

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
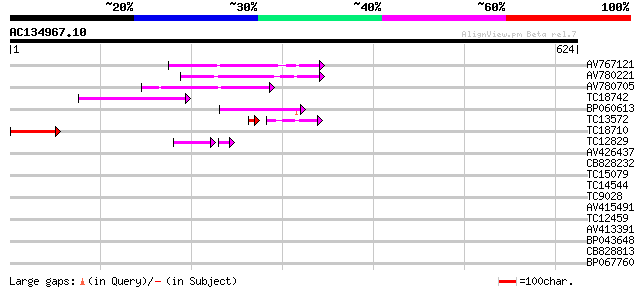
Query= AC134967.10 + phase: 0 /pseudo
(624 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV767121 127 7e-30
AV780221 125 2e-29
AV780705 72 2e-13
TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%) 53 2e-07
BP060613 49 2e-06
TC13572 40 1e-05
TC18710 44 8e-05
TC12829 39 2e-04
AV426437 37 0.009
CB828232 31 0.52
TC15079 similar to UP|Q84KA9 (Q84KA9) RING/C3HC4/PHD zinc finger... 31 0.52
TC14544 30 1.5
TC9028 weakly similar to UP|Q40077 (Q40077) Seed imbibition prot... 29 2.0
AV415491 28 4.4
TC12459 28 4.4
AV413391 28 5.7
BP043648 28 5.7
CB828813 27 7.5
BP067760 27 9.8
>AV767121
Length = 525
Score = 127 bits (318), Expect = 7e-30
Identities = 71/172 (41%), Positives = 96/172 (55%)
Frame = +2
Query: 175 LSGWNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSED 234
LS W + +SFGGR+ L++SVLT+LP++ LSFFK P G+ S L NF GG SE+
Sbjct: 2 LSKWKQKTVSFGGRIFLIQSVLTALPLFFLSFFKLPIGVGKSCVRLMRNFLWGG---SEN 172
Query: 235 NRKISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGEL 294
KI+ + W VC KE GGLG++ L FN ALLGK WR L + LW RV+ A+
Sbjct: 173 ENKIA*VKWTDVCKPKELGGLGIKDLFTFNKALLGK*RWRYLTEPDSLWRRVIEAQPDHY 352
Query: 295 GGRLEVGGRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSD 346
S WW+++ L + GWF + + VG+G T FWS+
Sbjct: 353 S--------CGSSWWNDILSL---CPEDADGWFSSGLKKLVGEGDQTKFWSE 475
>AV780221
Length = 461
Score = 125 bits (314), Expect = 2e-29
Identities = 67/159 (42%), Positives = 95/159 (59%), Gaps = 1/159 (0%)
Frame = +3
Query: 189 LVLLKSVLTSLPVYALSFFKAPTG-IISSIDSLFINFFLGGGGGSEDNRKISWLSWQTVC 247
L L++SVL +LP++ L FFK PTG +++++ FF G E +KI+W+ W TVC
Sbjct: 3 LCLIRSVLIALPLFHLPFFKLPTGGFYTNVNN**DPFF---GREVEGGKKIAWVKWSTVC 173
Query: 248 LKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGGRLEVGGRSVSC 307
K+ GGLGV+ L FN ALLGKW WRML +R GLWY+VL+ +Y +S S
Sbjct: 174 RPKDEGGLGVQNLGLFNKALLGKWRWRMLKERDGLWYKVLLIKY------KNAIPQSASS 335
Query: 308 WWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSD 346
WW+++ DGG GW + RR+G+G FW++
Sbjct: 336 WWNDLH--STCFEDGGGGWMQRGLCRRIGEGTEVKFWNE 446
>AV780705
Length = 524
Score = 72.4 bits (176), Expect = 2e-13
Identities = 46/148 (31%), Positives = 74/148 (49%), Gaps = 2/148 (1%)
Frame = -2
Query: 146 PFLYLDMPI--GGNPRRLCFWEPIVNRIKSRLSGWNSRFLSFGGRLVLLKSVLTSLPVYA 203
P YL +P G N W I ++K +L GW L+ G+ VL+K+++ ++P YA
Sbjct: 493 PGHYLGLPAIWGRNKSHSLVW--IEEKVKEKLEGWKETLLNQAGKEVLIKAIIQAIPSYA 320
Query: 204 LSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKISWLSWQTVCLKKECGGLGVRKLKEF 263
++ P +S+ +L +F+ G E + + L KE GG+G R L+
Sbjct: 319 MTIVHFPKTFCNSLYALVADFWWKSQG--EGAVEYTGAVGTK*PLSKEKGGVGFRDLRTQ 146
Query: 264 NLALLGKWCWRMLVDRGGLWYRVLVARY 291
NLA L + WR+L + LW RV+ + Y
Sbjct: 145 NLAFLARQAWRVLTNPEALWVRVMKSLY 62
>TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%)
Length = 912
Score = 52.8 bits (125), Expect = 2e-07
Identities = 36/124 (29%), Positives = 64/124 (51%)
Frame = -3
Query: 76 VSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSMLVGVNIGSSWLYEAA 135
+SHL FAD+ ++ + ++ + L +F+ +SGL N KS V + + + +
Sbjct: 373 ISHLCFADNLMVFSNGYLESIAIINNVLHIFQHLSGLTPNPAKSE-VFILVKETTKNQIT 197
