

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
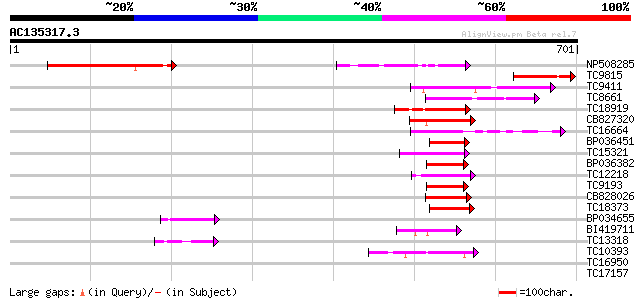
Query= AC135317.3 + phase: 0
(701 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
NP508285 myc-like regulatory protein [Lotus japonicus] 169 1e-42
TC9815 weakly similar to UP|Q9FEA1 (Q9FEA1) Anthocyanin 1, parti... 125 3e-29
TC9411 similar to UP|Q41102 (Q41102) Phaseolin G-box binding pro... 90 1e-18
TC8661 weakly similar to UP|O81348 (O81348) Symbiotic ammonium t... 86 3e-17
TC18919 similar to GB|AAN73298.1|25141207|BT002301 At4g16430/dl4... 76 2e-14
CB827320 62 4e-10
TC16664 similar to UP|BAD09830 (BAD09830) DNA-binding protein-li... 58 6e-09
BP036451 57 1e-08
TC15321 similar to UP|BAD09664 (BAD09664) BHLH protein family-li... 57 1e-08
BP036382 48 5e-06
TC12218 47 1e-05
TC9193 similar to GB|AAN28890.1|23506061|AY143951 At4g02590/T10P... 46 2e-05
CB828026 45 4e-05
TC18373 45 5e-05
BP034655 44 7e-05
BI419711 42 3e-04
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 42 4e-04
TC10393 41 7e-04
TC16950 similar to UP|RGP1_HUMAN (P46060) Ran GTPase-activating ... 40 0.001
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 40 0.002
>NP508285 myc-like regulatory protein [Lotus japonicus]
Length = 1529
Score = 169 bits (428), Expect = 1e-42
Identities = 88/165 (53%), Positives = 112/165 (67%), Gaps = 5/165 (3%)
Frame = +3
Query: 47 NGSIKTRKTVQPMEVSAEEASLQRSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESE 106
NG IK KTVQ ME A++ LQRS+QLRELY+ L GE +P +RP ASLSPEDL++SE
Sbjct: 3 NGDIKQMKTVQTMETKADKIGLQRSEQLRELYKFLLVGEADPLAKRPSASLSPEDLSDSE 182
Query: 107 WFYLMCVSFSFPPGVGLPGRAYTKRQHIWLTGANEVDSKIFSRAILAK-----TVVCIPV 161
W+YL+C+SF F P LPG+A + +WL A + DSK FSR++LAK TVVC P
Sbjct: 183 WYYLVCMSFVFYPNQSLPGKALETGETVWLCNAQQADSKFFSRSLLAKSASIQTVVCFPY 362
Query: 162 LDGVVEFGTTDKVQEDLNFIKHVKSFFLDGHSLPPKPALSEHSTS 206
L GV+E GTT+ V ED N I+HVK+ FL+ KP S+ S+S
Sbjct: 363 LGGVIEIGTTELVSEDPNLIQHVKACFLE----ISKPTCSDKSSS 485
Score = 54.7 bits (130), Expect = 5e-08
Identities = 44/165 (26%), Positives = 78/165 (46%)
Frame = +3
Query: 405 EDTHYSQTVSTILQNQSTQWTIDSSPSINYITSSNQSSFTNWTNHHNFHPLPPPETTTTT 464
E +Y++T+ +L N S S +++SSF W P
Sbjct: 924 EVLYYTRTLCAVLGNSS---------SFAQNLCASKSSFVKWNKGGVSERKWP-----RL 1061
Query: 465 SQCLLKYILFTVPYLHTKNHDETSPQTHDAGVDPSSKLRGKGTPQDELSANHVLAERRRR 524
Q +LK LF VP++H + Q + + +SKL D N V ++++R
Sbjct: 1062 QQMMLKKTLFDVPFMHL-SCSSLKLQKENGRKEWTSKLENA----DNFMGN-VFSDKKRE 1223
Query: 525 EKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLET 569
