

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135230.11 + phase: 0 /pseudo
(954 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
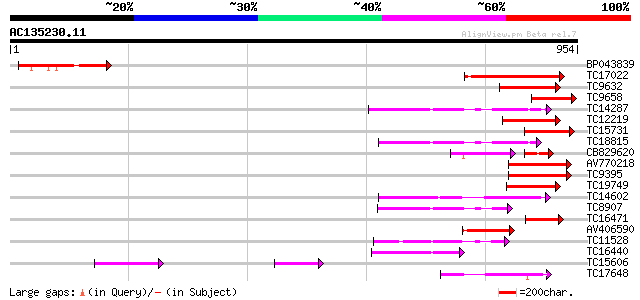
Score E
Sequences producing significant alignments: (bits) Value
BP043839 169 2e-42
TC17022 homologue to UP|Q39183 (Q39183) Serine/threonine protein... 162 2e-40
TC9632 similar to PIR|T47546|T47546 protein kinase-like - Arabid... 130 7e-31
TC9658 similar to UP|Q8H934 (Q8H934) Phototropin, partial (8%) 129 2e-30
TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein k... 124 5e-29
TC12219 similar to UP|Q9LZS4 (Q9LZS4) Protein kinase-like protei... 123 2e-28
TC15731 similar to UP|Q07526 (Q07526) Protein kinase PSPK-5 (Pr... 117 9e-27
TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein ki... 117 1e-26
CB829620 76 2e-22
AV770218 100 2e-21
TC9395 homologue to UP|KPK1_PHAVU (P15792) Protein kinase PVPK-1... 100 2e-21
TC19749 weakly similar to UP|KPK1_PHAVU (P15792) Protein kinase ... 92 3e-19
TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinas... 92 3e-19
TC8907 similar to UP|Q8H0X3 (Q8H0X3) Serine/threonine protein ki... 91 9e-19
TC16471 similar to UP|Q41384 (Q41384) Protein kinase (Fragment) ... 85 5e-17
AV406590 85 6e-17
TC11528 homologue to UP|Q39868 (Q39868) Protein kinase , partial... 78 8e-15
TC16440 similar to UP|Q93Y18 (Q93Y18) SNF1 related protein kinas... 75 6e-14
TC15606 weakly similar to GB|AAL15406.1|16323302|AY058232 At2g02... 74 8e-14
TC17648 similar to UP|Q9AYN9 (Q9AYN9) NQK1 MAPKK, partial (51%) 67 2e-11
>BP043839
Length = 534
Score = 169 bits (427), Expect = 2e-42
Identities = 104/182 (57%), Positives = 120/182 (65%), Gaps = 26/182 (14%)
Frame = +2
Query: 16 SFPRDPRGSLEVFNPTSNST------SPVRSPSHLKTW---TETEEQHKDFIST--DEVT 64
+FPRDPRGSLEVFNP+S+ T SP+R+ K W T ++ K + DEV
Sbjct: 11 AFPRDPRGSLEVFNPSSSLTTEKPTNSPLRTQHSWKDWKDPTRADDPPKPIAGSNADEVA 190
Query: 65 ------NTSWMAIKEGET-------GAAAQRAAEWGLVLRTDAETGKPQGVGVRNSGDDE 111
TSWMAI E + AAAQRAAEWGLVL+TDAETGKPQGVGVR+SG
Sbjct: 191 VAISDPPTSWMAISETASPPPVVRESAAAQRAAEWGLVLKTDAETGKPQGVGVRSSG--- 361
Query: 112 QNGKFSGKRNSNNSGRVSGDSSD--GGDPRGFPRVSEDLKDALSAFQQTFVVSDATKPDY 169
+R+S S R SG+SSD GG+ RG PRVSEDL+DALSAFQQTFVVSDATKPDY
Sbjct: 362 -----GSRRDSKTSVRSSGESSDDGGGEYRGIPRVSEDLRDALSAFQQTFVVSDATKPDY 526
Query: 170 PI 171
PI
Sbjct: 527 PI 532
>TC17022 homologue to UP|Q39183 (Q39183) Serine/threonine protein kinase
(Protein kinase 5) (AT5G47750/MCA23_7) , partial (32%)
Length = 775
Score = 162 bits (411), Expect = 2e-40
