

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
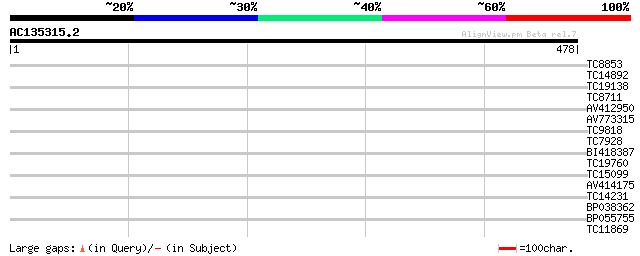
Query= AC135315.2 - phase: 0
(478 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8853 similar to PIR|T09217|T09217 protein sam2B - spinach {Spi... 30 0.86
TC14892 similar to UP|Q9Z0W6 (Q9Z0W6) Pax transcription activati... 29 1.9
TC19138 weakly similar to UP|Q9LZJ3 (Q9LZJ3) Alpha galactosyltra... 29 1.9
TC8711 weakly similar to UP|VCLC_PEA (P13918) Vicilin precursor,... 28 3.3
AV412950 28 3.3
AV773315 27 5.6
TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, ... 27 5.6
TC7928 similar to UP|Q93YR3 (Q93YR3) HSP associated protein like... 27 5.6
BI418387 27 5.6
TC19760 weakly similar to UP|CPR2_PETCR (Q99090) Light-inducible... 27 7.3
TC15099 weakly similar to UP|Q8LQG1 (Q8LQG1) Vacuolar sorting re... 27 7.3
AV414175 27 7.3
TC14231 27 7.3
BP038362 27 7.3
BP055755 27 9.6
TC11869 27 9.6
>TC8853 similar to PIR|T09217|T09217 protein sam2B - spinach {Spinacia
oleracea;}, partial (43%)
Length = 1157
Score = 30.0 bits (66), Expect = 0.86
Identities = 16/45 (35%), Positives = 28/45 (61%), Gaps = 3/45 (6%)
Frame = +3
Query: 412 DGKDEQKTHENLMLKKELVKVRRELEEK---DELLMRDSKRARGR 453
+GK+E + + + +KKELVKV +E E+ +L+ ++A GR
Sbjct: 381 EGKEETEEEKRMRVKKELVKVAKEQAERRATAQLMFDLGQKAYGR 515
>TC14892 similar to UP|Q9Z0W6 (Q9Z0W6) Pax transcription activation domain
interacting protein PTIP, partial (3%)
Length = 1502
Score = 28.9 bits (63), Expect = 1.9
Identities = 13/29 (44%), Positives = 17/29 (57%)
Frame = -1
Query: 411 LDGKDEQKTHENLMLKKELVKVRRELEEK 439
L KDEQ TH+NL K + + RE+ K
Sbjct: 1193 LSPKDEQTTHQNLNFSKHVSHLAREITTK 1107
>TC19138 weakly similar to UP|Q9LZJ3 (Q9LZJ3) Alpha
galactosyltransferase-like protein
(AT3g62720/F26K9_150), partial (12%)
Length = 458
Score = 28.9 bits (63), Expect = 1.9
Identities = 20/68 (29%), Positives = 30/68 (43%), Gaps = 8/68 (11%)
Frame = +1
Query: 341 EAYTQWVINRAEE--------IGMPYPAMRHVAASAPSIPLPLPPTTQEMYQEHLAMESR 392
E +T+ RAE+ I P PA +A P +PLPLP ++ H + R
Sbjct: 169 ERWTEVWCGRAEQQNAGTVLRISPPPPAAARLAPRLPHLPLPLP------HRHHASRHHR 330
Query: 393 KKQMWKAR 400
+Q+ R
Sbjct: 331 SRQVRNPR 354
>TC8711 weakly similar to UP|VCLC_PEA (P13918) Vicilin precursor, partial
(74%)
Length = 1591
Score = 28.1 bits (61), Expect = 3.3
Identities = 12/26 (46%), Positives = 16/26 (61%), Gaps = 1/26 (3%)
Frame = +3
Query: 196 RTLKG-RGYILCCVPLLYRWFISHLP 220
R LK RG++ CC + WF+S LP
Sbjct: 42 RVLKSKRGFLCCCCWEFFSWFLSPLP 119
>AV412950
Length = 431
Score = 28.1 bits (61), Expect = 3.3
Identities = 15/46 (32%), Positives = 25/46 (53%)
Frame = -1
Query: 98 SLTPLVIAKDLYLKTSDVSKHLITKSHIRGFTSKYLLVQANLETTC 143
SL+ + IA DL +DVS ++ K + F S + ++ +L TC
Sbjct: 410 SLSFMNIASDLLCHLTDVSFRILAKRDMIHFHSSRIFIRNHLNITC 273
>AV773315
Length = 519
Score = 27.3 bits (59), Expect = 5.6
Identities = 15/45 (33%), Positives = 21/45 (46%)
Frame = -2
Query: 425 LKKELVKVRRELEEKDELLMRDSKRARGRRNFYARYCGSDSESES 469
++ ++K+ L K R + RGR ARYC ESES
Sbjct: 452 IRSRMLKLSLMLPSKLFFYHRSRSKTRGRHRRLARYCDYG*ESES 318
>TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(62%)
Length = 1162
Score = 27.3 bits (59), Expect = 5.6
Identities = 14/36 (38%), Positives = 20/36 (54%), Gaps = 4/36 (11%)
Frame = +1
Query: 310 PTGQREAFTRAW----SKVRRKSVRHLGVRSGIAHE 341
