

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135100.8 - phase: 0
(420 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
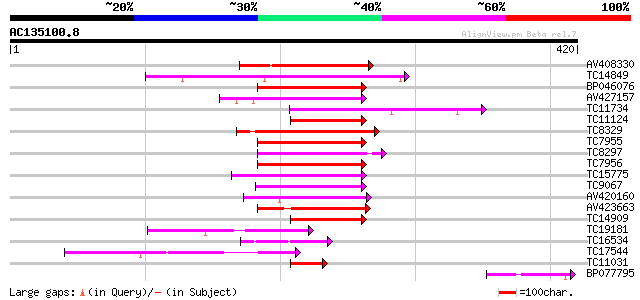
Sequences producing significant alignments: (bits) Value
AV408330 140 4e-34
TC14849 similar to UP|Q40090 (Q40090) SPF1 protein, partial (37%) 113 5e-26
BP046076 111 3e-25
AV427157 100 8e-22
TC11734 similar to UP|WR49_ARATH (Q9FHR7) Probable WRKY transcri... 87 5e-18
TC11124 similar to UP|Q8H9D7 (Q8H9D7) WRKY-type DNA binding prot... 85 3e-17
TC8329 similar to UP|BAD06717 (BAD06717) WRKY transcription fact... 82 1e-16
TC7955 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcrip... 82 1e-16
TC8297 similar to UP|Q40828 (Q40828) WRKY3, partial (27%) 80 8e-16
TC7956 similar to GB|AAM61419.1|21537078|AY084855 putaive DNA-bi... 79 1e-15
TC15775 similar to UP|WR17_ARATH (Q9SJA8) Probable WRKY transcri... 77 4e-15
TC9067 similar to UP|WR15_ARATH (O22176) Probable WRKY transcrip... 75 3e-14
AV420160 75 3e-14
AV423663 72 2e-13
TC14909 similar to UP|WR21_ARATH (O04336) Probable WRKY transcri... 70 6e-13
TC19181 similar to UP|Q40090 (Q40090) SPF1 protein, partial (19%) 65 2e-11
TC16534 similar to UP|WR41_ARATH (Q8H0Y8) Probable WRKY transcri... 57 6e-09
TC17544 similar to PIR|T04919|T04919 DNA-binding protein homolog... 54 6e-08
TC11031 homologue to UP|WR65_ARATH (Q9LP56) Probable WRKY transc... 52 2e-07
BP077795 50 7e-07
>AV408330
Length = 316
Score = 140 bits (353), Expect = 4e-34
Identities = 64/99 (64%), Positives = 79/99 (79%)
Frame = +3
Query: 171 NDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSY 230
++ ++ KKQLK KK N+ ++ P R F TKS+VDHL+DGYRWRKYGQK VKNSPFPRSY
Sbjct: 21 DEHNKTKKQLKAKKTNQKRQREP-RFAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSY 197
Query: 231 YRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLSPVL 269
YRCT+ +C VKKR+ERS D SIV+T+YEG H H SPV+
Sbjct: 198 YRCTSASCNVKKRVERSFTDPSIVVTTYEGQHTHSSPVM 314
>TC14849 similar to UP|Q40090 (Q40090) SPF1 protein, partial (37%)
Length = 1389
Score = 113 bits (283), Expect = 5e-26
Identities = 68/212 (32%), Positives = 110/212 (51%), Gaps = 16/212 (7%)
Frame = +2
Query: 101 PAPSPAASEVLNNPATPVSVSSSISS-------SSNAAGVADKVMVEENEHEHEEELEGD 153
P P ++ + P+ S+ IS+ +++ +G D V EN + + +
Sbjct: 56 PKPQATRRNSSSSSSLPIPHSNHISNDIQDQSYATHGSGQMDSVATPENSSISIGDDDFE 235
Query: 154 EAEEKDEDHGIRLVENQNDQDQMKKQLKPKKKNK--NKKLRPARVTFKTKSDVDHLDDGY 211
++ +K G L E++ D + K + + + + ++ +R RV +T SD+D LDDGY
Sbjct: 236 QSSQKCRSGGDELDEDEPDSKRWKVEGENEGISAPGSRTVREPRVVVQTTSDIDILDDGY 415
Query: 212 RWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLSPVLLR 271
RWRKYGQK VK +P PRSYY+CT C V+K +ER++ D V+T+YEG H H P