Identities = 86/171 (50%), Positives = 112/171 (65%), Gaps = 2/171 (1%)
Frame = +1
Query: 765 TSC-KPQLIIPANEDKKKRKKKKKKGQQKTQQIPTFMAEPMRA-SNSFVGTEEYIAPEII 822
TSC P+L +++ KK RK K + G Q +P +AEP A S SFVGT EY+APEII
Sbjct: 19 TSCFSPRLF--SSKSKKDRKPKNEIGNQ-VSPLPELIAEPTDARSMSFVGTHEYLAPEII 189
Query: 823 TGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQL 882
G GH SAVDWW GI LYE+L+G TPF+G + T N++ + L+FP++ VS A+ L
Sbjct: 190 KGEGHGSAVDWWTFGIFLYELLFGKTPFKGSGNRATLFNVVGQPLRFPEAPVVSFAARDL 369
Query: 883 IYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPELDAPILLE 933
I LL ++P++RL GA EIK HPFF+ VNWALIRC PPE+ + E
Sbjct: 370 IRGLLVKEPQHRLAYKRGATEIKQHPFFEGVNWALIRCATPPEIPKAVEFE 522
>TC9632 similar to PIR|T47546|T47546 protein kinase-like - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (12%)
Length = 650
Score = 130 bits (328), Expect = 7e-31
Identities = 59/103 (57%), Positives = 77/103 (74%)
Frame = +3
Query: 824 GSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLI 883
G GH +AVDWW G+ LYE+LYG TPF+G ++T AN++ + L+FP S VS QA+ LI
Sbjct: 6 GEGHGAAVDWWTFGVFLYELLYGRTPFKGSNNEETLANVVLQGLRFPDSPFVSFQARDLI 185
Query: 884 YWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPEL 926
LL ++P+NRLG+ +GA EIK HPFF+ +NWALIRC PPEL
Sbjct: 186 RGLLVKEPENRLGTEKGAAEIKQHPFFEGLNWALIRCAIPPEL 314
>TC9658 similar to UP|Q8H934 (Q8H934) Phototropin, partial (8%)
Length = 536
Score = 129 bits (325), Expect = 2e-30
Identities = 60/75 (80%), Positives = 67/75 (89%)
Frame = +2
Query: 879 AKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPELDAPILLENDEKK 938
AKQLIY LLHRDPKNRLGS EGANEIK HPFF+ V+WALIRCMKPPELDAP+L E +E K
Sbjct: 5 AKQLIYRLLHRDPKNRLGSQEGANEIKRHPFFRGVDWALIRCMKPPELDAPLLQETEEDK 184
Query: 939 EAKDIDPGLDDLQKN 953
EAKD+DP L+DLQ+N
Sbjct: 185 EAKDVDPRLEDLQEN 229
>TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial
(75%)
Length = 1999
Score = 124 bits (312), Expect = 5e-29
Identities = 87/308 (28%), Positives = 144/308 (46%), Gaps = 1/308 (0%)
Frame = +3
Query: 605 ENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTER 664
EN I + + LG G+ V+ T + A+K + K + V + E
Sbjct: 402 ENPRNIIFNKYEMGRVLGQGNFAKVYHGRNLATNENVAIKVIKKEKLKKDRLVKQIKREV 581
Query: 665 EILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEV 724
++ ++ HP + L TK + L+ +Y GGELF +++ L ED R Y ++
Sbjct: 582 SVMRLVRHPHIVELKEVMATKGKIFLVMEYVKGGELFTKVNKGK---LNEDDARKYFQQL 752
Query: 725 LIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKK 784
+ A+++ H +G+ +RDLKPEN+L+ N + ++DF LS L ++
Sbjct: 753 ISAVDFCHSRGVTHRDLKPENLLLDENEDLKVSDFGLSALP-----------------EQ 881