+ +L+S+ P +++K S+LGDTI+YLK+L ++++LE+
Sbjct: 1224 SR---NIQVLKSVAPSACEVEKISVLGDTIQYLKKLEARVEELES 1349
>TC9815 weakly similar to UP|Q9FEA1 (Q9FEA1) Anthocyanin 1, partial (11%)
Length = 634
Score = 125 bits (313), Expect = 3e-29
Identities = 68/77 (88%), Positives = 71/77 (91%)
Frame = +1
Query: 623 IESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRAKVKENGGNGK 682
IESDALLE+EC HREGLLLDVM MLRE+RIEV GVQSSLNNGVFVAELRAKVKE +GK
Sbjct: 1 IESDALLELECPHREGLLLDVMQMLREMRIEVTGVQSSLNNGVFVAELRAKVKEY-VSGK 177
Query: 683 KVSIVEVKRALNQIIPH 699
KVSIVEVKRALNQIIPH
Sbjct: 178 KVSIVEVKRALNQIIPH 228
>TC9411 similar to UP|Q41102 (Q41102) Phaseolin G-box binding protein PG2
(Fragment), partial (30%)
Length = 705
Score = 90.1 bits (222), Expect = 1e-18
Identities = 61/191 (31%), Positives = 96/191 (49%), Gaps = 12/191 (6%)
Frame = +2
Query: 496 VDPSSKLRGKGTPQD---ELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGD 552
V+P + R +G E NHV AER+RREKLN+RF LR++VP V+KMDKAS+LGD
Sbjct: 122 VEPEKRPRKRGRKPANGREEPLNHVEAERQRREKLNQRFYALRAVVPNVSKMDKASLLGD 301
Query: 553 TIEYLKQLRRKIQDLETRNRQI---------ETEQQSRSGVTVLVGPTDKKKVRIVEECG 603
I Y+ +L+ K+Q E+ + E E+ S + P +K K
Sbjct: 302 AISYITELKTKLQSSESDKTGLQKQFDAMKKELEKTSEQSSSPTPPPPNKNK------SF 463
Query: 604 ATRAKAVETEVVSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNN 663
++ + + +V + V II DA++ ++C + +M L EL +EV S+ N
Sbjct: 464 SSSSSSSNQILVEDIDVKIIGWDAMIRVQCSKKNHPAAILMAALMELDLEVNHASVSVVN 643
Query: 664 GVFVAELRAKV 674
+ + K+
Sbjct: 644 DTMIQQATVKM 676
>TC8661 weakly similar to UP|O81348 (O81348) Symbiotic ammonium
transporter, partial (34%)
Length = 1007
Score = 85.5 bits (210), Expect = 3e-17
Identities = 48/141 (34%), Positives = 80/141 (56%)
Frame = +1
Query: 515 NHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQI 574
+H++AERRRREKL++ I L +L+P + KMDKAS+LGD I+Y+K L+ +++ LE +N+
Sbjct: 28 DHIIAERRRREKLSQSLIALAALIPGLKKMDKASVLGDAIKYVKVLKERLRLLEEQNK-- 201
Query: 575 ETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVETEVVSSVQVSIIESDALLEIECL 634
++ V V+ P C E E + V+ + E D LL + C
Sbjct: 202 ---NRAMESVVVVNKPQISNDDNSSSSCDDGTIIGSE-EALPHVEARVSEKDVLLRLHCK 369
Query: 635 HREGLLLDVMVMLRELRIEVI 655
++GLLL ++ ++ L + V+
Sbjct: 370 KQKGLLLKILFEIQNLHLFVV 432
>TC18919 similar to GB|AAN73298.1|25141207|BT002301 At4g16430/dl4240w
{Arabidopsis thaliana;}, partial (29%)
Length = 604
Score = 75.9 bits (185), Expect = 2e-14
Identities = 44/94 (46%), Positives = 60/94 (63%)
Frame = +3
Query: 476 VPYLHTKNHDETSPQTHDAGVDPSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILR 535