Query: 136 SVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWNSRFLSFGGRLVLLKSV 195
++L K G +P L +P + + +V+R S+L W R LS+ GRL L+ S+
Sbjct: 196 NMLGYKEGKLPVR*LGVPQLTTKLKASDCKVLVDRKPSKLQHWTGRSLSYAGRLQLINSI 17
Query: 196 LTSL 199
L S+
Sbjct: 16 LFSM 5
>BP060613
Length = 378
Score = 49.3 bits (116), Expect = 2e-06
Identities = 32/100 (32%), Positives = 47/100 (47%), Gaps = 6/100 (6%)
Frame = +1
Query: 232 SEDNRKISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARY 291
S+ R + W W + K+ GGLG + + N ALL K WR+L + LW ++L A Y
Sbjct: 55 SKGTRGVHWRKWDLLTELKDEGGLGFKDFEIQNQALLAKQAWRILHNPDALWVQILKALY 234
Query: 292 GELGGRLEVGGRSVSCW-WSEVRR-----LRDGVGDGGSG 325
L+ R+ + W WS + L+ G GSG
Sbjct: 235 FPHHDFLQTTKRTGASWVWSSLLHGRELLLKQGQWQLGSG 354
>TC13572
Length = 534
Score = 40.4 bits (93), Expect(2) = 1e-05
Identities = 21/62 (33%), Positives = 33/62 (52%)
Frame = -3
Query: 283 WYRVLVARYGELGGRLEVGGRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTS 342
W RV++A+YG+ G ++S WW +V+ G G +GWF + V + + G T
Sbjct: 478 WCRVVLAKYGD------GGCTNISSWWRDVQT----AGGGENGWFEEGVHKVLRSGNHTR 329
Query: 343 FW 344
FW
Sbjct: 328 FW 323
Score = 25.8 bits (55), Expect(2) = 1e-05
Identities = 9/12 (75%), Positives = 10/12 (83%)
Frame = -1
Query: 264 NLALLGKWCWRM 275
N ALLGKW WR+
Sbjct: 534 NKALLGKWLWRL 499
>TC18710
Length = 843
Score = 43.9 bits (102), Expect = 8e-05
Identities = 22/56 (39%), Positives = 34/56 (60%)
Frame = +2
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKS 56
W K I VS + + +NG + +RGLRQGDP SP+LFL E L++++++
Sbjct: 653 WVKMIMILVSGVSYNYKINGVVGPKLLPQRGLRQGDPFSPYLFLFTMEVLSLLIQN 820
>TC12829
Length = 448
Score = 38.9 bits (89), Expect(2) = 2e-04
Identities = 19/46 (41%), Positives = 27/46 (58%)
Frame = +3
Query: 181 RFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFL 226
+FL GR VL+KSV ++P Y +S F P I S I+ + F+L
Sbjct: 255 KFLYRAGREVLIKSVTKAIPTYIMSCFALPDSICSQIEGMVGKFYL 392
Score = 23.1 bits (48), Expect(2) = 2e-04
Identities = 7/17 (41%), Positives = 9/17 (52%)
Frame = +1
Query: 231 GSEDNRKISWLSWQTVC 247
G R I WL W+ +C
Sbjct: 397 GDVSRRSIHWLGWKKLC 447
>AV426437
Length = 303
Score = 37.0 bits (84), Expect = 0.009
Identities = 14/30 (46%), Positives = 21/30 (69%)
Frame = +1
Query: 233 EDNRKISWLSWQTVCLKKECGGLGVRKLKE 262
E+ RKI+W++W+T+C K GLGV+ E
Sbjct: 214 ENQRKIAWINWETICKPKWLRGLGVKNWGE 303
>CB828232
Length = 532
Score = 31.2 bits (69), Expect = 0.52
Identities = 16/31 (51%), Positives = 21/31 (67%)
Frame = +1
Query: 178 WNSRFLSFGGRLVLLKSVLTSLPVYALSFFK 208
W + LS G R+ K VL+SLP++ LSFFK
Sbjct: 388 WKTLCLS-GARVCCFKMVLSSLPLFFLSFFK 477
>TC15079 similar to UP|Q84KA9 (Q84KA9) RING/C3HC4/PHD zinc finger-like
protein, partial (20%)
Length = 921
Score = 31.2 bits (69), Expect = 0.52
Identities = 16/30 (53%), Positives = 17/30 (56%)
Frame = -2
Query: 295 GGRLEVGGRSVSCWWSEVRRLRDGVGDGGS 324
GGR EVGG V C+W E DG G GS
Sbjct: 809 GGRREVGGGVVGCYW-EKASAADGAGGVGS 723
>TC14544
Length = 1160
Score = 29.6 bits (65), Expect = 1.5
Identities = 25/94 (26%), Positives = 41/94 (43%), Gaps = 12/94 (12%)
Frame = +2
Query: 259 KLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGGRLEVG----------GRSVSCW 308
