

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
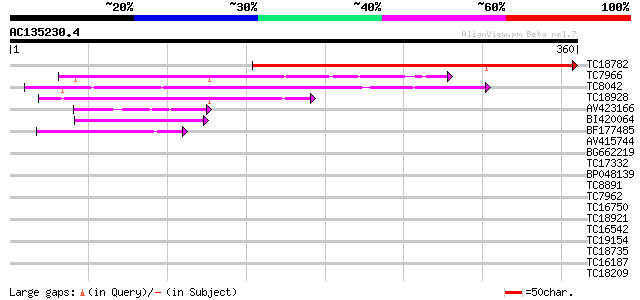
Query= AC135230.4 - phase: 0
(360 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18782 similar to UP|Q8LCU7 (Q8LCU7) Nuclear receptor binding f... 367 e-102
TC7966 similar to UP|Q84V25 (Q84V25) Quinone oxidoreductase, par... 74 3e-14
TC8042 homologue to UP|P93619 (P93619) Probable NADP-dependent o... 74 5e-14
TC18928 similar to UP|Q43677 (Q43677) Auxin-induced protein, par... 60 4e-10
AV423166 49 2e-06
BI420064 49 2e-06
BF177485 42 1e-04
AV415744 37 0.007
BG662219 36 0.009
TC17332 similar to UP|MTD_FRAAN (Q9ZRF1) Probable mannitol dehyd... 32 0.22
BP048139 32 0.22
TC8891 similar to GB|AAP68279.1|31711846|BT008840 At1g72680 {Ara... 32 0.22
TC7962 homologue to UP|Q84V97 (Q84V97) Alcohol dehydrogenase 1 ... 31 0.28
TC16750 similar to UP|CADH_MEDSA (P31656) Cinnamyl-alcohol dehyd... 30 0.48
TC18921 homologue to PIR|JQ1073|JQ1073 tryptophan synthase beta... 30 0.63
TC16542 similar to UP|Q93X81 (Q93X81) Sorbitol dehydrogenase, pa... 29 1.4
TC19154 similar to UP|AAS38575 (AAS38575) Short-chain dehydrogen... 28 1.8
TC18735 similar to UP|DSX_DROME (P23023) Doublesex protein, part... 28 1.8
TC16187 similar to UP|Q93122 (Q93122) AsSLR8.97, partial (19%) 28 2.4
TC18209 similar to UP|TRPD_ARATH (Q02166) Anthranilate phosphori... 28 3.1
>TC18782 similar to UP|Q8LCU7 (Q8LCU7) Nuclear receptor binding factor-like
protein, partial (57%)
Length = 832
Score = 367 bits (942), Expect = e-102
Identities = 189/216 (87%), Positives = 197/216 (90%), Gaps = 10/216 (4%)
Frame = +3
Query: 155 MEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRD 214
+EYAATITVNPLTA LMLE C++LNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRD
Sbjct: 3 LEYAATITVNPLTAFLMLEHCISLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRD 182
Query: 215 RPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLR 274
RPGV EVKERLKDLGADEVFTE ELEVKNVKSLLGGIPEPALGFNCVGGN+ASLVLKFLR
Sbjct: 183 RPGVDEVKERLKDLGADEVFTEKELEVKNVKSLLGGIPEPALGFNCVGGNAASLVLKFLR 362
Query: 275 RGGTMVTYGGMSKKPVTVSTSSFIFKN----------WLSTDKAEEGRRMIDRLLGLVQD 324
+GGTMVTYGGMS KPVTVSTSSFIFK+ WLST+KAEE R MID+LLGLVQD
Sbjct: 363 QGGTMVTYGGMSLKPVTVSTSSFIFKDLSLRGFWLQKWLSTEKAEESRGMIDKLLGLVQD 542
Query: 325 GKLKYKMELTPFNDFNTALDKALGKLGSQPKQVIKF 360
GKLKYKMELTPFNDFNTALDKALGKLGS PK VIKF
Sbjct: 543 GKLKYKMELTPFNDFNTALDKALGKLGSPPKPVIKF 650
>TC7966 similar to UP|Q84V25 (Q84V25) Quinone oxidoreductase, partial (98%)
Length = 1908
Score = 74.3 bits (181), Expect = 3e-14
Identities = 74/262 (28%), Positives = 119/262 (45%), Gaps = 12/262 (4%)
Frame = +2
Query: 32 FSSAVSPPS--KAIIYESHGQPDAVTKL-VDIPATEVKENDLCVKMLAAPINPSDINRIQ 88