V L N D +SP D P + R ++E NHV AER+RREKLN+RF LR
Sbjct: 303 VSSLDLGNEDASSPHLDDR--KPRKRGRKPANGREE-PLNHVEAERQRREKLNQRFYALR 473
Query: 536 SLVPFVTKMDKASILGDTIEYLKQLRRKIQDLET 569
++VP ++KMDKAS+LGD I ++ L++KI+ LET
Sbjct: 474 AVVPNISKMDKASLLGDAITHITDLQKKIKLLET 575
>CB827320
Length = 522
Score = 61.6 bits (148), Expect = 4e-10
Identities = 31/85 (36%), Positives = 53/85 (61%), Gaps = 3/85 (3%)
Frame = +1
Query: 495 GVDPSSKLRGKGTPQDELSA---NHVLAERRRREKLNERFIILRSLVPFVTKMDKASILG 551
G+D + +P S+ ++++ER RR+KLNER LRS+VP ++KMDKASI+
Sbjct: 160 GLDEALSYYDSSSPDGAASSAASKNIVSERNRRKKLNERLFALRSVVPNISKMDKASIIK 339
Query: 552 DTIEYLKQLRRKIQDLETRNRQIET 576
D IEY++ L + + ++ ++E+
Sbjct: 340 DAIEYIEHLHEQEKIIQAEIMELES 414
>TC16664 similar to UP|BAD09830 (BAD09830) DNA-binding protein-like, partial
(22%)
Length = 1067
Score = 57.8 bits (138), Expect = 6e-09
Identities = 51/195 (26%), Positives = 86/195 (43%), Gaps = 3/195 (1%)
Frame = +1
Query: 496 VDPSSKLRGKG-TPQDELSA--NHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGD 552
V+ +L KG +P+ + A NH AERRRR ++N LR+++P KMDKAS+L +
Sbjct: 223 VEAPMQLERKGVSPERSIEALRNHSEAERRRRARINAHLDTLRTVIPGANKMDKASLLAE 402
Query: 553 TIEYLKQLRRKIQDLETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVET 612
I +LK+L+ Q+ G+ + P D ++R+ E+ G
Sbjct: 403 VITHLKELK-------------TNAAQASEGLMI---PKDNDEIRVEEQEGGLNG----- 519
Query: 613 EVVSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRA 672
S++ S+ C ++ GLL D+ L L + + +RA
Sbjct: 520 -FPYSIRASLC---------CEYKPGLLSDIRQALDALHLMI---------------MRA 624
Query: 673 KVKENGGNGKKVSIV 687
++ GG K V ++
Sbjct: 625 EIATLGGRMKNVFVI 669
>BP036451
Length = 511
Score = 57.0 bits (136), Expect = 1e-08
Identities = 24/49 (48%), Positives = 40/49 (80%)
Frame = +2
Query: 520 ERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLE 568
E+ RRE+LN ++ ILRSL+P TK+D+AS++GD I+Y+++L R + +L+
Sbjct: 335 EKERREQLNGKYKILRSLIPSPTKIDRASVVGDAIDYIRELIRTLNELK 481
>TC15321 similar to UP|BAD09664 (BAD09664) BHLH protein family-like, partial
(23%)
Length = 1593
Score = 56.6 bits (135), Expect = 1e-08
Identities = 33/89 (37%), Positives = 49/89 (54%), Gaps = 2/89 (2%)
Frame = +2
Query: 482 KNHDETSPQTHDAGVDPSSKLRGKGTPQ-DELSANHVLAERRRREKLNERFIILRSLVPF 540
+ D S Q H P++ K T + + + H + E+RRR K+NERF ILR ++P
Sbjct: 287 EEEDFGSSQKHGPSSAPNTNKDAKATDKASAIRSKHSVTEQRRRSKINERFQILRDIIPH 466
Query: 541 -VTKMDKASILGDTIEYLKQLRRKIQDLE 568
K D AS L + IEY++ L+ K+Q E
Sbjct: 467 SEQKRDTASFLLEVIEYVQYLQEKVQKYE 553
>BP036382
Length = 511
