTDPMEAIEELVQLSDSMRQATAVLADDEDIEDSKRRPS--TFLHVVALGNVGAGKSAAL 264
Query: 233 RALLGRP 239
+L+G P
Sbjct: 265 NSLIGHP 285
>TC13228 homologue to UP|Q9LNN8 (Q9LNN8) F8L10.1 protein, partial (11%)
Length = 421
Score = 27.3 bits (59), Expect = 5.0
Identities = 18/49 (36%), Positives = 26/49 (52%)
Frame = +2
Query: 222 GSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILR 270
G ++ GKS+LL ALLG F N+ E+ GT++ LIL+
Sbjct: 242 GGQSDGKSSLLEALLGFRF--------------NVREVEMGTRRPLILQ 346
>TC19894
Length = 583
Score = 27.3 bits (59), Expect = 5.0
Identities = 15/46 (32%), Positives = 27/46 (58%), Gaps = 3/46 (6%)
Frame = +1
Query: 78 GNDLKLRDDFLPVPSKK---ASDQSVELSGAAIEFLKGVFRLVDTD 120
G+DL+ RD+ LP+PS++ AS ++ A I+ V + ++D
Sbjct: 199 GSDLEGRDEILPLPSRQLPVASSHALAEEHAGIQLAGSVCKRKESD 336
>TC12350 weakly similar to UP|Q9FL21 (Q9FL21) Similarity to, partial (12%)
Length = 458
Score = 26.9 bits (58), Expect = 6.5
Identities = 10/24 (41%), Positives = 16/24 (66%)
Frame = -2
Query: 382 HPHLSIPETEFVRKRKQHQQLLHT 405
HPH +P + +K +Q+Q+ LHT
Sbjct: 76 HPHQPVPPGKGAKKSEQYQKDLHT 5
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,246,668
Number of Sequences: 28460
Number of extensions: 74261
Number of successful extensions: 323
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 323
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 323
length of query: 432
length of database: 4,897,600
effective HSP length: 93
effective length of query: 339
effective length of database: 2,250,820
effective search space: 763027980
effective search space used: 763027980
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC135317.6


