

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
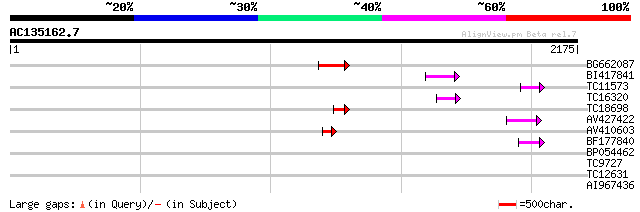
Query= AC135162.7 + phase: 0 /pseudo
(2175 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG662087 137 1e-32
BI417841 73 4e-13
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 63 6e-10
TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial ... 60 3e-09
TC18698 57 2e-08
AV427422 55 1e-07
AV410603 53 6e-07
BF177840 48 1e-05
BP054462 30 5.2
TC9727 weakly similar to UP|Q9FV67 (Q9FV67) Fatty acid elongase ... 29 6.8
TC12631 29 8.9
AI967436 29 8.9
>BG662087
Length = 373
Score = 137 bits (346), Expect = 1e-32
Identities = 65/119 (54%), Positives = 86/119 (71%)
Frame = +1
Query: 1184 GKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNQIKMSPEDREK 1243
GK RM VD+ DLNKA PKD++PLP ID LVD + +++ S MD +SGY+QIKM P D +K
Sbjct: 16 GKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDEDK 195
Query: 1244 TSFITPWGTFCYKVMPFGLINAGATYQRGMTTLFHDMIHKEVEVYVDDMIVKSADEEQH 1302
T+F+T +CY+ +PFGL NAGATYQ M +F D + + +EVY+D+MIVKSA H
Sbjct: 196 TAFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRANH 372
>BI417841
Length = 617
Score = 73.2 bits (178), Expect = 4e-13
Identities = 40/131 (30%), Positives = 66/131 (49%), Gaps = 2/131 (1%)
Frame = +1
Query: 1596 LVFDGAV--NAYGKGIGAVIVSPQGHHIPFTTRILFECTNNMAEYEACIFGIEEAIDMRI 1653
L FDG+ N G GAV+ + G + + + TNN AEY I G++ A +
Sbjct: 142 LEFDGSSKGNPGSAGAGAVLRAEDGSKV-YLREGVGNQTNNQAEYRGLILGLKHAHEQGY 318
Query: 1654 KHLDIYGDSALIINQIKGEWETHHANLIPYRDYARRLLTYFTKVELHHIPRDENQMADAL 1713
+H+++ GDS L+ Q++G W+ + N+ + A+ L + F +++H+PR N AD
Sbjct: 319 QHINVKGDSQLVCKQVEGSWKARNPNIASLCNEAKELKSKFQSFDINHVPRQYNSEADVQ 498
Query: 1714 ATLSSMFRVNH 1724
A L H
Sbjct: 499 ANLGVNLPAGH 531
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 62.8 bits (151), Expect = 6e-10
Identities = 36/96 (37%), Positives = 49/96 (50%), Gaps = 1/96 (1%)
Frame = +2
Query: 1958 DNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEAANKNI-KRIVQKMVTTYKDWHE 2016
+NG + Q C+ I+ SS PQ NG EAANK I K I +++ W +
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 2017 MLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEV 2052
LP L Y T +SS TPF L YG + +LP+E+
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEI 295
>TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (11%)
Length = 632
Score = 60.5 bits (145), Expect = 3e-09
Identities = 29/91 (31%), Positives = 45/91 (48%)
Frame = +2
Query: 1638 YEACIFGIEEAIDMRIKHLDIYGDSALIINQIKGEWETHHANLIPYRDYARRLLTYFTKV 1697
Y I G++ AI KH+ + GDS L+ NQ++G W+ + N+ A+ L F
Sbjct: 2 YRGLILGLKHAIKEGYKHIQVKGDSMLVCNQVQGLWKIKNQNIASLCSEAKELKNKFLSF 181
Query: 1698 ELHHIPRDENQMADALATLSSMFRVNHWNDV 1728
+++HIPR+ N AD A R +V
Sbjct: 182 KINHIPREYNSEADVQANFGISLRAGQVEEV 274
>TC18698
Length = 808
Score = 57.4 bits (137), Expect = 2e-08
Identities = 31/62 (50%), Positives = 39/62 (62%)
Frame = -2
Query: 1241 REKTSFITPWGTFCYKVMPFGLINAGATYQRGMTTLFHDMIHKEVEVYVDDMIVKSADEE 1300
++KT+ + Y+VMP GL N TYQR M +FH I K VEVYV+DMIVKS+ E
Sbjct: 807 KKKTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSSQE* 628
Query: 1301 QH 1302
H
Sbjct: 627 FH 622
>AV427422
Length = 417
Score = 55.1 bits (131), Expect = 1e-07
Identities = 33/133 (24%), Positives = 63/133 (46%), Gaps = 2/133 (1%)
Frame = +1
Query: 1907 SNGHRFILVAIDYFTKWVEAASYTN-VTKQVVAKFIKNNIICRYGVPSKIITDNGTNLNN 1965
S G+ +LV +D +K+ + T +V+A ++ +GVP I++D +
Sbjct: 19 SKGYEAVLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLFMS 198
Query: 1966 NVVQALCEEFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQKMVTTY-KDWHEMLPYALHG 2024