Sbjct: 416 RWRKYGQKVVKGNPNPRSYYKCTHPGCPVRKHVERASHDLRAVITTYEGKHNHDVPAARG 595
Query: 272 AANLGIMSD-PSGFQEQV------HQHHNTTS 296
+ N + P+ + HQHH+ T+
Sbjct: 596 SGNHSVTRPLPNNTANAIRPSSVTHQHHHNTN 691
>BP046076
Length = 593
Score = 111 bits (277), Expect = 3e-25
Identities = 51/81 (62%), Positives = 60/81 (73%)
Frame = -2
Query: 184 KKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKR 243
K +KLR R F+T+SDVD LDDGY+WRKYGQK VKNS PRSYYRCT NC VKKR
Sbjct: 529 KVKVRRKLREPRFCFQTRSDVDVLDDGYKWRKYGQKVVKNSLHPRSYYRCTHSNCRVKKR 350
Query: 244 IERSAADSSIVLTSYEGHHIH 264
+ER + D +V+T+YEG H H
Sbjct: 349 VERLSEDCRMVITTYEGRHNH 287
>AV427157
Length = 379
Score = 99.8 bits (247), Expect = 8e-22
Identities = 49/118 (41%), Positives = 71/118 (59%), Gaps = 9/118 (7%)
Frame = +3
Query: 156 EEKDEDHGIRL-----VENQNDQDQMKKQ----LKPKKKNKNKKLRPARVTFKTKSDVDH 206
EE+++D G + + D+ + K++ + + +R RV +T S+VD
Sbjct: 24 EEEEDDQGTHGSVSLGYDGEGDESESKRRKLESYATELSGATRAIREPRVVVQTTSEVDI 203
Query: 207 LDDGYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIH 264
LDDGYRWRKYGQK VK +P PRSYY+CT C V+K +ER++ D V+T+YEG H H
Sbjct: 204 LDDGYRWRKYGQKVVKGNPNPRSYYKCTNAGCMVRKHVERASHDLKSVITTYEGKHNH 377
>TC11734 similar to UP|WR49_ARATH (Q9FHR7) Probable WRKY transcription
factor 49 (WRKY DNA-binding protein 49), partial (28%)
Length = 1009
Score = 87.0 bits (214), Expect = 5e-18
Identities = 56/155 (36%), Positives = 72/155 (46%), Gaps = 9/155 (5%)
Frame = +3
Query: 208 DDGYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLS- 266
DDGY+WRKYGQK +KNSP PRSYYRCT C KK++ERS D ++ +YEG H+H +
Sbjct: 333 DDGYKWRKYGQKSIKNSPNPRSYYRCTNPRCSAKKQVERSNEDPETLIITYEGLHLHFAY 512
Query: 267 PVLLRAANLGIMSDP----SGF-QEQVHQHHNTTSGVVGSSVLGNEFAMSQSQQYQKFQD 321
P L S P S F QV +V + ++ QD
Sbjct: 513 PCFLVGQPQQSQSHPPVKKSKFTSSQVQAQAQAQDFIVNEAQASASVGLTSPTLLNSPQD 692
Query: 322 QNQNVLHHQ---RQAVPSLLYNSDYNNSFNISPLS 353
Q L Q VP ++ N N + S LS
Sbjct: 693 MAQQNLGSQGLLEDMVPFMVRNPSNNLNSGYSKLS 797
>TC11124 similar to UP|Q8H9D7 (Q8H9D7) WRKY-type DNA binding protein,
partial (45%)
Length = 517
Score = 84.7 bits (208), Expect = 3e-17
Identities = 36/56 (64%), Positives = 41/56 (72%)
Frame = +1
Query: 209 DGYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIH 264
DGYRWRKYGQK VKN+ FPRSYYRCT C VKK + R D +V+T+YEG H H
Sbjct: 1 DGYRWRKYGQKAVKNNKFPRSYYRCTHHGCHVKKPVPRLTQDEGVVVTTYEGVHTH 168
>TC8329 similar to UP|BAD06717 (BAD06717) WRKY transcription factor 1,
partial (41%)
Length = 1717
Score = 82.4 bits (202), Expect = 1e-16
Identities = 39/107 (36%), Positives = 65/107 (60%), Gaps = 1/107 (0%)
Frame = +1
Query: 169 NQNDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPR 228
+ D+D KK P+++ K+ A V + + DGY+WRKYGQK +++P PR
