

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135162.11 + phase: 0 /pseudo
(385 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
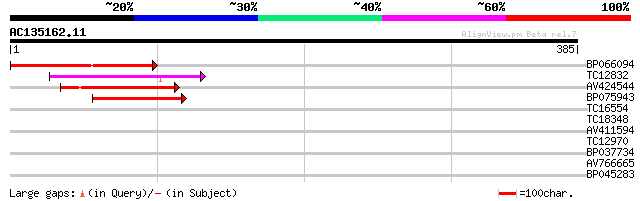
Score E
Sequences producing significant alignments: (bits) Value
BP066094 82 2e-16
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 69 1e-12
AV424544 59 1e-09
BP075943 57 4e-09
TC16554 homologue to UP|AAN71719 (AAN71719) Glycogen synthase ki... 27 4.4
TC18348 similar to UP|BAD09200 (BAD09200) Short-chain dehydrogen... 27 4.4
AV411594 24 6.7
TC12970 27 7.5
BP037734 26 9.8
AV766665 26 9.8
BP045283 26 9.8
>BP066094
Length = 532
Score = 81.6 bits (200), Expect = 2e-16
Identities = 41/100 (41%), Positives = 65/100 (65%)
Frame = +2
Query: 1 NEDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLY 60
N D IF+L K+LYGLKQ+ RAW +R+ +FL++ E V+ K + ++ K KD L+ + +Y
Sbjct: 236 NSDHIFKLKKSLYGLKQAPRAWYERLSSFLLENEXVRGKVDTTLFCKTYKDDILI-VQIY 412
Query: 61 VDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGME 100
VDD+I +N + + F M +F M +G+L YFLG++
Sbjct: 413 VDDIIFGSANPSLCKEFSEMMQAEFEMRMMGELKYFLGIQ 532
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 69.3 bits (168), Expect = 1e-12
Identities = 33/108 (30%), Positives = 59/108 (54%), Gaps = 2/108 (1%)
Frame = +2
Query: 28 NFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDLIVTDSNLAEIEAFKGHMMKKFNM 87
+F++ + + +++ Y K D++ + + LYVDD++V N ++ K + ++F+M
Sbjct: 8 SFIMSLGYNRLSSDHCTYHKRFDDNDFIILLLYVDDMLVVGPNKDRVQELKAQLAREFDM 187
Query: 88 TDLGKLTYFLGMEF--TKTSEGLVKHQKKYASDILKRFNMLNCNPAST 133
DLG LGM+ + + QK Y +L+RFNM +CNP ST
Sbjct: 188 KDLGPANKILGMQIHRDRKDRRIWLSQKNYLQKVLRRFNMQDCNPIST 331
>AV424544
Length = 276
Score = 58.9 bits (141), Expect = 1e-09
Identities = 29/81 (35%), Positives = 52/81 (63%)
Frame = +3
Query: 35 FVKCKAEYNVYVKNSKDSNLVHICLYVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLT 94
+++ +++++ K +D++ I +YVDDLI+ ++L EI+ K + +F + DLG L
Sbjct: 36 YIQSAHDHSLFTK-FRDASFTVILVYVDDLILAGNDLNEIQCVKNKLDIQFRIKDLGTLK 212
Query: 95 YFLGMEFTKTSEGLVKHQKKY 115
YFLG+E ++S GL Q+KY
Sbjct: 213 YFLGLEVARSSCGLFLSQRKY 275
>BP075943
Length = 547
Score = 57.4 bits (137), Expect = 4e-09
Identities = 23/64 (35%), Positives = 44/64 (67%)
Frame = -3
Query: 57 ICLYVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYA 116
+ LYVDD+++ +++ EI+ K + +F + DLG+ YFLG+E +++ G+V +Q+KYA
Sbjct: 521 LLLYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIARSTSGIVLNQRKYA 342
Query: 117 SDIL 120
++
Sbjct: 341 LQLI 330
>TC16554 homologue to UP|AAN71719 (AAN71719) Glycogen synthase kinase 3 beta
protein kinase DWARF12 , partial (71%)
Length = 927
Score = 27.3 bits (59), Expect = 4.4
Identities = 22/79 (27%), Positives = 38/79 (47%), Gaps = 8/79 (10%)
Frame = -1
Query: 24 KRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDD--------LIVTDSNLAEIE 75
KRI ++++ + + +V ++ S + NLVHI L++DD +++ D
Sbjct: 648 KRINQYILRLQISVANSRNSVDIRQSPE-NLVHIKLHIDDRHSLISITVVLDDPIHGFWN 472