K+K F++ + CWR L L + A ++GG+L V G+ + W
Sbjct: 452 KIKGFDVK---RGCWRELKGLNSLPIFLCGATMADVGGKLVVVWEYQAVKGGIGKEIDIW 622
Query: 309 WS--EVRRLRDGVGDGGSGWFGDQVLRRVGDGVV 340
+ EV + ++G GG WFG + G +V
Sbjct: 623 CAEIEVNKKQNGELWGGVCWFGKVLSVPKGSSIV 724
>TC9028 weakly similar to UP|Q40077 (Q40077) Seed imbibition protein,
partial (5%)
Length = 652
Score = 29.3 bits (64), Expect = 2.0
Identities = 18/38 (47%), Positives = 21/38 (54%), Gaps = 7/38 (18%)
Frame = +2
Query: 305 VSCWWS-EVRRLRDGVGDGGSGW------FGDQVLRRV 335
V CWWS R+R G G G GW FGDQ +R+V
Sbjct: 203 VQCWWSCRGPRVRGG-GRGQRGWIGW*SSFGDQGVRQV 313
>AV415491
Length = 365
Score = 28.1 bits (61), Expect = 4.4
Identities = 16/41 (39%), Positives = 21/41 (51%)
Frame = +1
Query: 237 KISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLV 277
+ SW+ + T C + GL L F ALLG W WR +V
Sbjct: 157 RASWMVFLTSC*HRRGQGL----LLWFLEALLGTWFWRFIV 267
>TC12459
Length = 797
Score = 28.1 bits (61), Expect = 4.4
Identities = 31/113 (27%), Positives = 47/113 (41%)
Frame = +3
Query: 227 GGGGGSEDNRKISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRV 286
GGGGG E ++ + + + G+ R+ LG R+ V GG+ R+
Sbjct: 138 GGGGGGEGRTEVVVVDETAIGEVRGVAGVSARQRVH-----LGPLPVRV-VGEGGVVERL 299
Query: 287 LVARYGELGGRLEVGGRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGV 339
+A+ L+ GR + L G D G+ WF Q L+R DGV
Sbjct: 300 RLAQRNPKC--LDAFGRVFDICGDDGDELPGG--DDGARWFSLQYLQRTSDGV 446
>AV413391
Length = 428
Score = 27.7 bits (60), Expect = 5.7
Identities = 12/26 (46%), Positives = 13/26 (49%)
Frame = +1
Query: 290 RYGELGGRLEVGGRSVSCWWSEVRRL 315
R GGRL R +CWW RRL
Sbjct: 217 RASTCGGRLRRRVRGSTCWWRSPRRL 294
>BP043648
Length = 403
Score = 27.7 bits (60), Expect = 5.7
Identities = 25/75 (33%), Positives = 31/75 (41%), Gaps = 4/75 (5%)
Frame = -2
Query: 227 GGGGGSEDNRKISWLSWQTVCLK----KECGGLGVRKLKEFNLALLGKWCWRMLVDRGGL 282
GGGG S R + LSW+ C + GL +RK F W + G
Sbjct: 198 GGGGVS---RALRLLSWRPFCSEILPMAVVVGLEIRKTTSFRRE------WTCX*NECGF 46
Query: 283 WYRVLVARYGELGGR 297
W RV R E+GGR
Sbjct: 45 W*RV-SGRVDEIGGR 4
>CB828813
Length = 465
Score = 27.3 bits (59), Expect = 7.5
Identities = 19/37 (51%), Positives = 20/37 (53%), Gaps = 1/37 (2%)
Frame = -2
Query: 290 RYGE-LGGRLEVGGRSVSCWWSEVRRLRDGVGDGGSG 325
R GE LGG L GG V EV +R GVG GG G
Sbjct: 341 RAGEPLGGGL--GGGGVGIGGLEVEEVRVGVGGGGGG 237
>BP067760
Length = 535
Score = 26.9 bits (58), Expect = 9.8
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = -2
Query: 254 GLGVRKLKEFNLALLGKWCWRMLVDRGGLWY 284
G+G + LALL + WR+L+D G L+Y
Sbjct: 315 GVGALDIGFTPLALLYRSYWRLLLDFGNLYY 223
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.348 0.155 0.570
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,341,181
Number of Sequences: 28460
Number of extensions: 218327
Number of successful extensions: 2605
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 2539
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2595
length of query: 624
length of database: 4,897,600
effective HSP length: 96
effective length of query: 528
effective length of database: 2,165,440
effective search space: 1143352320
effective search space used: 1143352320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC134967.10


