

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135160.2 + phase: 0 /pseudo
(319 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
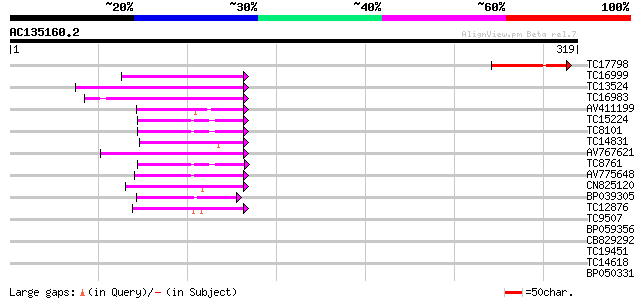
Sequences producing significant alignments: (bits) Value
TC17798 similar to GB|AAM19907.1|20453337|AY097391 At2g42750/F7D... 67 4e-12
TC16999 similar to UP|Q9T024 (Q9T024) DNAJ-like protein (AT4g391... 59 1e-09
TC13524 similar to UP|O48678 (O48678) F3I6.4 protein, partial (19%) 57 5e-09
TC16983 similar to UP|Q9ZSY2 (Q9ZSY2) ALTERED response to gravit... 55 1e-08
AV411199 51 3e-07
TC15224 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein ... 49 1e-06
TC8101 similar to UP|O24074 (O24074) DNAJ-like protein, partial ... 49 1e-06
TC14831 homologue to UP|P93499 (P93499) DnaJ-like protein (Fragm... 48 3e-06
AV767621 47 3e-06
TC8761 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein P... 47 3e-06
AV775648 47 6e-06
CN825120 44 3e-05
BP039305 43 8e-05
TC12876 42 1e-04
TC9507 similar to UP|Q8VZB6 (Q8VZB6) DnaJ-like protein, partial ... 37 0.003
BP059356 29 0.008
CB829292 30 0.41
TC19451 similar to GB|AAM65667.1|21593700|AY088122 J8-like prote... 30 0.54
TC14618 homologue to UP|AAQ76040 (AAQ76040) Ubiquitin extension ... 29 0.92
BP050331 29 1.2
>TC17798 similar to GB|AAM19907.1|20453337|AY097391 At2g42750/F7D19.25
{Arabidopsis thaliana;}, partial (29%)
Length = 548
Score = 67.0 bits (162), Expect = 4e-12
Identities = 34/45 (75%), Positives = 37/45 (81%)
Frame = +1
Query: 272 DYENSEEEVKERAKRAAAAARKWREYSRKGGAVKPSTVKLPEATS 316
DY+ SE+EVKERAKRAAAAAR+WREYSRK G KP T LPEA S
Sbjct: 178 DYQRSEDEVKERAKRAAAAARRWREYSRK-GVDKPPTFTLPEAAS 309
>TC16999 similar to UP|Q9T024 (Q9T024) DNAJ-like protein
(AT4g39150/T22F8_50), partial (35%)
Length = 648
Score = 58.9 bits (141), Expect = 1e-09
Identities = 29/71 (40%), Positives = 39/71 (54%)
Frame = +3
Query: 64 TSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEV 123
+ + V YY VLG+ DA+ IKKAYY + HPD + DP+ + E Y+V
Sbjct: 102 SQEMVKESAYYDVLGVNVDASAADIKKAYYIKARIVHPDKNPGDPKAAENFQMLGEAYQV 281
Query: 124 LSDPVQRRVYD 134
LSDP +R YD
Sbjct: 282 LSDPEKREAYD 314
>TC13524 similar to UP|O48678 (O48678) F3I6.4 protein, partial (19%)
Length = 485
Score = 56.6 bits (135), Expect = 5e-09
Identities = 31/97 (31%), Positives = 48/97 (48%)
Frame = +1
Query: 38 WRFATPSCKRRGFGRVRVATEQESFSTSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMK 97
W+ ++ RR F ++ E +D+ D Y VLG+ +T ++IK AY
Sbjct: 190 WKLSSRVS*RRRFKMPGPRSKSEKKHDADNQLRRDPYEVLGVSRSSTDQEIKTAYRKMAL 369
Query: 98 TCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRRVYD 134
HPD + NDP+ + + Y +LSDP +RR YD