Score = 48.1 bits (113), Expect = 5e-06
Identities = 24/52 (46%), Positives = 36/52 (69%)
Frame = +3
Query: 516 HVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDL 567
H +AER RRE++ ER L+ LVP K DKAS+L + I+Y+K L+ +++ L
Sbjct: 135 HSIAERLRRERIAERMKALQELVPNANKTDKASMLDEIIDYVKFLQLQVKVL 290
>TC12218
Length = 737
Score = 47.0 bits (110), Expect = 1e-05
Identities = 27/80 (33%), Positives = 45/80 (55%), Gaps = 2/80 (2%)
Frame = +2
Query: 498 PSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVP--FVTKMDKASILGDTIE 555
PS +LR + +S + E+ RR+KLNERF+ L S++ K DKA+I+ D +
Sbjct: 236 PSKRLR----TESSVSGSKACREKLRRDKLNERFLELSSILEPGRQPKTDKAAIISDAVR 403
Query: 556 YLKQLRRKIQDLETRNRQIE 575
+ QLR + + L+ N ++
Sbjct: 404 VVTQLRNEAEKLKEMNNDLQ 463
>TC9193 similar to GB|AAN28890.1|23506061|AY143951 At4g02590/T10P11_13
{Arabidopsis thaliana;}, partial (59%)
Length = 1474
Score = 45.8 bits (107), Expect = 2e-05
Identities = 22/52 (42%), Positives = 36/52 (68%)
Frame = +1
Query: 516 HVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDL 567
H +AER RRE++ ER + LVP V K D+A++L + ++Y+K LR +++ L
Sbjct: 466 HSIAERLRRERIAERIRA*QELVPSVNKTDRAAMLDEIVDYVKFLRLQVKVL 621
>CB828026
Length = 542
Score = 45.1 bits (105), Expect = 4e-05
Identities = 24/57 (42%), Positives = 39/57 (68%), Gaps = 1/57 (1%)
Frame = +2
Query: 515 NHVLAERRRREKLNERFIILRSLVPFVTKM-DKASILGDTIEYLKQLRRKIQDLETR 570
+H LAER RREK++ER L+ LVP +K+ KA +L + I Y++ L+R+++ L +
Sbjct: 146 SHSLAERVRREKISERMKFLQDLVPGCSKVTGKAVMLDEIINYVQSLQRQVEFLSMK 316
>TC18373
Length = 437
Score = 44.7 bits (104), Expect = 5e-05
Identities = 25/57 (43%), Positives = 36/57 (62%), Gaps = 2/57 (3%)
Frame = +1
Query: 520 ERRRREKLNERFIILRSLVP--FVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQI 574
E++RREKLNERF L S++ K DK +IL D I L QL+ + Q+L+ N ++
Sbjct: 262 EKQRREKLNERFCELSSVLEPGRPVKTDKPAILDDAIRVLSQLKTEAQELKETNEKL 432
>BP034655
Length = 517
Score = 44.3 bits (103), Expect = 7e-05
Identities = 24/73 (32%), Positives = 40/73 (53%)
Frame = +1
Query: 187 FFLDGHSLPPKPALSEHSTSNPTSSTDHIPTIMYTMADPPSTNPNQDDMDEDEEEEEEED 246
+FL PP P + S + I T AD ++ ++++ +E+EEEEEEE+
Sbjct: 13 YFLAAFKRPP-PLFATPMASPSEFHRNEIAAAHLTAADFGLSDDDEEEEEEEEEEEEEEE 189
Query: 247 EEDDEVESESEDE 259
EE++E E E ++E
Sbjct: 190 EEEEEEEEEEDEE 228
Score = 37.7 bits (86), Expect = 0.006
Identities = 14/27 (51%), Positives = 23/27 (84%)
Frame = +1
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESED 258
+++ +E+EEEEEEE++E+D+ E E ED
Sbjct: 175 EEEEEEEEEEEEEEEDEEDDEEDEDED 255
Score = 35.0 bits (79), Expect = 0.041