Query: 785 KKKKGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHT-SAVDWWALGILLYEM 843
++ G T GT Y+APE++ G+ S D W+ G++LY +
Sbjct: 882 RRDDGMLVTP----------------CGTPAYVAPEVLKKKGYDGSKADIWSCGVILYAL 1013
Query: 844 LYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANE 903
L GY PF+G+ + + + +FP+ +SPQAK LI LL DP+ R E
Sbjct: 1014LSGYLPFQGENVMRIYRKAFKAEYEFPEW--ISPQAKNLISNLLVADPEKRY----SIPE 1175
Query: 904 IKSHPFFK 911
I S P+F+
Sbjct: 1176IISDPWFQ 1199
>TC12219 similar to UP|Q9LZS4 (Q9LZS4) Protein kinase-like protein, partial
(11%)
Length = 587
Score = 123 bits (308), Expect = 2e-28
Identities = 56/98 (57%), Positives = 72/98 (73%)
Frame = +3
Query: 829 SAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLH 888
+AVDWW GI L+E+LYG TPF+G + T AN++ + LKFP + VS A+ LI LL
Sbjct: 3 NAVDWWTFGIFLFELLYGKTPFKGLANEDTLANVVSQSLKFPSAPIVSFHARDLIRGLLI 182
Query: 889 RDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPEL 926
+DP+NRLGS++GA EIK HPF + +NWALIRC PPEL
Sbjct: 183 KDPENRLGSVKGAAEIKQHPFSEGLNWALIRCAAPPEL 296
>TC15731 similar to UP|Q07526 (Q07526) Protein kinase PSPK-5 (Probable
serine/threonine-protein kinase PK5) , partial (17%)
Length = 940
Score = 117 bits (293), Expect = 9e-27
Identities = 54/84 (64%), Positives = 66/84 (78%)
Frame = +1
Query: 867 LKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPEL 926
LKFPKSK VS KQL+Y LL RDP +RLGS EGANEIK HPFF+ +NWAL+RC KPPEL
Sbjct: 1 LKFPKSKQVSLSGKQLMYRLLQRDPSSRLGSKEGANEIKRHPFFRGINWALVRCTKPPEL 180
Query: 927 DAPILLENDEKKEAKDIDPGLDDL 950
DAP+ +E+K+AK +DP +D+
Sbjct: 181 DAPLFGTTEEEKKAKYVDPVKEDM 252
>TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein kinase
CIPK25, partial (58%)
Length = 979
Score = 117 bits (292), Expect = 1e-26
Identities = 78/276 (28%), Positives = 132/276 (47%), Gaps = 1/276 (0%)
Frame = +2
Query: 621 LGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILDMLDHPFLPALYA 680
LG G V+ + +G+ A+K M K + + + E I+ ++ HP + L
Sbjct: 56 LGKGTLAKVYFAKEITSGEGVAIKVMSKARIKKEGMMDQIKREISIMRLVRHPNIVNLKE 235
Query: 681 SFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRD 740
TKT + I +Y GGELF + + LK+D R Y +++ A++Y H +G+ +RD
Sbjct: 236 VMATKTKIFFIMEYIRGGELFAKVAKGK---LKDDLARRYFQQLISAVDYCHSRGVSHRD 406
Query: 741 LKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKKGQQKTQQIPTFM 800
LKPEN+L+ N ++ ++DF LS L ++ ++ G TQ
Sbjct: 407 LKPENLLLDENENLKVSDFGLSGLP-----------------EQLRQDGLLHTQ------ 517
Query: 801 AEPMRASNSFVGTEEYIAPEIITGSGHTS-AVDWWALGILLYEMLYGYTPFRGKTRQKTF 859
GT Y+APE++ G+ D W+ G++LY +L G PF+ + +