Sbjct: 370 KYHPDKNANDPKAADMFKEVTFSYNILSDPDKRRQYD 480
>TC16983 similar to UP|Q9ZSY2 (Q9ZSY2) ALTERED response to gravity (ARG1
protein), partial (42%)
Length = 716
Score = 55.5 bits (132), Expect = 1e-08
Identities = 34/92 (36%), Positives = 45/92 (47%)
Frame = +2
Query: 43 PSCKRRGFGRVRVATEQESFSTSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPD 102
P C RR G A + TS V D Y VL + D+T ++IK AY HPD
Sbjct: 161 PRCFRRERG---AAMGSKMEGTSSPVIRRDPYEVLSVSRDSTDQEIKTAYRKLALKYHPD 331
Query: 103 LSGNDPETTNFCTFINEVYEVLSDPVQRRVYD 134
+ N+PE + + Y +LSDP +RR YD
Sbjct: 332 KNVNNPEASELFKEVAYSYSILSDPEKRRQYD 427
>AV411199
Length = 423
Score = 50.8 bits (120), Expect = 3e-07
Identities = 30/66 (45%), Positives = 38/66 (57%), Gaps = 3/66 (4%)
Frame = +2
Query: 72 DYYAVLGLLPDATPEQIKKAYYNCMKTCHPDL---SGNDPETTNFCTFINEVYEVLSDPV 128
DYY +L + AT E++KKAY HPD + ND ET I+E YEVLSDP
Sbjct: 68 DYYEILEVDHHATDEELKKAYRRLAMKWHPDKNPDNKNDAETK--FKLISESYEVLSDPQ 241
Query: 129 QRRVYD 134
+R +YD
Sbjct: 242 KRAIYD 259
>TC15224 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
partial (71%)
Length = 1031
Score = 48.9 bits (115), Expect = 1e-06
Identities = 26/62 (41%), Positives = 36/62 (57%)
Frame = +3
Query: 73 YYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRRV 132
YY +LG+ +A+ + +KKAY HPD G DPE + + YEVLSDP +R +
Sbjct: 177 YYEILGVPKNASQDDLKKAYKKAAIKNHPD-KGGDPEKFK---ELAQAYEVLSDPEKREI 344
Query: 133 YD 134
YD
Sbjct: 345 YD 350
>TC8101 similar to UP|O24074 (O24074) DNAJ-like protein, partial (90%)
Length = 1737
Score = 48.5 bits (114), Expect = 1e-06
Identities = 26/62 (41%), Positives = 36/62 (57%)
Frame = +1
Query: 73 YYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRRV 132
+Y VLG+ A+ ++IKKAY HPD G DPE + + YEVLSDP ++ +
Sbjct: 187 FYEVLGVPKSASEDEIKKAYRKAAMKNHPD-KGGDPEKFK---ELGQAYEVLSDPEKKEL 354
Query: 133 YD 134
YD
Sbjct: 355 YD 360
>TC14831 homologue to UP|P93499 (P93499) DnaJ-like protein (Fragment),
partial (63%)
Length = 851
Score = 47.8 bits (112), Expect = 3e-06
Identities = 23/63 (36%), Positives = 36/63 (56%), Gaps = 2/63 (3%)
Frame = +3
Query: 74 YAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTF--INEVYEVLSDPVQRR 131
Y +LG+ A+ ++IK AY + CHPD++ D + ++ F I+ Y LSDP +R
Sbjct: 249 YEILGIPAGASSQEIKAAYRRLARVCHPDVAAIDRKNSSADDFMKIHAAYSTLSDPEKRA 428
Query: 132 VYD 134
YD
Sbjct: 429 TYD 437
>AV767621
Length = 560
Score = 47.4 bits (111), Expect = 3e-06
Identities = 28/84 (33%), Positives = 42/84 (49%), Gaps = 1/84 (1%)
Frame = +1
Query: 52 RVRVATEQESFSTSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLS-GNDPET 110
R + E+++ + DYY +L + A+ + +KKAY HPD + N E
Sbjct: 1 RTKRRREEKTGEGEEEAMGVDYYKILQVDRGASDDDLKKAYRKLAMKWHPDKNPNNKKEA 180
