

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135234.5 - phase: 0 /pseudo
(256 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
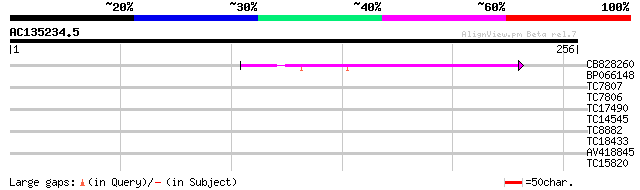
Score E
Sequences producing significant alignments: (bits) Value
CB828260 91 1e-19
BP066148 32 0.082
TC7807 homologue to UP|Q946J9 (Q946J9) Aquaporin protein PIP1,1,... 28 2.0
TC7806 homologue to UP|Q946J9 (Q946J9) Aquaporin protein PIP1,1,... 28 2.0
TC17490 similar to GB|AAO64817.1|29028876|BT005882 At3g04570 {Ar... 27 3.4
TC14545 similar to UP|Q9FT78 (Q9FT78) P23 co-chaperone, partial ... 27 3.4
TC8882 homologue to UP|Q43715 (Q43715) Chloroplastic outer envel... 26 5.9
TC18433 26 5.9
AV418845 26 5.9
TC15820 similar to UP|Q9SWF9 (Q9SWF9) Zinc finger protein, parti... 26 7.7
>CB828260
Length = 557
Score = 91.3 bits (225), Expect = 1e-19
Identities = 54/131 (41%), Positives = 75/131 (57%), Gaps = 3/131 (2%)
Frame = +3
Query: 105 KRPYLTTQSERSGIYKPPKTRKKPNL-EGIGSMKCNCPFRLKGFFDKD--TNDWWLAILC 161
++ Y+ E+ G Y P + P+L EG + K CPFRLKG K+ DW L ++
Sbjct: 174 RKAYVMLGCEKHGKYVP---YRDPDLVEGTRTQKTECPFRLKGRPMKNGTDRDWRLKVME 344
Query: 162 GMHNHDLEEKLQGHLIAGRLSAEEKKKVIDMTKSLTVPRNILTNLKENNKESVTTIKQVY 221
G HNH+ L GH GRL++EEK++V M+KS +PR +L LKENN ++TTI Q+Y
Sbjct: 345 GAHNHEPARSLLGHNFVGRLNSEEKEQVEKMSKSWVLPRKMLLTLKENNPLNLTTISQIY 524
Query: 222 NVRTRWCKGER 232
R K R
Sbjct: 525 GACKRLRKSLR 557
>BP066148
Length = 477
Score = 32.3 bits (72), Expect = 0.082
Identities = 13/23 (56%), Positives = 17/23 (73%)
Frame = +1
Query: 199 PRNILTNLKENNKESVTTIKQVY 221
PR +L LKENN ++TTI Q+Y
Sbjct: 88 PRKMLLTLKENNPSNLTTISQIY 156
>TC7807 homologue to UP|Q946J9 (Q946J9) Aquaporin protein PIP1,1, partial
(27%)
Length = 1289
Score = 27.7 bits (60), Expect = 2.0
Identities = 10/17 (58%), Positives = 11/17 (63%)
Frame = -2
Query: 71 TTNARWREREELLTWVR 87
TTN RWR R + WVR
Sbjct: 250 TTNHRWRRRSRMCPWVR 200
>TC7806 homologue to UP|Q946J9 (Q946J9) Aquaporin protein PIP1,1,
complete
Length = 1413
Score = 27.7 bits (60), Expect = 2.0
Identities = 10/17 (58%), Positives = 11/17 (63%)
Frame = +1
Query: 71 TTNARWREREELLTWVR 87
TTN RWR R + WVR
Sbjct: 55 TTNHRWRRRSRMCPWVR 105
>TC17490 similar to GB|AAO64817.1|29028876|BT005882 At3g04570 {Arabidopsis
thaliana;}, partial (9%)
Length = 604
Score = 26.9 bits (58), Expect = 3.4
Identities = 13/23 (56%), Positives = 13/23 (56%)
Frame = +1
Query: 7 DDDVEPKPAKTEASGEGSGDVKP 29
DDD P A T ASG GSG P
Sbjct: 67 DDDEGPSGAATAASGGGSGSSPP 135
>TC14545 similar to UP|Q9FT78 (Q9FT78) P23 co-chaperone, partial (46%)
Length = 1127
Score = 26.9 bits (58), Expect = 3.4
Identities = 16/46 (34%), Positives = 22/46 (47%), Gaps = 5/46 (10%)
Frame = +1
Query: 64 RDTASFFTTNARWRER-----EELLTWVRRQGVKAGFGVCIDKSVL 104
R + SF T + R + E W RRQ V G G C++K+ L
Sbjct: 262 RSSLSFLTRSM*RRAKLILGCEAYSLWCRRQRVDGGKGCCVEKASL 399
>TC8882 homologue to UP|Q43715 (Q43715) Chloroplastic outer envelope
membrane protein (OEP75) precursor (OEP75), partial
(32%)
Length = 1246
Score = 26.2 bits (56), Expect = 5.9
Identities = 19/68 (27%), Positives = 31/68 (44%), Gaps = 1/68 (1%)
Frame = -2
Query: 150 KDTNDWWLAILCGMHNHDLEEKLQGHLIAGRLSAEEKKKVIDMTKSLTVPRNI-LTNLKE 208
+D + WW +H H LEE Q L+ E+ ++ TK+L +I ++N
Sbjct: 420 EDKHRWWWFTSPFLHRHQLEEACQSKLV-----TIEEWELAANTKALIYLEHISISNYSS 256
Query: 209 NNKESVTT 216
+K V T
Sbjct: 255 IHKTCVIT 232
>TC18433
Length = 428
Score = 26.2 bits (56), Expect = 5.9
Identities = 17/46 (36%), Positives = 23/46 (49%)
Frame = -2
Query: 53 KRVPIGVVCEARDTASFFTTNARWREREELLTWVRRQGVKAGFGVC 98
KRVP GVV + D + E+EE W++R+G G VC
Sbjct: 139 KRVPCGVVVTSGD-------DG*CEEKEEPW*WLQREGEMDGECVC 23
>AV418845
Length = 293
Score = 26.2 bits (56), Expect = 5.9
Identities = 13/28 (46%), Positives = 15/28 (53%)
Frame = -2
Query: 2 VDNVADDDVEPKPAKTEASGEGSGDVKP 29
V A D VEP A T + GEG G +P
Sbjct: 127 VSESAIDVVEPCQASTNSGGEGGGGRRP 44
>TC15820 similar to UP|Q9SWF9 (Q9SWF9) Zinc finger protein, partial (18%)
Length = 663
Score = 25.8 bits (55), Expect = 7.7
Identities = 11/20 (55%), Positives = 13/20 (65%)
Frame = -1
Query: 8 DDVEPKPAKTEASGEGSGDV 27
DD EPK + EAS EG D+
Sbjct: 237 DDDEPKRRRPEASAEGGADM 178
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,506,550
Number of Sequences: 28460
Number of extensions: 59816
Number of successful extensions: 319
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 318
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 318
length of query: 256
length of database: 4,897,600
effective HSP length: 88
effective length of query: 168
effective length of database: 2,393,120
effective search space: 402044160
effective search space used: 402044160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC135234.5


