

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
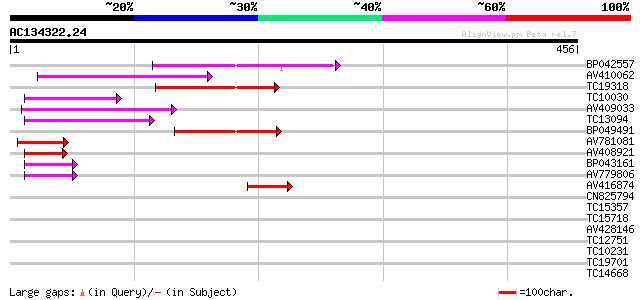
Query= AC134322.24 + phase: 0
(456 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP042557 92 1e-19
AV410062 84 5e-17
TC19318 similar to UP|Q94JM3 (Q94JM3) AT5g62000/mtg10_20, partia... 81 4e-16
TC10030 similar to UP|Q94JM3 (Q94JM3) AT5g62000/mtg10_20, partia... 72 1e-13
AV409033 72 1e-13
TC13094 similar to UP|Q94JM3 (Q94JM3) AT5g62000/mtg10_20, partia... 71 3e-13
BP049491 68 4e-12
AV781081 53 9e-08
AV408921 46 1e-05
BP043161 44 6e-05
AV779806 42 2e-04
AV416874 41 5e-04
CN825794 37 0.009
TC15357 similar to UP|Q9ZV06 (Q9ZV06) At2g39020 protein, partial... 33 0.13
TC15718 32 0.28
AV428146 30 0.83
TC12751 similar to UP|LEU3_BRANA (P29102) 3-isopropylmalate dehy... 30 0.83
TC10231 similar to UP|O23110 (O23110) AP2 domain containing prot... 30 1.1
TC19701 similar to UP|AAR04332 (AAR04332) Eukaryotic translation... 29 1.8
TC14668 UP|BAC98256 (BAC98256) MKIAA1813 protein (Fragment), par... 29 1.8
>BP042557
Length = 545
Score = 92.4 bits (228), Expect = 1e-19
Identities = 55/154 (35%), Positives = 84/154 (53%), Gaps = 3/154 (1%)
Frame = +2
Query: 116 FVKTLTRSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGF 175
F KTLT SD + F +PR A+ VFP LD + +Q L D++ KF H+ RG
Sbjct: 65 FCKTLTASDTSTHGGFSVPRRAAEKVFPPLDYSQQPPAQELIARDLNDNEWKFRHIFRGQ 244
Query: 176 PKRNMLYISEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDEL 233
PKR++L + W+ FV K+LVAGDSV+F+ + ++ +GIRR +Q + ++ D +
Sbjct: 245 PKRHLL-TTGWSVFVSAKRLVAGDSVLFIWNEKNQLLLGIRRANRSQTIMPSSVLSSDSM 421
Query: 234 EKAVM-EALKLAEENKAFEIVYYPQGDDWCDFVV 266
++ A A N F I Y P+ +FV+
Sbjct: 422 HIGLLAAAAHAASTNSRFTIFYNPRASP-SEFVI 520
>AV410062
Length = 435
Score = 84.0 bits (206), Expect = 5e-17
Identities = 49/142 (34%), Positives = 67/142 (46%), Gaps = 1/142 (0%)
Frame = +3
Query: 23 VKIPKLHSRVYYFPQGHLENASPSSSSITHTH-SFLQSFRPFTLCIVSAVDLLADPHTDE 81
V +P L SRV YFPQGH E + S++ H S P +C + V + AD TDE
Sbjct: 6 VSLPALGSRVVYFPQGHSEQVAVSTNKEVDAHIPNYPSLPPQLICQLHNVTMHADAETDE 185
Query: 82 VFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSFHIPRFCADNV 141
V+ ++ L P+ E + F KTLT SD + F +PR A+ V
Sbjct: 186 VYAQMTLQPLNPQEQKEAYLPAELGTPSKQPTNYFCKTLTASDTSTHGGFSVPRRAAEKV 365
Query: 142 FPQLDLEAESSSQHLFVTDVHG 163
FP LD + +Q L D+HG
Sbjct: 366 FPPLDFSQQPPAQELIARDLHG 431
>TC19318 similar to UP|Q94JM3 (Q94JM3) AT5g62000/mtg10_20, partial (23%)
Length = 585
Score = 80.9 bits (198), Expect = 4e-16