FSS + P+ KA Y +G+ + K ++P E+KE+ + +K++AA +NP D R++
Sbjct: 704 FSSMAAIPTHIKAWTYSEYGKSTDILKFDPNVPLPELKEDHVLIKVVAAALNPIDYKRME 883
Query: 89 GVYPVRPEP-PAVGGYEGVGEVHSVGSAVTCFSPGDWV--------IPSPPSFGTWQTYI 139
++ P P GY+ G V VGS V F GD V + P G+ Y
Sbjct: 884 ALFKDTDSPLPTAPGYDVAGVVVKVGSEVKKFKVGDEVYGDINENALEYPKVVGSLAEYT 1063
Query: 140 VKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQ 199
++ + K + AA++ + TA LE ++G +I+ G VG VIQ
Sbjct: 1064AVEEKLLAHKPKNLSFVEAASLPLAIETAYEGLER-AGFSAGKSILVLGGAGGVGTHVIQ 1240
Query: 200 LAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFN 259
LAK + + + G++ E L+ LGAD ++ V+N+ + F+
Sbjct: 1241LAKH--VFGASKVAATSSTGKL-ELLRKLGADMPIDYTKENVENLPEKFDVV------FD 1393
Query: 260 CVGGNSASLVLKFLRRGGTMVT 281
VG A K L+ GG VT
Sbjct: 1394AVG--DADTTFKPLKEGGKFVT 1453
>TC8042 homologue to UP|P93619 (P93619) Probable NADP-dependent
oxidoreductase TED2 , complete
Length = 1244
Score = 73.6 bits (179), Expect = 5e-14
Identities = 78/300 (26%), Positives = 125/300 (41%), Gaps = 4/300 (1%)
Frame = +2
Query: 10 SPSSLFNFTSLRSRILNTHTHAF---SSAVSPPSKAIIYESHGQPDAVTKLVDIPATEVK 66
+PSS + +LR+R + A SS+ + KAI G P + K D+ + K
Sbjct: 20 TPSSSSSLITLRARATSLTAKASLSSSSSATEMVKAIRVHELGGPQ-ILKWEDVEIGDPK 196
Query: 67 ENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVI 126
EN++ VK A +N D+ +GVY P G E VG V +VG T GD V
Sbjct: 197 ENEVRVKNKAIGVNFIDVYFRKGVYKASTS-PFTPGMEAVGVVTAVGGGPTGVQVGDLVG 373
Query: 127 PSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQ 186
+ G++ + + + AA++ + +TA +L C + G I+
Sbjct: 374 YAGQPIGSYVEEQILHAGKVVPIPSSIDPSVAASVMLKGMTAQFLLRRCFKVERGHTILV 553
Query: 187 NGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELE-VKNVK 245
+ A VG + Q A + G I + + + KE G V E + V V
Sbjct: 554 HAAAGGVGSLLCQWANALGATVIGTVSNEEKAAQAKED----GCHHVIIYKEEDFVARVN 721
Query: 246 SLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKNWLST 305
+ G + ++ VG ++ L L++ G MV++G S P VS S K+ T
Sbjct: 722 EITSGNGVEVV-YDSVGKDTFEGSLACLKQRGYMVSFGQSSGSPDPVSLSFLAAKSLFLT 898
>TC18928 similar to UP|Q43677 (Q43677) Auxin-induced protein, partial (53%)
Length = 612
Score = 60.5 bits (145), Expect = 4e-10
Identities = 50/186 (26%), Positives = 86/186 (45%), Gaps = 10/186 (5%)
Frame = +1
Query: 19 SLRSRILNTHTHAFSSAVSPPSKAIIYESHGQPDAVTKL-VDIPATEVKENDLCVKMLAA 77
+L ++ ++T + + + A+ KA Y HG+ V K ++P E+KE+ + +K+ AA
Sbjct: 28 TLETQFISTSSSSMA-AIPTHIKAWAYSEHGKSVDVLKFDPNLPLPELKEDQVLIKVAAA 204
Query: 78 PINPSDINRIQGVYPVRPEP-PAVGGYEGVGEVHSVGSAVTCFSPGDWV--------IPS 128
+NP D R++G + P P GY+ G V VGS V F GD V +
Sbjct: 205 SLNPIDYKRMEGAFKASDSPLPTAPGYDVAGVVVKVGSEVKKFKVGDEVYGDINEQAMVY 384