Sbjct: 518 ----------CGTPAYVAPEVLRKKGYDGFKTDTWSCGVILYALLAGCLPFQHENLMTMY 667
Query: 860 ANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRL 895
+L + +FP SP++K+LI +L DP R+
Sbjct: 668 NKVLRAEFQFPPW--FSPESKKLISKILVADPNRRI 769
>CB829620
Length = 548
Score = 75.9 bits (185), Expect(2) = 2e-22
Identities = 40/115 (34%), Positives = 61/115 (52%), Gaps = 6/115 (5%)
Frame = +1
Query: 742 KPENVLIQRNGHVSLTDFDL----SCLTSCKPQLIIPANEDKKKRKKKKKKGQQKTQQ-- 795
KP+N+L+ RNGH+ L+DF L C + +N + + ++TQQ
Sbjct: 1 KPDNLLLDRNGHMKLSDFGLCKPLDCSNLQENDFSTGSNRSGALQSNGRPVAPKRTQQEQ 180
Query: 796 IPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPF 850
+ + + S VGT +YIAPE++ G+ DWW+LG ++YEML GY PF
Sbjct: 181 LQHWQKNRRTLAYSTVGTPDYIAPEVLLKKGYGMECDWWSLGGIMYEMLVGYPPF 345
Score = 47.8 bits (112), Expect(2) = 2e-22
Identities = 25/49 (51%), Positives = 36/49 (73%)
Frame = +3
Query: 867 LKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNW 915
LKFP+ +SP+AK LI LL + + RLG+ +GA+EIK+HP+FK + W
Sbjct: 399 LKFPEEAKLSPEAKDLISRLLC-NVQQRLGT-KGADEIKAHPWFKGIEW 539
>AV770218
Length = 567
Score = 99.8 bits (247), Expect = 2e-21
Identities = 49/106 (46%), Positives = 65/106 (61%)
Frame = -1
Query: 840 LYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLE 899
LYE+L G TPF+G + T N++ + L FP S VS A+ LI LL ++P++RL
Sbjct: 567 LYELLSGRTPFKGSAYRATLFNVVGQPLSFPVSPAVSFAARDLIRGLLVKEPQHRLAYRR 388
Query: 900 GANEIKSHPFFKNVNWALIRCMKPPELDAPILLENDEKKEAKDIDP 945
GA EIK HPFF NVNWALIRC PPE+ P L+ ++ + P
Sbjct: 387 GATEIKQHPFFHNVNWALIRCTNPPEVPRPTLMRPSPTEKELGVKP 250
>TC9395 homologue to UP|KPK1_PHAVU (P15792) Protein kinase PVPK-1 ,
partial (17%)
Length = 590
Score = 99.8 bits (247), Expect = 2e-21
Identities = 48/106 (45%), Positives = 67/106 (62%)
Frame = +2
Query: 840 LYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLE 899
LYE+L+G TPF+G + T N++ + L+FP+S VS A+ LI LL ++P++RL
Sbjct: 5 LYELLFGRTPFKGSANRATLFNVVGQPLRFPESPSVSFAARDLIRGLLVKEPQHRLAYRR 184
Query: 900 GANEIKSHPFFKNVNWALIRCMKPPELDAPILLENDEKKEAKDIDP 945
GA EIK HPFF NVNWALIRC PPE+ + ++ A + P
Sbjct: 185 GATEIKQHPFFHNVNWALIRCASPPEVPRQAMKAVAAERVAPGVKP 322
>TC19749 weakly similar to UP|KPK1_PHAVU (P15792) Protein kinase PVPK-1 ,
partial (14%)
Length = 524
Score = 92.4 bits (228), Expect = 3e-19
Identities = 46/93 (49%), Positives = 60/93 (64%), Gaps = 3/93 (3%)
Frame = +3
Query: 837 GILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSP---QAKQLIYWLLHRDPKN 893
GI +YEM+YG TPF G + T +IL K L FP + P S A+ LI LL++DP
Sbjct: 6 GIFIYEMVYGRTPFAGPSNDVTLRSILKKPLVFPTATPSSALEMHARDLISGLLNKDPNR 185