Identities = 41/100 (41%), Positives = 61/100 (61%)
Frame = +3
Query: 118 KTLTRSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPK 177
KTLT SD + F + R AD P LD+ + +Q L D+HG +F H+ G P+
Sbjct: 39 KTLTASDTSTDGGFSVLRRHADECLPPLDMSKQPPTQALVAKDLHGSEWRFRHIFLGQPR 218
Query: 178 RNMLYISEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRR 217
R++L +S W+ FV K+LVAGD+ IF++ G++ VG+RR
Sbjct: 219 RHLL-LSGWSVFVSSKRLVAGDAFIFLRGENGELRVGVRR 335
>TC10030 similar to UP|Q94JM3 (Q94JM3) AT5g62000/mtg10_20, partial (13%)
Length = 588
Score = 72.4 bits (176), Expect = 1e-13
Identities = 33/78 (42%), Positives = 45/78 (57%)
Frame = +3
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P+ RV+YFPQGH+E S++ + H + P LC V V
Sbjct: 354 ELWHACAGPLVTVPREGERVFYFPQGHIEQVEASTNQVAELHMPVYDLPPKILCRVINVM 533
Query: 73 LLADPHTDEVFVKLLLTP 90
L A+P TDEVF ++ L P
Sbjct: 534 LKAEPDTDEVFAQVTLLP 587
>AV409033
Length = 418
Score = 72.4 bits (176), Expect = 1e-13
Identities = 40/125 (32%), Positives = 58/125 (46%)
Frame = +3
Query: 10 VRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVS 69
+ E+WH CA V +P + S V YFPQGH E + S T + +C++
Sbjct: 42 INSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKETDFIPSYPNLPSKLICMLH 221
Query: 70 AVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNAR 129
V L ADP TDEV+ ++ L P+ ++ +R F KTLT SD +
Sbjct: 222 NVALHADPETDEVYAQMTLQPVNKYDKDAILASDMGLKQNRQPTEFFCKTLTASDTSTHG 401
Query: 130 SFHIP 134
F +P
Sbjct: 402 GFSVP 416
>TC13094 similar to UP|Q94JM3 (Q94JM3) AT5g62000/mtg10_20, partial (18%)
Length = 685
Score = 71.2 bits (173), Expect = 3e-13
Identities = 39/104 (37%), Positives = 51/104 (48%)
Frame = +2
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P+ RV+YFPQGH+E S++ + H + LC V V
Sbjct: 374 ELWHACAGPLVTVPREGERVFYFPQGHIEQVEASTNQVADEHMPVYDLPSKILCRVINVQ 553
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSF 116
L A+P TDEVF ++ L P N KE R V SF
Sbjct: 554 LKAEPDTDEVFAQVTLLPEPNQDENAVEKEPPPPPXXRFHVHSF 685
>BP049491
Length = 516
Score = 67.8 bits (164), Expect = 4e-12
Identities = 33/86 (38%), Positives = 54/86 (62%)
Frame = -3
Query: 133 IPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKR 192
+PR A++ P LD + + SQ L D+HG KF H+ RG P+ ++L + W+ FV +
Sbjct: 514 VPRRAAEDCLPPLDYKQQRPSQELVAKDLHGVEWKFCHIYRG*PRWHLL-TTGWSIFVSQ 338
Query: 193 KKLVAGDSVIFMKDSTGKIFVGIRRN 218
LV+GD+V F++ G++ +GIRR+
Sbjct: 337 NNLVSGDAVSFLRGENGELRLGIRRS 260
>AV781081
Length = 528
Score = 53.1 bits (126), Expect = 9e-08
Identities = 20/41 (48%), Positives = 27/41 (65%)