Identities = 16/28 (57%), Positives = 23/28 (82%)
Frame = +1
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
+++ +E+EEEEEEE EED+E + E EDE
Sbjct: 172 EEEEEEEEEEEEEE-EEDEEDDEEDEDE 252
>BI419711
Length = 539
Score = 42.0 bits (97), Expect = 3e-04
Identities = 33/97 (34%), Positives = 49/97 (50%), Gaps = 17/97 (17%)
Frame = +1
Query: 479 LHTKNHDETSPQTHDAGV-DPSS----------KLRGKGTPQDELSAN----HVLAERRR 523
LH+ N D +S + +A D SS + R K E N H+ ER R
Sbjct: 247 LHSNNWDSSSHTSPEACTFDQSSPTPFSSSRRKRRRTKSAKNMEEIENQRMTHIAVERNR 426
Query: 524 REKLNERFIILRSLVP--FVTKMDKASILGDTIEYLK 558
R+++NE LRSL+P +V + D+ASI+G I ++K
Sbjct: 427 RKQMNEYLSALRSLMPSSYVQRGDQASIIGGAINFVK 537
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 41.6 bits (96), Expect = 4e-04
Identities = 23/79 (29%), Positives = 38/79 (47%)
Frame = +3
Query: 180 FIKHVKSFFLDGHSLPPKPALSEHSTSNPTSSTDHIPTIMYTMADPPSTNPNQDDMDEDE 239
++ H + DG PPKP +++ + D + + DD D+D+
Sbjct: 189 WLVHDPEIYPDG---PPKPVQTDNEEEEDDDEEE----------DDEDEDDDDDDDDDDD 329
Query: 240 EEEEEEDEEDDEVESESED 258
EEEE+D++DDE E SE+
Sbjct: 330 GEEEEDDDDDDEEEEGSEE 386
>TC10393
Length = 1063
Score = 40.8 bits (94), Expect = 7e-04
Identities = 44/161 (27%), Positives = 66/161 (40%), Gaps = 25/161 (15%)
Frame = +1
Query: 444 TNWTNHHNF-HPLPPPETTTTTSQCLLKYILFTVPYLHTKNHDET--------SPQTHDA 494
TNW + +P PE T T FT P N S D+
Sbjct: 208 TNWLFDYGLIDDIPAPEVTFTVPPSG-----FTWPSSQPLNSSSNVVGVEIDGSLGDSDS 372
Query: 495 GVDPSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVP--FVTKMDKASILGD 552
+P SK R + S+ E+ RR+KLN++F+ L S++ K DKA+IL D
Sbjct: 373 LKEPGSKKRVRSESCSATSSK-ACREKLRRDKLNDKFVELGSILEPGRPPKTDKAAILID 549
Query: 553 TIEYLKQLR--------------RKIQDLETRNRQIETEQQ 579
+ + QLR KI++L+T ++ E+Q
Sbjct: 550 AVRMVTQLRGEAQKMKDTNMGLQEKIKELKTEKNELRDEKQ 672
>TC16950 similar to UP|RGP1_HUMAN (P46060) Ran GTPase-activating protein 1,
partial (3%)
Length = 643
Score = 40.4 bits (93), Expect = 0.001
Identities = 16/41 (39%), Positives = 29/41 (70%)
Frame = +1
Query: 219 MYTMADPPSTNPNQDDMDEDEEEEEEEDEEDDEVESESEDE 259
M T PP +P ++D +E+E++EEEE+E+ D+ ES+ ++
Sbjct: 289 MKTPLPPPIADPPEEDEEEEEKKEEEEEEDPDKEESDKPEQ 411
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein
1-like protein 3, partial (81%)
Length = 1298
Score = 39.7 bits (91), Expect = 0.002
Identities = 15/42 (35%), Positives = 28/42 (65%)
Frame = +2
Query: 224 DPPSTNPNQDDMDEDEEEEEEEDEEDDEVESESEDETGGRIR 265
D DD DE+EE+++++DE+D+E E +S+ ++G + R
Sbjct: 788 DGDEDEDEDDDDDEEEEDDDDDDEDDEEGEGKSKSKSGSKAR 913
Score = 38.9 bits (89), Expect = 0.003
