

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135231.7 + phase: 0 /pseudo
(943 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
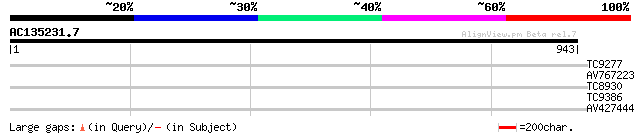
Score E
Sequences producing significant alignments: (bits) Value
TC9277 similar to GB|AAP13413.1|30023760|BT006305 At3g55530 {Ara... 30 2.3
AV767223 30 2.3
TC8930 UP|Q40204 (Q40204) RAB1D, complete 28 5.2
TC9386 28 6.8
AV427444 28 8.8
>TC9277 similar to GB|AAP13413.1|30023760|BT006305 At3g55530 {Arabidopsis
thaliana;}, partial (79%)
Length = 1297
Score = 29.6 bits (65), Expect = 2.3
Identities = 13/21 (61%), Positives = 13/21 (61%)
Frame = +2
Query: 322 GHR*IFGLTFYDRSRSKCYFC 342
G R* F TFYDR R KC C
Sbjct: 686 GFR*CFYCTFYDRGRDKCSSC 748
>AV767223
Length = 256
Score = 29.6 bits (65), Expect = 2.3
Identities = 13/26 (50%), Positives = 18/26 (69%)
Frame = -3
Query: 328 GLTFYDRSRSKCYFCLHQGSCLAENQ 353
G +R+ S+CYFCL GSCL+ +Q
Sbjct: 239 GFAGVERAYSECYFCL--GSCLSNDQ 168
>TC8930 UP|Q40204 (Q40204) RAB1D, complete
Length = 943
Score = 28.5 bits (62), Expect = 5.2
Identities = 14/34 (41%), Positives = 18/34 (52%), Gaps = 2/34 (5%)
Frame = -3
Query: 457 SGVSNF*SSLLPLWLLSHGHCWAQPQLC--LAQH 488
SG+ + LP WL S HCW +C LA+H
Sbjct: 707 SGLGRAAPTFLPNWLSSDLHCWRPSIICAGLARH 606
>TC9386
Length = 751
Score = 28.1 bits (61), Expect = 6.8
Identities = 17/47 (36%), Positives = 24/47 (50%)
Frame = +2
Query: 667 YACFLCLSFFYTGLE*NRSFGVRCNTPFLPLPQLRKLYFICWRLCRL 713
Y LCLS F TGLE R F + +L L ++ L ++ + L L
Sbjct: 83 YFSILCLSSFVTGLEPLRFFFSTSQSSYLFLQVIQTLAYVLYLLLLL 223
>AV427444
Length = 428
Score = 27.7 bits (60), Expect = 8.8
Identities = 14/40 (35%), Positives = 22/40 (55%)
Frame = -3
Query: 778 PLNNIESRGSLPWPEDTNVTSMRPSLLTSTVQVLVFVFAM 817
P+NN E+ P P N++ M+PSL + L F+F +
Sbjct: 405 PINNFETTTR*PCP---NISCMQPSLFINDFSRLSFIFVI 295
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.365 0.165 0.642
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,647,729
Number of Sequences: 28460
Number of extensions: 325637
Number of successful extensions: 4623
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 2558
Number of HSP's successfully gapped in prelim test: 172
Number of HSP's that attempted gapping in prelim test: 1980
Number of HSP's gapped (non-prelim): 2916
length of query: 943
length of database: 4,897,600
effective HSP length: 99
effective length of query: 844
effective length of database: 2,080,060
effective search space: 1755570640
effective search space used: 1755570640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC135231.7


