

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
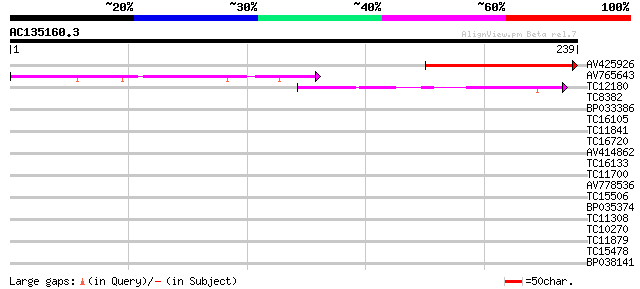
Query= AC135160.3 - phase: 0
(239 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV425926 76 6e-15
AV765643 51 2e-07
TC12180 43 6e-05
TC8382 similar to UP|Q84WK9 (Q84WK9) At4g33985, partial (32%) 33 0.044
BP033386 31 0.17
TC16105 similar to UP|Q84L33 (Q84L33) RAD23-like protein, partia... 31 0.22
TC11841 weakly similar to GB|BAC57345.1|28411900|AP005473 recept... 29 0.83
TC16720 homologue to UP|Q9XH46 (Q9XH46) THA4 protein precursor, ... 28 1.1
AV414862 28 1.1
TC16133 similar to UP|Q39449 (Q39449) Specific tissue protein 1,... 28 1.8
TC11700 homologue to GB|AAL06971.1|15810091|AY056083 AT4g30260/F... 28 1.8
AV778536 27 4.1
TC15506 similar to UP|Q9SEE9 (Q9SEE9) Arginine/serine-rich prote... 27 4.1
BP035374 26 5.4
TC11308 similar to UP|CENB_HUMAN (P07199) Major centromere autoa... 26 7.0
TC10270 homologue to UP|Q8VXT0 (Q8VXT0) Translation initiation f... 25 9.2
TC11879 weakly similar to UP|OREX_HUMAN (O43612) Orexin precurso... 25 9.2
TC15478 homologue to UP|AAQ08016 (AAQ08016) Expansin, partial (33%) 25 9.2
BP038141 25 9.2
>AV425926
Length = 408
Score = 75.9 bits (185), Expect = 6e-15
Identities = 33/64 (51%), Positives = 46/64 (71%)
Frame = -3
Query: 176 EEHKKIDENVLSDNKPYLSEAWDVFDQRERKNPLVNWRVPDKGSEIDMKDNLKFWAHAVA 235
EE ++ D N +PYLSEAW+ +R+++NPL+NW+VP +EIDMKD+L+ WAH VA
Sbjct: 406 EEEEEEDSNESLVPRPYLSEAWEFHGKRKKENPLMNWKVPSLKNEIDMKDSLRLWAHTVA 227
Query: 236 SIAR 239
S R
Sbjct: 226 STVR 215
>AV765643
Length = 452
Score = 50.8 bits (120), Expect = 2e-07
Identities = 49/149 (32%), Positives = 74/149 (48%), Gaps = 18/149 (12%)
Frame = +1
Query: 1 MSAEQILTLLDSFWFETTILTNKNPSK--------VEEILPQDTNLLLVPTPNL----DV 48
M AE ++ L DSFW + IL + SK VE + ++++ VP +
Sbjct: 19 MEAEHVMNLFDSFWLDQNILNKTSSSKSSTNSAENVENQVVKESSDSEVPKLHRIRTGHT 198
Query: 49 RFYSEQNLSIIGSVFSDSPSPNSVLTSSKLRTIPSEREFGEF-STGTSSENHHQKEDVAY 107
R S+Q SI S +S SPNSVL + KL+TI S +E + + T HH +V
Sbjct: 199 RSMSDQ--SIATSFNHESLSPNSVLLAPKLQTIFSGKEVVDSEAEETPVVQHH---EVLL 363
Query: 108 TKKKQ-----SQSYRRRRRKSRSLSELEF 131
++K S +++ R S+SLS+LEF
Sbjct: 364 SEKNNNGNSTSCEKKKKNRGSKSLSDLEF 450
>TC12180
Length = 565
Score = 42.7 bits (99), Expect = 6e-05
Identities = 35/115 (30%), Positives = 52/115 (44%), Gaps = 1/115 (0%)
Frame = +3
Query: 122 KSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDEEHKKI 181
