

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135311.8 + phase: 0
(189 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
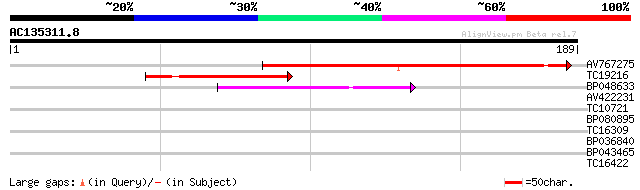
Sequences producing significant alignments: (bits) Value
AV767275 114 1e-26
TC19216 similar to UP|MCPI_SOLTU (P01075) Metallocarboxypeptidas... 49 5e-07
BP048633 49 5e-07
AV422231 28 0.99
TC10721 similar to UP|MTD_MEDSA (O82515) Probable mannitol dehyd... 27 2.2
BP080895 27 2.9
TC16309 similar to UP|AXI7_ARATH (Q38825) Auxin-responsive prote... 26 4.9
BP036840 26 4.9
BP043465 25 6.4
TC16422 similar to PIR|T05653|T05653 amino acid transport protei... 25 6.4
>AV767275
Length = 485
Score = 114 bits (285), Expect = 1e-26
Identities = 56/104 (53%), Positives = 70/104 (66%), Gaps = 1/104 (0%)
Frame = -3
Query: 85 KGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITISA-NGRNVVAMVVDECDS 143
+GGDGG S+CD+++H VVALSTGW+N SRC I I+A NG++ A VVDECDS
Sbjct: 483 EGGDGGAESKCDEKFHDSSERVVALSTGWYNKGSRCGKMIRITAMNGKSTTAKVVDECDS 304
Query: 144 RKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLDITWS 187
GCD +H PPC NIVD S+AVW ALG+ + G +TWS
Sbjct: 303 VNGCDSEHAGLPPCRKNIVDGSQAVWNALGLDGDV-GEEHVTWS 175
>TC19216 similar to UP|MCPI_SOLTU (P01075) Metallocarboxypeptidase inhibitor
(MCPI) (Carboxypeptidase inhibitor), partial (33%)
Length = 406
Score = 48.9 bits (115), Expect = 5e-07
Identities = 23/49 (46%), Positives = 30/49 (60%)
Frame = +2
Query: 46 CNQENDSDCCVQGKMYTTYVCSPSVSTHTKAYLTLNSFQKGGDGGGPSE 94
C + +C +GK Y Y CSP V + T+A LTLN F +GGDGG S+
Sbjct: 266 CKSSGNLNC--KGKSYPQYRCSPPVLSSTQALLTLNDFSEGGDGGAESK 406
>BP048633
Length = 380
Score = 48.9 bits (115), Expect = 5e-07
Identities = 28/66 (42%), Positives = 36/66 (54%)
Frame = +1
Query: 70 VSTHTKAYLTLNSFQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITISAN 129
V++HT+ L K GGGPS+ D YH +D P VAL T ++ C NI IS N
Sbjct: 43 VTSHTRGISHLEQLGKKSRGGGPSKGDN*YHPEDNPGVALGTEG-TPQAGCHQNIAISTN 219
Query: 130 GRNVVA 135
R+V A
Sbjct: 220 SRSVEA 237
>AV422231
Length = 466
Score = 28.1 bits (61), Expect = 0.99
Identities = 22/75 (29%), Positives = 34/75 (45%)
Frame = +2
Query: 2 KRFCPKVSFIFLAITLIFTSFVYSEAQNCRPSGRIRGKKAPPGKCNQENDSDCCVQGKMY 61
K+F SF+ L L F S +E C P+ G+ PPG +E+ D C++ +
Sbjct: 74 KQFRFPSSFLDL---LRFRSHRQNERHRCLPANPADGQVHPPGS-RRESQRDLCLRRRRI 241
Query: 62 TTYVCSPSVSTHTKA 76
CS ST ++
Sbjct: 242 QHRSCSWLKSTRRRS 286
>TC10721 similar to UP|MTD_MEDSA (O82515) Probable mannitol dehydrogenase
(NAD-dependent mannitol dehydrogenase) , complete
Length = 1297
Score = 26.9 bits (58), Expect = 2.2
Identities = 15/41 (36%), Positives = 22/41 (53%), Gaps = 1/41 (2%)