Sbjct: 475 SSTDEDSFKK---PREETIKAKISRAYVRTEATDTGFVVKDGYQWRKYGQKVTRDNPCPR 645
Query: 229 SYYRCT-AGNCEVKKRIERSAADSSIVLTSYEGHHIHLSPVLLRAAN 274
+Y++C+ A +C VKK+++RS D S+++ +YEG H H P + A +
Sbjct: 646 AYFKCSFAPSCPVKKKVQRSVDDQSVLVATYEGEHNHPQPSQMEATS 786
>TC7955 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcription
factor 11 (WRKY DNA-binding protein 11), partial (27%)
Length = 579
Score = 82.4 bits (202), Expect = 1e-16
Identities = 36/82 (43%), Positives = 50/82 (60%), Gaps = 1/82 (1%)
Frame = +3
Query: 184 KKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYYRC-TAGNCEVKK 242
KK KN+ + RV + D D Y WRKYGQKP+K SP+PR YY+C T C +K
Sbjct: 18 KKRKNRVKKTVRVPAISSKTADIPPDEYSWRKYGQKPIKGSPYPRGYYKCSTVRGCPARK 197
Query: 243 RIERSAADSSIVLTSYEGHHIH 264
+ER+ D ++++ +YEG H H
Sbjct: 198 HVERATDDPTMLIVTYEGEHRH 263
>TC8297 similar to UP|Q40828 (Q40828) WRKY3, partial (27%)
Length = 617
Score = 79.7 bits (195), Expect = 8e-16
Identities = 38/97 (39%), Positives = 58/97 (59%), Gaps = 1/97 (1%)
Frame = +1
Query: 184 KKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYYRCTA-GNCEVKK 242
KK K + R RV + D D ++WRKYGQKP+K SP+PR YY+C++ C +K
Sbjct: 97 KKKKLRVKRTIRVPAISSKTADIPTDDFQWRKYGQKPIKGSPYPRGYYKCSSEKGCPARK 276
Query: 243 RIERSAADSSIVLTSYEGHHIHLSPVLLRAANLGIMS 279
+ER+ D ++++ +YEG H HL + A +G M+
Sbjct: 277 HVERAQDDPNMLIVTYEGEHRHL---ITTTAAVGFMA 378
>TC7956 similar to GB|AAM61419.1|21537078|AY084855 putaive DNA-binding
protein {Arabidopsis thaliana;} , partial (45%)
Length = 1257
Score = 79.0 bits (193), Expect = 1e-15
Identities = 35/82 (42%), Positives = 50/82 (60%), Gaps = 1/82 (1%)
Frame = +3
Query: 184 KKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYYRC-TAGNCEVKK 242
KK K++ + RV + D D Y WRKYGQKP+K SP+PR YY+C T C +K
Sbjct: 579 KKRKSRVKQTIRVPAVSSKIADIPPDEYSWRKYGQKPIKGSPYPRGYYKCSTVRGCPARK 758
Query: 243 RIERSAADSSIVLTSYEGHHIH 264
+ER+ D ++++ +YEG H H
Sbjct: 759 HVERAQDDPNMLIVTYEGEHRH 824
>TC15775 similar to UP|WR17_ARATH (Q9SJA8) Probable WRKY transcription
factor 17 (WRKY DNA-binding protein 17), partial (53%)
Length = 1284
Score = 77.4 bits (189), Expect = 4e-15
Identities = 36/101 (35%), Positives = 54/101 (52%), Gaps = 1/101 (0%)
Frame = +3
Query: 165 RLVENQNDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNS 224
R E+ D + K+ KN+ + RV + D D Y WRKYGQKP+K +
Sbjct: 609 RCHEHSGDPSGSNTKCHCTKRRKNRVKKTIRVPAVSSKVADIPADEYSWRKYGQKPIKGT 788
Query: 225 PFPRSYYRC-TAGNCEVKKRIERSAADSSIVLTSYEGHHIH 264
P+PR YY+C T C +K +ER+ D ++++ +YE H H
Sbjct: 789 PYPRGYYKCSTVRGCPARKHVERATDDPAMLIVTYEDEHDH 911
>TC9067 similar to UP|WR15_ARATH (O22176) Probable WRKY transcription
factor 15 (WRKY DNA-binding protein 15), partial (31%)
Length = 660
