

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135230.3 + phase: 0 /pseudo
(605 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
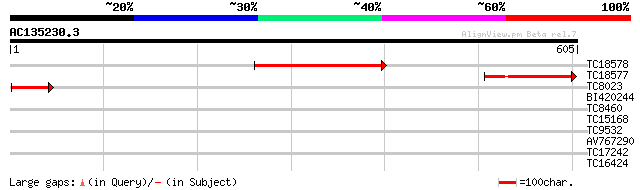
Score E
Sequences producing significant alignments: (bits) Value
TC18578 238 3e-63
TC18577 167 4e-42
TC8023 47 7e-06
BI420244 32 0.22
TC8460 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F... 32 0.29
TC15168 similar to UP|O04084 (O04084) Serine carboxypeptidase is... 31 0.65
TC9532 similar to UP|THIC_MYCLE (Q9ZBL0) Thiamine biosynthesis p... 28 5.5
AV767290 27 7.2
TC17242 27 9.4
TC16424 similar to UP|Q9FZF1 (Q9FZF1) T2E6.18, partial (39%) 27 9.4
>TC18578
Length = 600
Score = 238 bits (606), Expect = 3e-63
Identities = 117/141 (82%), Positives = 127/141 (89%)
Frame = +3
Query: 262 RPSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASEL 321
RPSFGIAFDIDGVILLGN+PVGGSP ALR+LY+ DG LK P+VFLTNGGG PEA RASEL
Sbjct: 144 RPSFGIAFDIDGVILLGNSPVGGSPGALRRLYDADGRLKIPFVFLTNGGGFPEATRASEL 323
Query: 322 SELLGLNVSASQVLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSEYGFKNVISIDAYAS 381
S+LLGL+V SQVLQGHSPF+QLVNRFE+KLIVA GKGEPA MSEYGFKNV+S+D YAS
Sbjct: 324 SQLLGLDVLPSQVLQGHSPFKQLVNRFENKLIVAVGKGEPAAAMSEYGFKNVLSMDEYAS 503
Query: 382 RFENIDPLAPYKKWTTKLATT 402
FENIDPLA YKKWTTKLATT
Sbjct: 504 CFENIDPLATYKKWTTKLATT 566
>TC18577
Length = 536
Score = 167 bits (423), Expect = 4e-42
Identities = 83/98 (84%), Positives = 89/98 (90%)
Frame = +2
Query: 507 PHPSVFKNAEIVLQKHVPRVDEDSYDINHKNAQPFQTLYMIGDNPAVDIRGARQTGHPWF 566
PHPSVFKNAE VLQ+ VP VD + D N+KNAQ FQTLYMIGDNPAVDIRGA+QTGHPWF
Sbjct: 2 PHPSVFKNAENVLQQFVPCVDLN--DFNNKNAQYFQTLYMIGDNPAVDIRGAQQTGHPWF 175
Query: 567 SILTRTGVFKGKENHDKFPADLVVDTVEEAVDYILAKE 604
SILTRTGVFKGKENHDKFPADLVVDTVEEAVD+IL +E
Sbjct: 176 SILTRTGVFKGKENHDKFPADLVVDTVEEAVDFILTRE 289
>TC8023
Length = 1089
Score = 47.4 bits (111), Expect = 7e-06
Identities = 23/45 (51%), Positives = 30/45 (66%), Gaps = 1/45 (2%)
Frame = +3
Query: 3 PFDIEAYFQKRDVEDTYIVNRFIQRRKKLEEGSASRT-RKYFNRD 46
PFD+EAY ++ ++ED YI N F +RR + EG R RKY NRD
Sbjct: 54 PFDVEAYKRQSEIEDRYIFNLFRKRRNLILEGYTPRVKRKYLNRD 188
>BI420244
Length = 571
Score = 32.3 bits (72), Expect = 0.22
Identities = 29/97 (29%), Positives = 45/97 (45%), Gaps = 8/97 (8%)
Frame = -1
Query: 259 LLIRPSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPY--------VFLTNGG 310
+L R F + DIDG LL G +PAA G K V G
Sbjct: 412 ILYRKIFRLGKDIDGDDLLSGVTGGATPAA-------SGPNKAVRRRVV*ALEVSSRPSG 254
Query: 311 GIPEAKRASELSELLGLNVSASQVLQGHSPFRQLVNR 347
G+ + +A+ ++LLG+N+S VL+G + + +L+ R
Sbjct: 253 GVEMSNKANLEADLLGMNLSE*IVLEGDA*WLELIMR 143
>TC8460 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F18_180
{Arabidopsis thaliana;}, partial (36%)