Query: 129 PPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNG 188
P G+ Y ++ + + + AA++ + TA LE ++G +I+ G
Sbjct: 385 PKVVGSLAEYTAAEEKLLSHKPQNLSFAEAASLPLTIETAYNGLE-LAGFSAGKSILVLG 561
Query: 189 ATSMVG 194
VG
Sbjct: 562 GAGGVG 579
>AV423166
Length = 452
Score = 48.5 bits (114), Expect = 2e-06
Identities = 41/90 (45%), Positives = 51/90 (56%), Gaps = 2/90 (2%)
Frame = +2
Query: 41 KAIIYESHGQPDAVTKL-VDIP-ATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPP 98
KA I G+P V ++ VD P ATEV+ VKML A I +DI I G +P P
Sbjct: 83 KAAICWGLGEPVTVEEIQVDPPKATEVR-----VKMLCASICHTDITSIHG-FPHGIFPL 244
Query: 99 AVGGYEGVGEVHSVGSAVTCFSPGDWVIPS 128
A+G +EGVG V SVG VT GD +IP+
Sbjct: 245 ALG-HEGVGIVESVGDQVTNLKEGDMIIPT 331
>BI420064
Length = 410
Score = 48.5 bits (114), Expect = 2e-06
Identities = 29/87 (33%), Positives = 47/87 (53%), Gaps = 2/87 (2%)
Frame = +3
Query: 42 AIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDIN-RIQGVYPVRPEP-PA 99
A+ Y S+G + K V++P ++++ +K+ AA INP D +I + P+ P P
Sbjct: 117 AVQYNSYGGGPSALKQVEVPIPAPSKDEVLIKLEAASINPFDWKVQIGKLRPLLPRKFPY 296
Query: 100 VGGYEGVGEVHSVGSAVTCFSPGDWVI 126
+ G + GEV +G V F PGD V+
Sbjct: 297 IPGTDIAGEVVEIGQGVKKFKPGDKVV 377
>BF177485
Length = 334
Score = 42.4 bits (98), Expect = 1e-04
Identities = 31/97 (31%), Positives = 44/97 (44%), Gaps = 1/97 (1%)
Frame = +1
Query: 18 TSLRSRIL-NTHTHAFSSAVSPPSKAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLA 76
TS+ I H S++ KA++ E G + + D+P E KE ++ VK
Sbjct: 10 TSINPHIAPRAHNIRAMSSLPKTMKAVLIEKTGGTEVLQYKTDVPVPEPKEGEVLVKNEF 189
Query: 77 APINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVG 113
IN D GVY P+ P + G EG G + SVG
Sbjct: 190 IGINYIDTYFRSGVYK-PPQFPYILGREGAGTIASVG 297
>AV415744
Length = 415
Score = 36.6 bits (83), Expect = 0.007
Identities = 35/132 (26%), Positives = 54/132 (40%), Gaps = 7/132 (5%)
Frame = +1
Query: 178 LNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVF--- 234
L SG ++ GA VG +Q+ K+ G I + R E + LK G D V
Sbjct: 43 LTSGQVLLVLGAAGGVGLAAVQIGKACGAIVIAVARG----AEKVQLLKSHGVDHVVDLT 210
Query: 235 ----TESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPV 290
TES E + L G + ++ VGG L+ L+ G ++ G S +
Sbjct: 211 NENVTESVKEFLKARKLKG----IDVLYDPVGGKLTKECLRLLKWGANILIIGFASGEIP 378
Query: 291 TVSTSSFIFKNW 302
+ + + KNW
Sbjct: 379 VIPANIALVKNW 414
>BG662219
Length = 366
Score = 36.2 bits (82), Expect = 0.009
Identities = 24/78 (30%), Positives = 36/78 (45%)
Frame = +2
Query: 50 QPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEV 109
+P+ + D+ + ++ VK+L + +D G P P + G+E G V
Sbjct: 104 EPNKPLTIEDVQVAPPQAGEVRVKILYTALCHTDAYTWSGKDP-EGLFPCILGHEAAGVV 280
Query: 110 HSVGSAVTCFSPGDWVIP 127
SVG VT PGD VIP
Sbjct: 281 ESVGEGVTEVQPGDHVIP 334