Query: 111 TNFCTFINEVYEVLSDPVQRRVYD 134
I+E Y+VLSDP +R VYD
Sbjct: 181 EAKFKQISEAYDVLSDPQKRAVYD 252
>TC8761 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
partial (74%)
Length = 1050
Score = 47.4 bits (111), Expect = 3e-06
Identities = 25/63 (39%), Positives = 36/63 (56%)
Frame = +3
Query: 73 YYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRRV 132
YY +LG+ A+ + +KKAY HPD G DPE + + YEVL+DP +R +
Sbjct: 120 YYDILGVSKTASQDDLKKAYKKAAIKNHPD-KGGDPEKFK---ELAQAYEVLNDPEKREI 287
Query: 133 YDD 135
YD+
Sbjct: 288 YDN 296
>AV775648
Length = 371
Score = 46.6 bits (109), Expect = 6e-06
Identities = 23/64 (35%), Positives = 33/64 (50%)
Frame = +1
Query: 71 EDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQR 130
+D Y ++G+ +A +IKKAYY HPD + DPE+ + YE+L D R
Sbjct: 136 DDCYDLVGVSQNANSSEIKKAYYKLSLKHHPD-TNPDPESKKLLVKVANAYEILKDEATR 312
Query: 131 RVYD 134
YD
Sbjct: 313 EQYD 324
>CN825120
Length = 667
Score = 44.3 bits (103), Expect = 3e-05
Identities = 24/71 (33%), Positives = 35/71 (48%), Gaps = 2/71 (2%)
Frame = +3
Query: 66 DSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGND--PETTNFCTFINEVYEV 123
D Y +LG+ A+ ++IK AY + HPD++ D +TN I+ Y
Sbjct: 186 DMASCTSLYEILGIAAAASDQEIKAAYRRLARVRHPDVAAVDRKDSSTNEFMKIHAAYST 365
Query: 124 LSDPVQRRVYD 134
LSDP +R YD
Sbjct: 366 LSDPEKRASYD 398
>BP039305
Length = 555
Score = 42.7 bits (99), Expect = 8e-05
Identities = 24/59 (40%), Positives = 33/59 (55%)
Frame = +3
Query: 72 DYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQR 130
DYY LG+ AT ++IK AY + HPD+ +P T+ I+ YEVLSD +R
Sbjct: 381 DYYNTLGVPKSATVKEIKAAYRRLARQYHPDVY-KEPGATDKFKEISNAYEVLSDDKKR 554
>TC12876
Length = 820
Score = 42.0 bits (97), Expect = 1e-04
Identities = 27/71 (38%), Positives = 35/71 (49%), Gaps = 6/71 (8%)
Frame = -1
Query: 70 AEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPD---LSGN---DPETTNFCTFINEVYEV 123
+ D+YAVLGL + T +++ AY HPD SGN E I E Y V
Sbjct: 241 SNDFYAVLGLNKECTESELRNAYKKLALKWHPDRCSASGNLKFVEEAKKKFQSIQEAYSV 62
Query: 124 LSDPVQRRVYD 134
LSD +R +YD
Sbjct: 61 LSDANKRLMYD 29
>TC9507 similar to UP|Q8VZB6 (Q8VZB6) DnaJ-like protein, partial (40%)
Length = 905
Score = 37.4 bits (85), Expect = 0.003
Identities = 26/106 (24%), Positives = 41/106 (38%), Gaps = 14/106 (13%)
Frame = +1
Query: 43 PSCKRRGFGRVRVATEQESFSTSDSVG--------------AEDYYAVLGLLPDATPEQI 88
PS +RR A S S+S S ++YY +LGL T E +
Sbjct: 388 PSVRRRSSASAAAAPVPPSSSSSSSASYTTEQVSIIREIKRKKNYYEILGLEKSCTVEDV 567
Query: 89 KKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRRVYD 134
+K+Y HPD P +++ ++ L + +R YD
Sbjct: 568 RKSYRKLSLKVHPD-KNKAPGAEEAFKAVSKAFQCLGNEESKRKYD 702
>BP059356
Length = 436