Identities = 15/59 (25%), Positives = 34/59 (57%)
Frame = +2
Query: 231 NQDDMDEDEEEEEEEDEEDDEVESESEDETGGRIRRATSMTAIVEPSELMQIEMPDDIR 289
+ +D DEDE+E++++DEE+++ + + ED+ G + + + P + E P + +
Sbjct: 779 DDEDGDEDEDEDDDDDEEEEDDDDDDEDDEEGEGKSKSKSGSKARPGKDQPTERPPECK 955
Score = 35.0 bits (79), Expect = 0.041
Identities = 12/27 (44%), Positives = 22/27 (81%)
Frame = +2
Query: 233 DDMDEDEEEEEEEDEEDDEVESESEDE 259
++ DED +E+E+ED++DDE E + +D+
Sbjct: 773 EEDDEDGDEDEDEDDDDDEEEEDDDDD 853
Score = 34.7 bits (78), Expect = 0.053
Identities = 11/27 (40%), Positives = 23/27 (84%)
Frame = +2
Query: 233 DDMDEDEEEEEEEDEEDDEVESESEDE 259
+D++ED+E+ +E+++EDD+ + E ED+
Sbjct: 764 EDIEEDDEDGDEDEDEDDDDDEEEEDD 844
Score = 32.0 bits (71), Expect = 0.34
Identities = 9/28 (32%), Positives = 23/28 (82%)
Frame = +2
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
++D ++ +E+E+E+D++D+E E + +D+
Sbjct: 773 EEDDEDGDEDEDEDDDDDEEEEDDDDDD 856
Score = 31.6 bits (70), Expect = 0.45
Identities = 11/28 (39%), Positives = 21/28 (74%)
Frame = +2
Query: 232 QDDMDEDEEEEEEEDEEDDEVESESEDE 259
Q D ++ EE++E+ DE++DE + + E+E
Sbjct: 752 QSDFEDIEEDDEDGDEDEDEDDDDDEEE 835
Score = 29.3 bits (64), Expect = 2.2
Identities = 31/112 (27%), Positives = 46/112 (40%), Gaps = 2/112 (1%)
Frame = +2
Query: 194 LPPKPALSEHSTSNPTSSTDHIPTIMYTMADPPSTNPNQDDMDEDEEEE-EEEDEEDDEV 252
L KP +T P + T+ + + +PP + DD+D+D EE + E D ++
Sbjct: 515 LKKKPKKGSKNTK-PITKTEKCESF-FNFFNPPQVPEDDDDIDDDAVEELQNLMEHDYDI 688
Query: 253 ESESEDETGGRIRRATS-MTAIVEPSELMQIEMPDDIRIGSPNDGSNHLDSD 303
S D+ I A S T E S+ IE D+ DG D D
Sbjct: 689 GSTIRDKI---IPHAVSWFTGEAEQSDFEDIEEDDE-------DGDEDEDED 814
Score = 27.7 bits (60), Expect = 6.5
Identities = 10/27 (37%), Positives = 19/27 (70%)
Frame = +2
Query: 233 DDMDEDEEEEEEEDEEDDEVESESEDE 259
+ D ++ EE++ED ++DE E + +DE
Sbjct: 749 EQSDFEDIEEDDEDGDEDEDEDDDDDE 829
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.131 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,305,058
Number of Sequences: 28460
Number of extensions: 189103
Number of successful extensions: 3110
Number of sequences better than 10.0: 223
Number of HSP's better than 10.0 without gapping: 1753
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2423
length of query: 701
length of database: 4,897,600
effective HSP length: 97
effective length of query: 604
effective length of database: 2,136,980
effective search space: 1290735920
effective search space used: 1290735920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC135317.3