Query: 76 AFKGHMMKKFNMTDLGKLT 94
F + KKF T GK T
Sbjct: 471 IFHH*IQKKFICTCSGKET 415
>TC18348 similar to UP|BAD09200 (BAD09200) Short-chain
dehydrogenase/reductase family protein-like, partial
(12%)
Length = 611
Score = 27.3 bits (59), Expect = 4.4
Identities = 13/35 (37%), Positives = 19/35 (54%), Gaps = 1/35 (2%)
Frame = +3
Query: 25 RIVNFLVQQ-EFVKCKAEYNVYVKNSKDSNLVHIC 58
RIVNF + C N+Y+ ++K L+HIC
Sbjct: 102 RIVNFFKPYFNILNCCLIVNLYITSNKSCMLIHIC 206
>AV411594
Length = 244
Score = 23.9 bits (50), Expect(2) = 6.7
Identities = 10/35 (28%), Positives = 18/35 (50%)
Frame = +2
Query: 96 FLGMEFTKTSEGLVKHQKKYASDILKRFNMLNCNP 130
FL K+ +G++ ++KYA +L + C P
Sbjct: 50 FLRT*IAKSKKGILLSRRKYALQLLDDTRNMGCKP 154
Score = 21.2 bits (43), Expect(2) = 6.7
Identities = 8/20 (40%), Positives = 12/20 (60%)
Frame = +3
Query: 81 MMKKFNMTDLGKLTYFLGME 100
+ K F + L + YFLG+E
Sbjct: 6 LAKSFKLKVLDDMKYFLGLE 65
>TC12970
Length = 565
Score = 26.6 bits (57), Expect = 7.5
Identities = 11/28 (39%), Positives = 18/28 (64%)
Frame = +2
Query: 324 CCMEEASILKQSFIFSENK*TKRLSSSY 351
CC++ +L SF+FS +K +K L S+
Sbjct: 410 CCLDRCIVLFFSFVFSSSKISKLLFFSF 493
>BP037734
Length = 490
Score = 26.2 bits (56), Expect = 9.8
Identities = 30/109 (27%), Positives = 45/109 (40%), Gaps = 5/109 (4%)
Frame = +2
Query: 12 LYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNS-----KDSNLVHICLYVDDLIV 66
LYGLKQ R W +R F++ KA +NV V K L C+ + L
Sbjct: 194 LYGLKQLPRDWFERFT-------FLRSKAVHNVKVLRL*GIQ*KGRELC*FCM*MKVLTG 352
Query: 67 TDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKY 115
D + +++ F + L F G + K +G+V Q+KY
Sbjct: 353 DD*DERQVKEFS---YSRI*NRRPRSLNMFFG-NYCKI*KGIVVSQRKY 487
>AV766665
Length = 601
Score = 26.2 bits (56), Expect = 9.8
Identities = 13/45 (28%), Positives = 23/45 (50%)
Frame = +1
Query: 56 HICLYVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGME 100
H+C+YVD + + + E + + + +F DL KL Y +E
Sbjct: 112 HLCVYVDHMTIVGDDEVEKQILREKLAPQF---DLKKLKYISVIE 237
>BP045283
Length = 484
Score = 26.2 bits (56), Expect = 9.8
Identities = 12/46 (26%), Positives = 25/46 (54%)
Frame = +3
Query: 29 FLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDLIVTDSNLAEI 74
FL + + +K K +++ + S L+H+CL ++ + S+L I
Sbjct: 309 FLEKMDLIKKKKNSILHLLTKQASQLIHVCLIIEAINKICSHLTII 446
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.366 0.163 0.562
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,725,124
Number of Sequences: 28460
Number of extensions: 68316
Number of successful extensions: 765
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 547
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 214
Number of HSP's gapped (non-prelim): 573
length of query: 385
length of database: 4,897,600
effective HSP length: 92
effective length of query: 293
effective length of database: 2,279,280
effective search space: 667829040
effective search space used: 667829040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC135162.11