Score = 74.7 bits (182), Expect = 3e-14
Identities = 34/83 (40%), Positives = 50/83 (59%), Gaps = 1/83 (1%)
Frame = +3
Query: 183 KKKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYYRCTA-GNCEVK 241
+K K + R RV + D D Y WRKYGQKP+K SP PR YY+C++ C +
Sbjct: 48 QKSRKMRLKRVVRVPAISLKMADIPPDDYSWRKYGQKPIKGSPHPRGYYKCSSVRGCPAR 227
Query: 242 KRIERSAADSSIVLTSYEGHHIH 264
K +ER+ D+++++ +YEG H H
Sbjct: 228 KHVERALDDAAMLVVTYEGEHNH 296
>AV420160
Length = 417
Score = 74.7 bits (182), Expect = 3e-14
Identities = 36/102 (35%), Positives = 55/102 (53%), Gaps = 7/102 (6%)
Frame = +1
Query: 174 DQMKKQLKPKKKNKNKKLRPARVTF------KTKSDVDHLDDGYRWRKYGQKPVKNSPFP 227
+ +K + KK + K R + K+K + D + WRKYGQKP+K SP+P
Sbjct: 34 EDIKTEAPSPKKRREMKKRVVTLPIGDVDGSKSKGETYPPSDSWAWRKYGQKPIKGSPYP 213
Query: 228 RSYYRCTAG-NCEVKKRIERSAADSSIVLTSYEGHHIHLSPV 268
R YYRC++ C +K++ERS D + ++ +Y H H PV
Sbjct: 214 RGYYRCSSSKGCPARKQVERSRVDPTKLMVTYSYEHNHSLPV 339
>AV423663
Length = 447
Score = 71.6 bits (174), Expect = 2e-13
Identities = 32/85 (37%), Positives = 54/85 (62%), Gaps = 1/85 (1%)
Frame = +1
Query: 184 KKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYYRCTAG-NCEVKK 242
K+ KN+ + +V+ + S D + WRKYGQKP+K SP+PR YYRC++ C +K
Sbjct: 22 KRRKNQFKKVCQVSAENLSS-----DIWAWRKYGQKPIKGSPYPRGYYRCSSSKGCLARK 186
Query: 243 RIERSAADSSIVLTSYEGHHIHLSP 267
++ER+ ++ ++ + +Y G H H +P
Sbjct: 187 QVERNKSEPAMFIVTYTGEHNHPAP 261
>TC14909 similar to UP|WR21_ARATH (O04336) Probable WRKY transcription
factor 21 (WRKY DNA-binding protein 21), partial (22%)
Length = 590
Score = 70.1 bits (170), Expect = 6e-13
Identities = 28/57 (49%), Positives = 39/57 (68%), Gaps = 1/57 (1%)
Frame = +3
Query: 209 DGYRWRKYGQKPVKNSPFPRSYYRCTA-GNCEVKKRIERSAADSSIVLTSYEGHHIH 264
D Y WRKYGQKP+K SP PR YY+C++ C +K +ER + +++L +YEG H H
Sbjct: 63 DDYSWRKYGQKPIKGSPHPRGYYKCSSMRGCPARKHVERCLEEPTMLLVTYEGEHNH 233
>TC19181 similar to UP|Q40090 (Q40090) SPF1 protein, partial (19%)
Length = 402
Score = 65.5 bits (158), Expect = 2e-11
Identities = 42/133 (31%), Positives = 65/133 (48%), Gaps = 10/133 (7%)
Frame = +3
Query: 103 PSPAASEVLNNPATPVS-VSSSISSSSNAAGVADKVMVEENE---------HEHEEELEG 152
P P A N+ T + VS+ I +++ +G D V EN + G
Sbjct: 24 PKPQAPHKRNSSFTTSNHVSTEIQGATHVSGQMDSVATPENSSLSMGGDDFEQSSMSKSG 203
Query: 153 DEAEEKDEDHGIRLVENQNDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYR 212
E ++DE HG + +M+ + + ++ ++ +RV +T SD+D LDDGYR
Sbjct: 204 GEELDEDEPHGKKW--------RMEGENEGMSAAGSRTVKESRVVVQTTSDIDILDDGYR 359
Query: 213 WRKYGQKPVKNSP 225
WRKYGQK VK +P
Sbjct: 360 WRKYGQKVVKGNP 398
>TC16534 similar to UP|WR41_ARATH (Q8H0Y8) Probable WRKY transcription
factor 41 (WRKY DNA-binding protein 41), partial (20%)
Length = 697