Query: 894 RLGSLEGANEIKSHPFFKNVNWALIRCMKPPEL 926
RLGS+ GA ++K HPFF +N ALIR + PPE+
Sbjct: 186 RLGSMRGAADVKKHPFFAGINLALIRMVAPPEV 284
>TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinase, complete
Length = 1256
Score = 92.4 bits (228), Expect = 3e-19
Identities = 74/294 (25%), Positives = 131/294 (44%), Gaps = 4/294 (1%)
Frame = +2
Query: 621 LGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILDMLD-HPFLPALY 679
+G G G++ + FA+K +DK ++ + H E + + +L HP + ++
Sbjct: 86 IGRGRFGTIFRCFHPISTDTFAVKLIDKSLLADSTDRHCLENEPKYMSLLSPHPNILQIF 265
Query: 680 ASFQTKTHVCLITDY-YPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIY 738
F+ + ++ + P L ++ T + + +A ++L A+ + H G+ +
Sbjct: 266 DVFEDDDVLSMVIELCQPLTLLDRIVAANGTSIPEVEAAGLMK-QLLEAVAHCHRLGVAH 442
Query: 739 RDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKKGQQKTQQIPT 798
RD+KP+NVL G + L DF +
Sbjct: 443 RDVKPDNVLFGGGGDLKLADFGSA-----------------------------------E 517
Query: 799 FMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKT 858
+ + R S VGT Y+APE++ G + VD W+ G++LY ML G PF G + +
Sbjct: 518 WFGDGRRMSG-VVGTPYYVAPEVLMGREYGEKVDVWSCGVILYIMLSGTPPFYGDSAAEI 694
Query: 859 FANILHKDLKFPKS--KPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFF 910
F ++ +L+FP + VSP AK L+ ++ RDP NR+ A + HP+F
Sbjct: 695 FEAVIRGNLRFPSRIFRNVSPAAKDLLRKMICRDPSNRI----SAEQALRHPWF 844
>TC8907 similar to UP|Q8H0X3 (Q8H0X3) Serine/threonine protein kinase-like
protein, partial (44%)
Length = 941
Score = 90.9 bits (224), Expect = 9e-19
Identities = 62/229 (27%), Positives = 106/229 (46%), Gaps = 1/229 (0%)
Frame = +3
Query: 619 KPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILDMLDHPFLPAL 678
K LG G+ V+ T + A+K + K + V + E ++ ++ HP + L
Sbjct: 354 KILGQGNFAKVYHGRNMETNESVAIKVIKKERLKKERLVKQIKREVSVMRLVRHPHIVEL 533
Query: 679 YASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIY 738
TKT + ++ +Y GGELF L + + E A R Y +++ A+++ H +G+ +
Sbjct: 534 KEVMATKTKIFMVVEYVKGGELFAKLTKGK---MTEVAARKYFQQLISAVDFCHSRGVTH 704
Query: 739 RDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKKGQQKTQQIPT 798
RDLKPEN+L+ N + ++DF LS L ++++ G T
Sbjct: 705 RDLKPENLLLDDNEDLKVSDFGLSSLP-----------------EQRRSDGMLLTP---- 821
Query: 799 FMAEPMRASNSFVGTEEYIAPEIITGSGHT-SAVDWWALGILLYEMLYG 846
GT Y+APE++ G+ S D W+ G++LY +L G
Sbjct: 822 ------------CGTPAYVAPEVLKKKGYDGSKADIWSCGVILYALLCG 932
>TC16471 similar to UP|Q41384 (Q41384) Protein kinase (Fragment) , partial
(12%)
Length = 617
Score = 85.1 bits (209), Expect = 5e-17