>TC17332 similar to UP|MTD_FRAAN (Q9ZRF1) Probable mannitol dehydrogenase
(NAD-dependent mannitol dehydrogenase) , partial (63%)
Length = 736
Score = 31.6 bits (70), Expect = 0.22
Identities = 25/67 (37%), Positives = 32/67 (47%), Gaps = 5/67 (7%)
Frame = +1
Query: 64 EVKENDLCVKMLAAPINPSDINRIQG-----VYPVRPEPPAVGGYEGVGEVHSVGSAVTC 118
E E D+ K+L I SD++ I+ VYP+ P G+E G V VGS V
Sbjct: 139 ETGEKDVTFKVLYCGICHSDLHMIKNEWGSSVYPLVP------GHEIAGVVTEVGSKVQK 300
Query: 119 FSPGDWV 125
F GD V
Sbjct: 301 FKVGDRV 321
>BP048139
Length = 567
Score = 31.6 bits (70), Expect = 0.22
Identities = 16/30 (53%), Positives = 19/30 (63%)
Frame = -2
Query: 98 PAVGGYEGVGEVHSVGSAVTCFSPGDWVIP 127
P + G+E VG V SVG VT + GD VIP
Sbjct: 428 PRILGHEAVGVVESVGKDVTEVTKGDVVIP 339
>TC8891 similar to GB|AAP68279.1|31711846|BT008840 At1g72680 {Arabidopsis
thaliana;} , complete
Length = 1636
Score = 31.6 bits (70), Expect = 0.22
Identities = 35/123 (28%), Positives = 51/123 (41%), Gaps = 5/123 (4%)
Frame = +1
Query: 8 KPSPSSLFNFTSLRSRILNTHTHAFSSAVSPPSKAIIYESHGQPDAVTKLVDIPATEVKE 67
+P P SL NF+S S + H H + S+ + + V V +
Sbjct: 25 RPLPFSLLNFSSSSSSL---HLHRKIMSSEGASEECLGWAATDTSGVLSPYKFSRRAVGD 195
Query: 68 NDLCVKMLAAPINPSDI----NRI-QGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPG 122
+D+ VK+ + +D+ N + VYP P G+E G V VGS V FS G
Sbjct: 196 DDVYVKISHCGVCYADVAWTRNTLGNSVYPCVP------GHEIAGIVTKVGSNVHGFSVG 357
Query: 123 DWV 125
D V
Sbjct: 358 DHV 366
>TC7962 homologue to UP|Q84V97 (Q84V97) Alcohol dehydrogenase 1 , complete
Length = 1489
Score = 31.2 bits (69), Expect = 0.28
Identities = 14/30 (46%), Positives = 17/30 (56%)
Frame = +1
Query: 98 PAVGGYEGVGEVHSVGSAVTCFSPGDWVIP 127
P + G+E G V SVG VT PGD +P
Sbjct: 304 PRIFGHEAGGIVESVGEGVTHLKPGDHALP 393
>TC16750 similar to UP|CADH_MEDSA (P31656) Cinnamyl-alcohol dehydrogenase
(CAD) , partial (34%)
Length = 442
Score = 30.4 bits (67), Expect = 0.48
Identities = 19/58 (32%), Positives = 34/58 (57%)
Frame = +1
Query: 68 NDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWV 125
+D+ +K+ + +D+++++ + P V G+E VGEV VGS VT F+ G+ V
Sbjct: 166 DDVYIKVHYCGVCHTDVHQVKNDLGMS-NYPMVPGHEVVGEVLEVGSDVTRFTVGEIV 336
>TC18921 homologue to PIR|JQ1073|JQ1073 tryptophan synthase beta-2 chain
precursor - Arabidopsis thaliana
{Arabidopsis thaliana;} , partial (26%)
Length = 563
Score = 30.0 bits (66), Expect = 0.63
Identities = 20/50 (40%), Positives = 26/50 (52%), Gaps = 4/50 (8%)
Frame = +2
Query: 207 HNINIIRDRPGVGEVKERLKDLGADEVFTESELEV----KNVKSLLGGIP 252
H+I+ D PGVG LKDLG E ++ ++ E K V L G IP
Sbjct: 83 HSISAGLDYPGVGPEHSFLKDLGRAEYYSITDEEALEAFKRVSRLEGIIP 232
>TC16542 similar to UP|Q93X81 (Q93X81) Sorbitol dehydrogenase, partial (60%)