Length = 880
Score = 32.0 bits (71), Expect = 0.29
Identities = 25/105 (23%), Positives = 44/105 (41%)
Frame = +3
Query: 61 NEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTTAMRML 120
+ P + + FRR +RM K F I +L S+ + + ++ I + + L
Sbjct: 357 SHPDFPEEEFRRSFRMSKATFEMICRELDSA----VTKKNTMLREAIPVRQRVAVCIWRL 524
Query: 121 AYGVAADAVDEYIKIGSSTTLECLLRFCKGIIRLYEQVYLRAPTQ 165
A G V + +G ST + +L C I + +LR P +
Sbjct: 525 ATGDPLRLVAKRFGLGISTCHKLVLEVCSAIKTVLMPKFLRWPDE 659
>TC15168 similar to UP|O04084 (O04084) Serine carboxypeptidase isolog,
30227-33069, partial (55%)
Length = 1005
Score = 30.8 bits (68), Expect = 0.65
Identities = 15/37 (40%), Positives = 18/37 (48%)
Frame = -2
Query: 291 KLYNYDGTLKFPYVFLTNGGGIPEAKRASELSELLGL 327
K++ Y T Y +L GG P K SELLGL
Sbjct: 911 KIFIYGITKSSQYAYLVEGGDSPRRKEEKNASELLGL 801
>TC9532 similar to UP|THIC_MYCLE (Q9ZBL0) Thiamine biosynthesis protein
thiC, partial (14%)
Length = 1259
Score = 27.7 bits (60), Expect = 5.5
Identities = 12/17 (70%), Positives = 14/17 (81%)
Frame = +3
Query: 244 IISLLLLCYPILINDLL 260
I+ LLLLCYPIL N L+
Sbjct: 555 ILLLLLLCYPILSNKLI 605
>AV767290
Length = 484
Score = 27.3 bits (59), Expect = 7.2
Identities = 20/73 (27%), Positives = 33/73 (44%), Gaps = 5/73 (6%)
Frame = -3
Query: 173 HVSETRGFPWMIGSIDCMH----WEWKNYPKAWEGKLAFANHVRARSRVASR-ITSRSSE 227
H + + F M+ DC + +EWK+ P + K+ SRV S+ I + S E
Sbjct: 419 HEGDWKDFVKMLPESDCRYAVVDFEWKDQPTVTKSKICLILWSPEYSRVRSKMIYAASQE 240
Query: 228 AHVDKV*NTETNL 240
A K+ + + L
Sbjct: 239 AVASKMADVQRQL 201
>TC17242
Length = 461
Score = 26.9 bits (58), Expect = 9.4
Identities = 10/17 (58%), Positives = 14/17 (81%)
Frame = -3
Query: 507 PHPSVFKNAEIVLQKHV 523
P P V++ AEI+LQKH+
Sbjct: 135 PSPEVYE*AEIILQKHI 85
>TC16424 similar to UP|Q9FZF1 (Q9FZF1) T2E6.18, partial (39%)
Length = 537
Score = 26.9 bits (58), Expect = 9.4
Identities = 13/48 (27%), Positives = 26/48 (54%)
Frame = +3
Query: 376 IDAYASRFENIDPLAPYKKWTTKLATTQNPKFDESGPQIDVFSERVQA 423
++AY ++ + +D L ++TTKL +P S + S+R++A
Sbjct: 207 VEAYLAKRDGVDKLLKISRYTTKLLLASSPNLLRSDTNL---SQRLKA 341
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.139 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,000,918
Number of Sequences: 28460
Number of extensions: 134245
Number of successful extensions: 604
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 602
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 602
length of query: 605
length of database: 4,897,600
effective HSP length: 96
effective length of query: 509
effective length of database: 2,165,440
effective search space: 1102208960
effective search space used: 1102208960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC135230.3