Identities = 41/64 (64%), Positives = 46/64 (71%)
Frame = +3
Query: 869 FPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPELDA 928
FP S PVS A+QLI LL RDP +RLGS GANEIK HPFF+ +NW LIR M PP LD
Sbjct: 3 FPSSIPVSLAARQLINALLQRDPASRLGSTTGANEIKQHPFFREINWPLIRNMSPPPLDV 182
Query: 929 PILL 932
P+ L
Sbjct: 183 PLQL 194
>AV406590
Length = 406
Score = 84.7 bits (208), Expect = 6e-17
Identities = 43/88 (48%), Positives = 54/88 (60%), Gaps = 1/88 (1%)
Frame = +3
Query: 762 SCLTSCKPQLIIPANEDKKKRKKKKKKGQQKTQQIPTFMAEPMRA-SNSFVGTEEYIAPE 820
SC T P+ + ++ KK+ K K + +P MAEP A S SFVGT EY+APE
Sbjct: 150 SCFT---PRFLSGKSKKDKKKSKPKTDIHNQVTPLPELMAEPTNARSMSFVGTHEYLAPE 320
Query: 821 IITGSGHTSAVDWWALGILLYEMLYGYT 848
I+ G GH SAVDWW GI LYE+L+G T
Sbjct: 321 IVKGEGHGSAVDWWTFGIFLYELLFGRT 404
>TC11528 homologue to UP|Q39868 (Q39868) Protein kinase , partial (56%)
Length = 640
Score = 77.8 bits (190), Expect = 8e-15
Identities = 63/233 (27%), Positives = 100/233 (42%), Gaps = 5/233 (2%)
Frame = +3
Query: 613 KHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILD--ML 670
+ + P+K LGSG+ G L + TG+ A+K +++G ++ N +REI++ L
Sbjct: 72 ERYEPLKELGSGNFGVARLARDKNTGELVAVKYIERGKKIDEN------VQREIINHRSL 233
Query: 671 DHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEY 730
HP + T TH+ ++ +Y GGELF + ED R++ +++ + Y
Sbjct: 234 RHPNIIRFKEVLLTPTHLAIVLEYASGGELFERICSAGR--FSEDEARYFFQQLISGVSY 407
Query: 731 LHCQGIIYRDLKPENVLIQRN--GHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKK 788
H I +RDLK EN L+ N + + DF S K
Sbjct: 408 CHSMEICHRDLKLENTLLDGNPSPRLKICDFGYS--------------------KSAILH 527
Query: 789 GQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAV-DWWALGILL 840
Q K S VGT YIAPE+++ + V D W+ G+ L
Sbjct: 528 SQPK----------------STVGTPAYIAPEVLSRKEYDGKVADVWSCGVTL 638
>TC16440 similar to UP|Q93Y18 (Q93Y18) SNF1 related protein kinase, partial
(30%)
Length = 549
Score = 74.7 bits (182), Expect = 6e-14
Identities = 48/157 (30%), Positives = 82/157 (51%), Gaps = 1/157 (0%)
Frame = +1
Query: 609 QISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILD 668
+I L ++ + LG G+ V+ G A+K +DK ++ + R E + +
Sbjct: 73 RIILGKYQLTRFLGRGNFAKVYQAVSLTDGTTVAVKMIDKSKTVDASMEPRIVREIDAMR 252
Query: 669 MLDH-PFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIA 727
L H P + ++ T+T + LI DY GGELF + ++ L E R Y +++ A
Sbjct: 253 RLQHHPNILKIHEVMATRTKIYLIVDYAGGGELFSKISRRGR--LPEPLARRYFQQLVSA 426
Query: 728 LEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCL 764
L + H G+ +RDLKP+N+L+ G++ ++DF LS L
Sbjct: 427 LCFCHRNGVAHRDLKPQNLLLDAEGNLKVSDFGLSAL 537