Score = 57.0 bits (136), Expect = 6e-09
Identities = 29/68 (42%), Positives = 39/68 (56%)
Frame = +1
Query: 172 DQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYY 231
D + K+ K K+K K + RV + + H DDGY WRKYGQK + + +PRSYY
Sbjct: 484 DHQEAKRDSK-KRKTMPKWIDQIRVVSENGLEGPH-DDGYNWRKYGQKDILGAKYPRSYY 657
Query: 232 RCTAGNCE 239
RCT N +
Sbjct: 658 RCTFRNTQ 681
>TC17544 similar to PIR|T04919|T04919 DNA-binding protein homolog T9A21.10 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(14%)
Length = 578
Score = 53.5 bits (127), Expect = 6e-08
Identities = 46/185 (24%), Positives = 78/185 (41%), Gaps = 10/185 (5%)
Frame = +1
Query: 41 IFDMIPPSQSDDQQSSSFAGYIDLLSTNDFTSSSATLFDWFPTIDTDMSAPPSTT----- 95
I + P + D S +F + +S + +++ P +T PS++
Sbjct: 97 ISNSFPSAYHHDSSSQAFDDPPNYMSFTECLQGRVMDYNYSPFGNTSFGLSPSSSEAFSS 276
Query: 96 -----QLPPLPAPSPAASEVLNNPATPVSVSSSISSSSNAAGVADKVMVEENEHEHEEEL 150
Q PP + AT +++SSIS+SS++ A++ ++ + ++ +
Sbjct: 277 VDQGNQKPPEGDVAGGGGGGGETLAT-TTLNSSISASSSSDAGAEEDSGKKKKDNNQVKE 453
Query: 151 EGDEAEEKDEDHGIRLVENQNDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDG 210
EG+ + KD KK KK + R F TKS+VDHL+DG
Sbjct: 454 EGNNSGNKD------------------------KKKGEKKQKEPRFAFMTKSEVDHLEDG 561
Query: 211 YRWRK 215
YRWRK
Sbjct: 562 YRWRK 576
>TC11031 homologue to UP|WR65_ARATH (Q9LP56) Probable WRKY transcription
factor 65 (WRKY DNA-binding protein 65), partial (15%)
Length = 453
Score = 51.6 bits (122), Expect = 2e-07
Identities = 18/27 (66%), Positives = 23/27 (84%)
Frame = +3
Query: 209 DGYRWRKYGQKPVKNSPFPRSYYRCTA 235
D + WRKYGQKP+K SP+PR YYRC++
Sbjct: 351 DSWAWRKYGQKPIKGSPYPRGYYRCSS 431
>BP077795
Length = 395
Score = 50.1 bits (118), Expect = 7e-07
Identities = 30/69 (43%), Positives = 39/69 (56%), Gaps = 3/69 (4%)
Frame = -2
Query: 354 NIVNSADNLMISSTSFGGFLQNQENYGGGNSSLSSSMMTSTND-MVRNNGLLQDMIAP-- 410
N+ N A N +++SFG FLQ G + SS M + N ++RNNGLLQDM P
Sbjct: 367 NVANPAPNTHFNTSSFGSFLQG---INGVDFVQSSRFMANNNQGLLRNNGLLQDMFVPAH 197
Query: 411 MEMGGAGAK 419
ME GG G +
Sbjct: 196 MEFGGGGGR 170
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.309 0.126 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,365,267
Number of Sequences: 28460
Number of extensions: 121099
Number of successful extensions: 1944
Number of sequences better than 10.0: 272
Number of HSP's better than 10.0 without gapping: 1590
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1824
length of query: 420
length of database: 4,897,600
effective HSP length: 93
effective length of query: 327
effective length of database: 2,250,820
effective search space: 736018140
effective search space used: 736018140
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC135100.8