+S+SL++ + +ELKG +DLGF FS D+ +L + +P L+ HK
Sbjct: 96 RSKSLTDDDLEELKGCVDLGFGFSY-DEIPELRNTLPALELC----------YSMSHK-- 236
Query: 182 DENVLSDNKPYLSEAWDVFDQRERKNPLVNWRVPDKGSEI-DMKDNLKFWAHAVA 235
S A D + + NW++ G D+K LKFWA AVA
Sbjct: 237 -----------FSSAADSSSDPPPADSIANWKISSPGDHPEDVKARLKFWAQAVA 368
>TC8382 similar to UP|Q84WK9 (Q84WK9) At4g33985, partial (32%)
Length = 1176
Score = 33.1 bits (74), Expect = 0.044
Identities = 31/127 (24%), Positives = 54/127 (42%), Gaps = 6/127 (4%)
Frame = +3
Query: 119 RRRKSRSLSELEFKELKGFMDLGFVFS---EEDKDSKLVSLIPGLQRLGRENDDAEEGED 175
+ R+++S+++ + ELK ++LGF F E + D +L +P L+ N E
Sbjct: 282 KNRRTKSVTDEDVDELKACIELGFGFESSPEVELDRRLSDTLPALELYHAVN--KSYSES 455
Query: 176 EEHKKIDENVLSDNKPYLSEAWDVFDQRERKNPLVNWRVPDKGSEID---MKDNLKFWAH 232
HK S + D+ +P+ + R G++ D +K L+ WA
Sbjct: 456 RFHKPPTATAPSSSA--------ASDRESTPSPISSPRTAIFGTDDDPQTVKTRLRQWAQ 611
Query: 233 AVASIAR 239
VA R
Sbjct: 612 VVACAVR 632
>BP033386
Length = 498
Score = 31.2 bits (69), Expect = 0.17
Identities = 14/30 (46%), Positives = 19/30 (62%), Gaps = 1/30 (3%)
Frame = -3
Query: 207 NPLVNWRVPDKGSEI-DMKDNLKFWAHAVA 235
+P+ NWR+ G + D+K LK WA AVA
Sbjct: 274 SPISNWRISSPGDDPRDVKARLKVWAQAVA 185
>TC16105 similar to UP|Q84L33 (Q84L33) RAD23-like protein, partial (34%)
Length = 608
Score = 30.8 bits (68), Expect = 0.22
Identities = 18/53 (33%), Positives = 27/53 (49%)
Frame = +3
Query: 156 LIPGLQRLGRENDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRERKNP 208
L P LQ LG++N DE H + + + N+P D+FDQ E++ P
Sbjct: 66 LQPVLQELGKQNPSLLRLIDEHHAEFLQLI---NEPVEGSEGDIFDQPEQEMP 215
>TC11841 weakly similar to GB|BAC57345.1|28411900|AP005473 receptor protein
kinase CLAVATA1 precursor-like porotein {Oryza sativa
(japonica cultivar-group);} , partial (11%)
Length = 504
Score = 28.9 bits (63), Expect = 0.83
Identities = 25/85 (29%), Positives = 36/85 (41%), Gaps = 5/85 (5%)
Frame = +2
Query: 7 LTLLDSFWFETTILTNKNPS-----KVEEILPQDTNLLLVPTPNLDVRFYSEQNLSIIGS 61
LT L ++ K P K E L NL P P F S QNLS+ G+
Sbjct: 95 LTKLVKLSMSNNFMSGKLPDNAADFKSLEFLDISNNLFSSPLPPEIGNFGSLQNLSLAGN 274
Query: 62 VFSDSPSPNSVLTSSKLRTIPSERE 86
FS PNS+ + ++++ R+
Sbjct: 275 NFS-GRIPNSISDMASIKSLDLSRK 346
>TC16720 homologue to UP|Q9XH46 (Q9XH46) THA4 protein precursor, partial
(42%)
Length = 567
Score = 28.5 bits (62), Expect = 1.1
Identities = 12/39 (30%), Positives = 24/39 (60%), Gaps = 2/39 (5%)