Length = 724
Score = 28.9 bits (63), Expect = 1.4
Identities = 20/64 (31%), Positives = 33/64 (51%), Gaps = 2/64 (3%)
Frame = +1
Query: 68 NDLCVKMLAAPINPSDINRIQGVYPVR--PEPPAVGGYEGVGEVHSVGSAVTCFSPGDWV 125
+D+ V+M A I SD++ ++ + + P V G+E G + +VG+ V PGD V
Sbjct: 163 HDVRVRMKAVGICGSDVHYLKKLRCADFIVKEPMVIGHECAGIIEAVGAEVKSLVPGDRV 342
Query: 126 IPSP 129
P
Sbjct: 343 AIEP 354
>TC19154 similar to UP|AAS38575 (AAS38575) Short-chain dehydrogenase Tic32,
partial (44%)
Length = 508
Score = 28.5 bits (62), Expect = 1.8
Identities = 23/105 (21%), Positives = 48/105 (44%)
Frame = +2
Query: 143 QNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAK 202
+++W KG P ++++ T +T + +G V GA+S +G ++
Sbjct: 74 ESMWPFSKKG-PSGFSSSSTAEQVTQGID-------GTGLTAVVTGASSGIGTETTRVLA 229
Query: 203 SRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSL 247
RG+H I +R+ +VKE + EL++ +++S+
Sbjct: 230 KRGVHVIMGVRNTAAGKDVKETILKENPSAKVDAMELDLSSMESV 364
>TC18735 similar to UP|DSX_DROME (P23023) Doublesex protein, partial (4%)
Length = 784
Score = 28.5 bits (62), Expect = 1.8
Identities = 17/44 (38%), Positives = 21/44 (47%)
Frame = +2
Query: 87 IQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPP 130
+QG YP P PP G G G+ +S G A G + P PP
Sbjct: 386 VQGYYPYHPTPPYGG---GAGDGNSGGGA----GGGSFGTPPPP 496
>TC16187 similar to UP|Q93122 (Q93122) AsSLR8.97, partial (19%)
Length = 500
Score = 28.1 bits (61), Expect = 2.4
Identities = 17/41 (41%), Positives = 26/41 (62%), Gaps = 3/41 (7%)
Frame = +3
Query: 1 MLRSLSHKPSPSSLFNFT---SLRSRILNTHTHAFSSAVSP 38
+L L KPS SSLF+F+ +L I ++H H +S+V+P
Sbjct: 48 LLHLLLSKPSTSSLFHFSIDLNLPDSI-SSHQHVLTSSVAP 167
>TC18209 similar to UP|TRPD_ARATH (Q02166) Anthranilate
phosphoribosyltransferase, chloroplast precursor ,
partial (41%)
Length = 774
Score = 27.7 bits (60), Expect = 3.1
Identities = 17/55 (30%), Positives = 27/55 (48%)
Frame = +2
Query: 7 HKPSPSSLFNFTSLRSRILNTHTHAFSSAVSPPSKAIIYESHGQPDAVTKLVDIP 61
+K P + + + + S +L+TH F SPP + +SH P+ T L D P
Sbjct: 5 NKVKPENHYGYGT--SSLLHTHFFPFLLLHSPPRS--LSQSHPPPNPATSLSDFP 157
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.135 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,294,408
Number of Sequences: 28460
Number of extensions: 93071
Number of successful extensions: 495
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 492
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 492
length of query: 360
length of database: 4,897,600
effective HSP length: 91
effective length of query: 269
effective length of database: 2,307,740
effective search space: 620782060
effective search space used: 620782060
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC135230.4