P + E R+W S RR+SV H GV + ++HE
Sbjct: 697 PQSRMENDARSWRTPISNPRRRSVSHAGVPTVVSHE 804
>TC7928 similar to UP|Q93YR3 (Q93YR3) HSP associated protein like, partial
(52%)
Length = 765
Score = 27.3 bits (59), Expect = 5.6
Identities = 20/52 (38%), Positives = 27/52 (51%), Gaps = 3/52 (5%)
Frame = +3
Query: 366 ASAPSIPLPLPP---TTQEMYQEHLAMESRKKQMWKARYNEAENLIMTLDGK 414
ASAPS PLPL P T + M + M S+ + + + NLI +L GK
Sbjct: 117 ASAPSFPLPLTPIRSTRRAMMI*KIFMRSK*RWKNQRKMMR*LNLISSLKGK 272
>BI418387
Length = 479
Score = 27.3 bits (59), Expect = 5.6
Identities = 11/28 (39%), Positives = 14/28 (49%)
Frame = +3
Query: 359 PAMRHVAASAPSIPLPLPPTTQEMYQEH 386
P H +S +PLPLP T M + H
Sbjct: 63 PLQPHTCSSHTLLPLPLPSTLHSMVETH 146
>TC19760 weakly similar to UP|CPR2_PETCR (Q99090) Light-inducible protein
CPRF-2, partial (21%)
Length = 573
Score = 26.9 bits (58), Expect = 7.3
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = -1
Query: 82 LGSPIAEKTPFTGPGASLTPLV 103
LG P + PFTG GA+ TP++
Sbjct: 321 LGGPSDKTPPFTGVGAAATPVL 256
>TC15099 weakly similar to UP|Q8LQG1 (Q8LQG1) Vacuolar sorting receptor-like
protein, partial (16%)
Length = 687
Score = 26.9 bits (58), Expect = 7.3
Identities = 11/29 (37%), Positives = 16/29 (54%)
Frame = -2
Query: 355 GMPYPAMRHVAASAPSIPLPLPPTTQEMY 383
G P+P++R+ A + P P P T Q Y
Sbjct: 209 GCPFPSIRYSAKYVKTRPTP*PSTVQRKY 123
>AV414175
Length = 226
Score = 26.9 bits (58), Expect = 7.3
Identities = 10/25 (40%), Positives = 12/25 (48%)
Frame = +3
Query: 206 CCVPLLYRWFISHLPSSFHDNSEDW 230
CC P W +H H NS+DW
Sbjct: 108 CCQPFWELWVWTHGRFHSHGNSQDW 182
>TC14231
Length = 803
Score = 26.9 bits (58), Expect = 7.3
Identities = 18/67 (26%), Positives = 31/67 (45%), Gaps = 4/67 (5%)
Frame = -2
Query: 241 PNEVVWITPAAQV----KEIITGCGDFLNVPLLGTRGGINYNPELAMRQFGFPMKTKPIN 296
PN +VW PAA K I +G G+ ++V L+ + + +L + F + +
Sbjct: 328 PNPIVWALPAACYL*CHKTISSGVGNEVHVHLISISSNKHISTKLLVESFDHINELWIVQ 149
Query: 297 LATSPEF 303
+S EF
Sbjct: 148 RCSSXEF 128
>BP038362
Length = 274
Score = 26.9 bits (58), Expect = 7.3
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = -3
Query: 69 FQLVPTLEAYSYLLGSPIAEKTPFTGPGASLTPLVI 104
F PT+EA+SYL ++ P LTPL++
Sbjct: 209 FPFSPTVEAFSYLGHGTSPWESSHPSPTLDLTPLIL 102
>BP055755
Length = 273
Score = 26.6 bits (57), Expect = 9.6
Identities = 11/31 (35%), Positives = 18/31 (57%)
Frame = -3
Query: 371 IPLPLPPTTQEMYQEHLAMESRKKQMWKARY 401
IP P P+T +QE A +R ++W+ +Y
Sbjct: 238 IPPPYDPSTFPSFQEGKAYATRPVEIWEDKY 146
>TC11869
Length = 753
Score = 26.6 bits (57), Expect = 9.6
Identities = 13/43 (30%), Positives = 21/43 (48%)
Frame = +1
Query: 153 LLIYGLILFPNLDNFVDMNAIEIFHSRNPVPTLLANMYHAIHD 195
LL+ G + FV +++ H +PTLLA+ A+ D
Sbjct: 7 LLLPGQSVLVETSGFVIQQTLQVAHMVTVIPTLLASRVAAVSD 135
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,229,804
Number of Sequences: 28460
Number of extensions: 116177
Number of successful extensions: 690
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 686
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 690
length of query: 478
length of database: 4,897,600
effective HSP length: 94
effective length of query: 384
effective length of database: 2,222,360
effective search space: 853386240
effective search space used: 853386240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC135315.2