Frame = -2
Query: 161 QRLGRENDDAEEGEDEE--HKKIDENVLSDNKPYLSEAW 197
Q LG +ND + G+D+E + + + V S++ ++S +W
Sbjct: 281 QLLGSKNDGGDAGDDDEFGNAEAKQGVASESSTFVSPSW 165
>AV414862
Length = 406
Score = 28.5 bits (62), Expect = 1.1
Identities = 19/63 (30%), Positives = 31/63 (49%)
Frame = +1
Query: 119 RRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDEEH 178
+R + R L+ F KG F +D+D +L L + R+NDD +EG DE+
Sbjct: 115 QRERMRLLTTAAFDRGKG----EDTFGAKDEDWQLYKL------MSRDNDDDDEGPDEDD 264
Query: 179 KKI 181
++
Sbjct: 265 TEL 273
>TC16133 similar to UP|Q39449 (Q39449) Specific tissue protein 1, partial
(40%)
Length = 934
Score = 27.7 bits (60), Expect = 1.8
Identities = 24/81 (29%), Positives = 42/81 (51%)
Frame = +3
Query: 149 KDSKLVSLIPGLQRLGRENDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRERKNP 208
KD ++ I GL +L E + G++ EHK +E+V+++N+ Y+ E VF + P
Sbjct: 141 KDEEMPEGIQGLLQLKSE---IKPGKNSEHKCDEEHVVTNNE-YIIEK-KVFTEELEPRP 305
Query: 209 LVNWRVPDKGSEIDMKDNLKF 229
++ D +D K N K+
Sbjct: 306 NISAYDDD---HVDTKANKKY 359
>TC11700 homologue to GB|AAL06971.1|15810091|AY056083 AT4g30260/F9N11_110
{Arabidopsis thaliana;}, partial (45%)
Length = 561
Score = 27.7 bits (60), Expect = 1.8
Identities = 26/77 (33%), Positives = 35/77 (44%), Gaps = 6/77 (7%)
Frame = +3
Query: 25 PSKVEEILPQD--TNLLLV----PTPNLDVRFYSEQNLSIIGSVFSDSPSPNSVLTSSKL 78
PS +LP T+LLL P+PNL+ QN S+ + P P S +S
Sbjct: 159 PSPSPPLLPSSIPTSLLLFLPNPPSPNLNF-----QNPSLSLLLLLPPPDPKSPPPASDP 323
Query: 79 RTIPSEREFGEFSTGTS 95
IPS +FG S+ S
Sbjct: 324 LLIPSPSQFGTPSSAIS 374
>AV778536
Length = 550
Score = 26.6 bits (57), Expect = 4.1
Identities = 19/49 (38%), Positives = 25/49 (50%)
Frame = -2
Query: 82 PSEREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELE 130
P ER G G SSE H +E+ K++ S + R R +SRS S E
Sbjct: 153 PVERHRG----GRSSERHE*EEEELQRKREISITPWRSRSRSRSRSRSE 19
>TC15506 similar to UP|Q9SEE9 (Q9SEE9) Arginine/serine-rich protein, partial
(24%)
Length = 677
Score = 26.6 bits (57), Expect = 4.1
Identities = 15/38 (39%), Positives = 20/38 (52%)
Frame = +2
Query: 61 SVFSDSPSPNSVLTSSKLRTIPSEREFGEFSTGTSSEN 98
S FSDSPSP V S+ R+ G S+ +SS +
Sbjct: 221 SSFSDSPSPRKVSRRSRSRSPRRPLRGGRASSNSSSSS 334
>BP035374
Length = 435
Score = 26.2 bits (56), Expect = 5.4
Identities = 15/38 (39%), Positives = 20/38 (52%)
Frame = -1
Query: 61 SVFSDSPSPNSVLTSSKLRTIPSEREFGEFSTGTSSEN 98