>TC15606 weakly similar to GB|AAL15406.1|16323302|AY058232
At2g02710/T20F6.15 {Arabidopsis thaliana;}, partial
(25%)
Length = 642
Score = 74.3 bits (181), Expect = 8e-14
Identities = 43/118 (36%), Positives = 63/118 (52%), Gaps = 2/118 (1%)
Frame = +1
Query: 143 RVSEDLKDALSAFQQT-FVVSDATKPDYPIMYASAGFFNMTGYTSKEVIGRNCRFLQGAD 201
R S +++L T F ++D + +PI++AS GF MTG++ EV+GR QG
Sbjct: 289 RYSRHARESLDELTDTSFTITDPSVSGHPIVFASLGFLKMTGFSRDEVVGRGGGMFQGPG 468
Query: 202 TDPQDVAKIREALEGGKSYCGRLLNYKKDGTPFWNLLTISPI-KDDDGNVLKLIGMLV 258
T + V IREA+ K LNY+KDGTPF LL + P+ G V+ + + V
Sbjct: 469 TCRRSVMAIREAVREEKEAQVVXLNYRKDGTPFXMLLLVCPVFSASSGGVVHFVAVQV 642
Score = 53.9 bits (128), Expect = 1e-07
Identities = 31/84 (36%), Positives = 47/84 (55%), Gaps = 1/84 (1%)
Frame = +1
Query: 446 LYFLELTEYSREEILGRNCRFLQGPETDPATVRKIREAIDNQTEVTVQLINYTRTGKKFW 505
L FL++T +SR+E++GR QGP T +V IREA+ + E V +NY + G F
Sbjct: 391 LGFLKMTGFSRDEVVGRGGGMFQGPGTCRRSVMAIREAVREEKEAQVVXLNYRKDGTPFX 570
Query: 506 NLFHLQPM-RDHKGEVQYFIGVQL 528
L + P+ G V +F+ VQ+
Sbjct: 571 MLLLVCPVFSASSGGVVHFVAVQV 642
>TC17648 similar to UP|Q9AYN9 (Q9AYN9) NQK1 MAPKK, partial (51%)
Length = 741
Score = 66.6 bits (161), Expect = 2e-11
Identities = 53/195 (27%), Positives = 87/195 (44%), Gaps = 8/195 (4%)
Frame = +3
Query: 725 LIALEYLHCQG-IIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRK 783
L L YLH + +I+RD KP ++L+ G V +TDF +S +
Sbjct: 3 LQGLVYLHNERHVIHRDSKPSHLLVNHQGEVKITDFGVSAM------------------- 125
Query: 784 KKKKKGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEM 843
+A M ++FVGT Y++P I+GS + + D W+LG+++ E
Sbjct: 126 ----------------LATSMGQRDTFVGTYNYMSPARISGSTYDYSSDIWSLGMVVLEC 257
Query: 844 LYGYTPFRGKTRQK---TFANILHKDLKFP----KSKPVSPQAKQLIYWLLHRDPKNRLG 896
G P+ Q+ +F +L ++ P S SP+ + + +DP++RL
Sbjct: 258 AIGRFPYIQSEDQQAWPSFYELLAAIVESPPPSAPSDQFSPEFCSFVSSCIQKDPQDRLT 437
Query: 897 SLEGANEIKSHPFFK 911
SL E+ HPF K
Sbjct: 438 SL----ELLDHPFIK 470
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,376,869
Number of Sequences: 28460
Number of extensions: 208891
Number of successful extensions: 1954
Number of sequences better than 10.0: 197
Number of HSP's better than 10.0 without gapping: 1549
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1739
length of query: 954
length of database: 4,897,600
effective HSP length: 99
effective length of query: 855
effective length of database: 2,080,060
effective search space: 1778451300
effective search space used: 1778451300
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC135230.11