Frame = +3
Query: 122 NNITISANGRNV-VAMVVDECDSRKGCDEQHDYQPPCPNNI 161
NN+T G V V ++VD C S + CD+ D + CP +
Sbjct: 321 NNVTKFKVGDKVGVGVIVDSCKSCENCDQ--DLENYCPQPV 437
>BP080895
Length = 404
Score = 26.6 bits (57), Expect = 2.9
Identities = 18/56 (32%), Positives = 26/56 (46%)
Frame = -1
Query: 36 IRGKKAPPGKCNQENDSDCCVQGKMYTTYVCSPSVSTHTKAYLTLNSFQKGGDGGG 91
+R + PP C + S G ++ +CS +V+THT A S G GGG
Sbjct: 395 MRSPRLPPRTCLPLHGSR---SGVLFADPICSSTVATHTLASFLFLS----GCGGG 249
>TC16309 similar to UP|AXI7_ARATH (Q38825) Auxin-responsive protein IAA7
(Indoleacetic acid-induced protein 7), partial (63%)
Length = 812
Score = 25.8 bits (55), Expect = 4.9
Identities = 20/77 (25%), Positives = 28/77 (35%), Gaps = 9/77 (11%)
Frame = -1
Query: 38 GKKAPPGKCNQENDSDCCVQGKMYTTYVCSPSVSTHTKAYLTLNSFQ---------KGGD 88
GK P K Q CC+ CSPSV + + ++L + Q +G
Sbjct: 626 GKVLHPWKL*QMPHQQCCLLSSQCLDPHCSPSVPSCSSCMISLVANQLLEP*LVAWQGP* 447
Query: 89 GGGPSECDKQYHSDDTP 105
GG +H D P
Sbjct: 446 GGSSPLKHSSHHHSDLP 396
>BP036840
Length = 584
Score = 25.8 bits (55), Expect = 4.9
Identities = 14/32 (43%), Positives = 17/32 (52%), Gaps = 6/32 (18%)
Frame = +3
Query: 155 PPCPNNIVDASKAVWKAL------GVPKEQWG 180
PPCP++ V AS AV GV K +WG
Sbjct: 21 PPCPDDNVTASAAVRGGTRHPVYRGVRKRRWG 116
>BP043465
Length = 559
Score = 25.4 bits (54), Expect = 6.4
Identities = 9/29 (31%), Positives = 15/29 (51%)
Frame = -3
Query: 153 YQPPCPNNIVDASKAVWKALGVPKEQWGG 181
+ PPCP N WK+ G+ +++ G
Sbjct: 320 FSPPCPPNASSLRSDEWKSEGLEEKRSSG 234
>TC16422 similar to PIR|T05653|T05653 amino acid transport protein homolog
F22I13.20 - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (38%)
Length = 685
Score = 25.4 bits (54), Expect = 6.4
Identities = 17/54 (31%), Positives = 26/54 (47%)
Frame = +2
Query: 83 FQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITISANGRNVVAM 136
FQK + G S Y +DTP+++ S + K + NI IS G V+ +
Sbjct: 89 FQKEPEAGSSSYSLPPYPREDTPLLSKSPP-LSSKFKTFANIFISIVGAGVLGL 247
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.134 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,410,503
Number of Sequences: 28460
Number of extensions: 65959
Number of successful extensions: 304
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 302
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 303
length of query: 189
length of database: 4,897,600
effective HSP length: 85
effective length of query: 104
effective length of database: 2,478,500
effective search space: 257764000
effective search space used: 257764000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC135311.8