S FSDSPSP V S+ R+ G S+ +SS +
Sbjct: 378 SSFSDSPSPRKVSRRSRSRSPRRPLRGGRASSDSSSSS 265
>TC11308 similar to UP|CENB_HUMAN (P07199) Major centromere autoantigen B
(Centromere protein B) (CENP-B), partial (4%)
Length = 756
Score = 25.8 bits (55), Expect = 7.0
Identities = 10/21 (47%), Positives = 16/21 (75%)
Frame = -2
Query: 164 GRENDDAEEGEDEEHKKIDEN 184
G E +AEEGE++E ++ DE+
Sbjct: 302 GEEEVEAEEGEEDEEEEDDED 240
>TC10270 homologue to UP|Q8VXT0 (Q8VXT0) Translation initiation factor
(eIF-1A), partial (98%)
Length = 619
Score = 25.4 bits (54), Expect = 9.2
Identities = 16/51 (31%), Positives = 28/51 (54%)
Frame = +3
Query: 75 SSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKSRS 125
S++ RTI + F+T T ++ HH +E ++ Q + R RRR+ R+
Sbjct: 36 STQSRTIATRES--RFTTTTVTD-HHAEEQGKGREESQERKERSRRREERA 179
>TC11879 weakly similar to UP|OREX_HUMAN (O43612) Orexin precursor
(Hypocretin) (Hcrt) [Contains: Orexin-A (Hypocretin-1)
(Hcrt1); Orexin-B (Hypocretin-2) (Hcrt2)], partial (17%)
Length = 683
Score = 25.4 bits (54), Expect = 9.2
Identities = 11/28 (39%), Positives = 16/28 (56%)
Frame = -1
Query: 98 NHHQKEDVAYTKKKQSQSYRRRRRKSRS 125
NH + +Y ++K + YRRR R RS
Sbjct: 518 NHTWGKRKSYLREKMQRRYRRRNRVDRS 435
>TC15478 homologue to UP|AAQ08016 (AAQ08016) Expansin, partial (33%)
Length = 629
Score = 25.4 bits (54), Expect = 9.2
Identities = 12/41 (29%), Positives = 21/41 (50%)
Frame = +1
Query: 53 EQNLSIIGSVFSDSPSPNSVLTSSKLRTIPSEREFGEFSTG 93
+ N ++G S + + TS+ L +PS +FG+ TG
Sbjct: 118 QSNSVLVGQSLSFRVTASDRRTSTSLNIVPSHWQFGQTFTG 240
>BP038141
Length = 493
Score = 25.4 bits (54), Expect = 9.2
Identities = 11/28 (39%), Positives = 17/28 (60%)
Frame = +3
Query: 98 NHHQKEDVAYTKKKQSQSYRRRRRKSRS 125
NHH + + + S++ RRRRR+S S
Sbjct: 3 NHHHEGEKLQLQSLTSKALRRRRRRSPS 86
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.130 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,876,437
Number of Sequences: 28460
Number of extensions: 50128
Number of successful extensions: 305
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 296
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 302
length of query: 239
length of database: 4,897,600
effective HSP length: 87
effective length of query: 152
effective length of database: 2,421,580
effective search space: 368080160
effective search space used: 368080160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC135160.3