N + L + + S+ Y P+ +G E N+ ++ ++ + K W +P+A +
Sbjct: 199 NFWKELFKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWAEYW 378
Query: 2025 YRTTVRSSTGATP 2037
Y T+ STG TP
Sbjct: 379 YNTSYHVSTGQTP 417
>AV410603
Length = 162
Score = 52.8 bits (125), Expect = 6e-07
Identities = 26/51 (50%), Positives = 34/51 (65%)
Frame = +1
Query: 1201 KDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNQIKMSPEDREKTSFITPWG 1251
KD+FP+P +D L+D S+ FS +D SGY+QI + PEDR KT F T G
Sbjct: 10 KDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTHHG 162
>BF177840
Length = 410
Score = 48.1 bits (113), Expect = 1e-05
Identities = 26/101 (25%), Positives = 49/101 (47%), Gaps = 1/101 (0%)
Frame = +2
Query: 1953 SKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQKMVT-TY 2011
+ I++D T ++ + L + + S+ PQ +G E NK + +++ ++
Sbjct: 11 TSIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNL 190
Query: 2012 KDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEV 2052
K W LP+ Y V S+T +PF +VYG + PL++
Sbjct: 191 KMWETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDL 313
>BP054462
Length = 422
Score = 29.6 bits (65), Expect = 5.2
Identities = 34/118 (28%), Positives = 50/118 (41%), Gaps = 6/118 (5%)
Frame = +1
Query: 2032 STGATPFSLVYGMEAVLPLE-VEIPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAMA 2090
S +PF +YG E + L+ IPS K++ Q D+L D A
Sbjct: 19 SAKMSPFQALYGREPPVLLQGTTIPS-------KIAAVNDLQVGRDEL------LSDLRA 159
Query: 2091 RGQSYQARMKTAFDKKVHPREFKVG-ELVLK----RRISQQPDPRGKWTPNYEGPYVV 2143
Q M+T +KK ++++G E+ LK RR S K +P Y GPY +
Sbjct: 160 NLLKSQDMMRTYANKKRRDVDYQIGDEVFLKLQPYRRRSLAKKMNEKLSPRYYGPYPI 333
>TC9727 weakly similar to UP|Q9FV67 (Q9FV67) Fatty acid elongase 1-like
protein, partial (40%)
Length = 731
Score = 29.3 bits (64), Expect = 6.8
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +1
Query: 891 LHLLHQEALRQIGMPLTFLQSCMFLSNVSCCF 922
LH LHQ L +G P L C +L + +C F
Sbjct: 238 LHFLHQRYL*HMGQPAIQLDICHYLLHSACIF 333
>TC12631
Length = 870
Score = 28.9 bits (63), Expect = 8.9
Identities = 33/104 (31%), Positives = 49/104 (46%), Gaps = 5/104 (4%)
Frame = -2
Query: 878 MLSLKKPLDLVSDLHLLHQEALRQIGMPLTFLQSCMFLSNVSCCF--NALFSK*QS---S 932
+LSLK+P+ +V+ L L ++G F S LSN SCCF N + S + S
Sbjct: 434 ILSLKEPV-VVASLCC----NLLELGCLEGFSMSTTTLSNFSCCFCSNGVVSCESAPTLS 270
Query: 933 RSAQSESDIFCKGMFVSALPLINKILSFLPRLFFFSAFSLGNNG 976
S+ S I C + L L + LP F+++FS+ G
Sbjct: 269 FSSYFSSSIICGIVIPFPLALTTAAI*LLP---FWNSFSISYRG 147
>AI967436
Length = 294
Score = 28.9 bits (63), Expect = 8.9
Identities = 18/70 (25%), Positives = 31/70 (43%), Gaps = 13/70 (18%)
Frame = +2
Query: 1771 LSREYPPGASKQDKKTLRRLAGRFLLDG------------NILYKRNYDMVLLRCVDEHE 1818
L++ Y P K+ K +R L+G L ++YKR + C+DE +
Sbjct: 80 LTKWYSPYTHKERNKIIRELSGMVLSRAPKLCNFVEWRGQKVVYKRYASLYFCMCIDEED 259
Query: 1819 AE-QLMHDVH 1827
E Q++ +H
Sbjct: 260 NERQVLEMIH 289
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.340 0.148 0.492
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,346,165
Number of Sequences: 28460
Number of extensions: 585476
Number of successful extensions: 4609
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 4470
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4606
length of query: 2175
length of database: 4,897,600
effective HSP length: 105
effective length of query: 2070
effective length of database: 1,909,300
effective search space: 3952251000
effective search space used: 3952251000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC135162.7