Score = 29.3 bits (64), Expect(2) = 0.008
Identities = 12/18 (66%), Positives = 15/18 (82%)
Frame = +1
Query: 117 INEVYEVLSDPVQRRVYD 134
I+E Y+VLSDP +R VYD
Sbjct: 232 ISEAYDVLSDPQKRAVYD 285
Score = 25.8 bits (55), Expect(2) = 0.008
Identities = 13/37 (35%), Positives = 17/37 (45%)
Frame = +3
Query: 65 SDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHP 101
S+ DYY VL + A + +KKAY HP
Sbjct: 72 SEKAMGVDYYKVLQVDRSAKDDDLKKAYRKLAMKWHP 182
>CB829292
Length = 544
Score = 30.4 bits (67), Expect = 0.41
Identities = 21/64 (32%), Positives = 30/64 (46%), Gaps = 4/64 (6%)
Frame = +1
Query: 65 SDSVGAEDYYAVLGLLPDA--TPEQIKKAYYNCMKTCHPDL--SGNDPETTNFCTFINEV 120
+ + A D +LG P++ TP Q+K AY + HPDL S P + I+E
Sbjct: 67 ASQMDANDARILLGFPPNSHPTPSQVKSAYKKKVWESHPDLFPSHEKPLAESKFKLISEA 246
Query: 121 YEVL 124
Y L
Sbjct: 247 YACL 258
>TC19451 similar to GB|AAM65667.1|21593700|AY088122 J8-like protein
{Arabidopsis thaliana;}, partial (35%)
Length = 480
Score = 30.0 bits (66), Expect = 0.54
Identities = 17/53 (32%), Positives = 26/53 (48%)
Frame = +2
Query: 72 DYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVL 124
D Y L + P A+ ++KKA+ + HPD+ F INE Y+V+
Sbjct: 227 DSYKTLRVQPGASQTEVKKAFRHLALQYHPDVCQGSKCGVQF-HHINEAYDVV 382
>TC14618 homologue to UP|AAQ76040 (AAQ76040) Ubiquitin extension protein,
partial (97%)
Length = 644
Score = 29.3 bits (64), Expect = 0.92
Identities = 22/57 (38%), Positives = 27/57 (46%), Gaps = 11/57 (19%)
Frame = -2
Query: 18 SSLCTRSSWQMLQQKTAVSPWRFA---------TPS--CKRRGFGRVRVATEQESFS 63
S C SSW + ++ S WR TPS C RRG GR+RVA + S S
Sbjct: 319 SCACA*SSWA*CRSSSS-SSWRHRGDGGRGGEWTPSGCCSRRGCGRLRVAFRRRSTS 152
>BP050331
Length = 519
Score = 28.9 bits (63), Expect = 1.2
Identities = 21/76 (27%), Positives = 36/76 (46%), Gaps = 4/76 (5%)
Frame = +3
Query: 63 STSDSVGAEDYYAVLGLLPDATPEQ----IKKAYYNCMKTCHPDLSGNDPETTNFCTFIN 118
+T+++ D+YA+L + D Q IK+ Y HPD + F ++
Sbjct: 240 TTTNNRNNLDFYAILQV--DHRDSQDLNLIKRQYRRLALLLHPDKNLFSFSHHTF-DLVS 410
Query: 119 EVYEVLSDPVQRRVYD 134
+ +LSDP Q+ +YD
Sbjct: 411 HAWALLSDPAQKEIYD 458
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.135 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,708,813
Number of Sequences: 28460
Number of extensions: 76442
Number of successful extensions: 459
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 448
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 449
length of query: 319
length of database: 4,897,600
effective HSP length: 90
effective length of query: 229
effective length of database: 2,336,200
effective search space: 534989800
effective search space used: 534989800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC135160.2