Frame = +2
Query: 7 PSHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSS 47
P + PE+WH CA V +P+L S YYFPQGH+E + S+
Sbjct: 398 PKGINPELWHACAGPLVSLPQLGSLAYYFPQGHIEQVAAST 520
>AV408921
Length = 429
Score = 46.2 bits (108), Expect = 1e-05
Identities = 18/34 (52%), Positives = 22/34 (63%)
Frame = +2
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPS 46
E+WH CA V +P+ RVYYFPQGH+E S
Sbjct: 326 ELWHACAGPLVTLPREGERVYYFPQGHMEQLEAS 427
>BP043161
Length = 509
Score = 43.9 bits (102), Expect = 6e-05
Identities = 18/42 (42%), Positives = 24/42 (56%)
Frame = +2
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTH 54
E+WH CA V +P +RV YFPQGH E S +++ H
Sbjct: 347 ELWHACAGPLVSLPTAGTRVVYFPQGHSEQVSATTNREVDGH 472
>AV779806
Length = 546
Score = 42.0 bits (97), Expect = 2e-04
Identities = 18/42 (42%), Positives = 24/42 (56%)
Frame = +2
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTH 54
E+ H CA V +P + SRV YFPQGH E + S++ H
Sbjct: 410 ELGHACAGPLVSLPPIGSRVVYFPQGHSEQVAASTNREVDAH 535
>AV416874
Length = 256
Score = 40.8 bits (94), Expect = 5e-04
Identities = 19/36 (52%), Positives = 25/36 (68%)
Frame = +2
Query: 192 RKKLVAGDSVIFMKDSTGKIFVGIRRNTQFVAAAAE 227
RKKLVAGDSV+F++ G + VGIRR + + AE
Sbjct: 2 RKKLVAGDSVVFLRAENGDLCVGIRRAKRGIGGGAE 109
>CN825794
Length = 212
Score = 36.6 bits (83), Expect = 0.009
Identities = 19/53 (35%), Positives = 31/53 (57%)
Frame = +2
Query: 235 KAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKM 287
+A++EA LA + FE+ YYP+G +F ++V+ +M+I MR KM
Sbjct: 35 EALIEAANLAANKEPFEVAYYPRGST-PEFCAKTSLVESAMQITRRSGMRFKM 190
>TC15357 similar to UP|Q9ZV06 (Q9ZV06) At2g39020 protein, partial (69%)
Length = 1578
Score = 32.7 bits (73), Expect = 0.13
Identities = 36/148 (24%), Positives = 59/148 (39%), Gaps = 7/148 (4%)
Frame = +3
Query: 2 SLQKQPSHVRPEIWHTCATAAVKIPKLHSRVYYFPQ----GHLENASPSSSSITHTHSFL 57
SL + +H P +P +H Y HL + + SS+T T F
Sbjct: 411 SLPETATHRHPVFTRIRLAVPSDVPHIHKMTYQMAVFERLTHL--FATTESSLTST-LFS 581
Query: 58 QSFRPFTLCIVSAVDLLADPHTDEVFVK-LLLTPITNDVHLENPKEEVANLNDRNEVVS- 115
+PF V +++ ++P TD F P+T VHLE P ++ RN++ +
Sbjct: 582 PDNKPFHSFTVFILEVSSNPFTDTHFDNDPFYKPVTKTVHLELPLDDPEKETFRNQLGNE 761
Query: 116 -FVKTLTRSDVNNARSFHIPRFCADNVF 142
FV N + P F +++F
Sbjct: 762 VFVAGFVLFSPNYSTFLAKPGFYVEDLF 845
>TC15718
Length = 722
Score = 31.6 bits (70), Expect = 0.28
Identities = 14/40 (35%), Positives = 24/40 (60%), Gaps = 1/40 (2%)
Frame = +2
Query: 11 RPEIWHTCATAAV-KIPKLHSRVYYFPQGHLENASPSSSS 49
R +IW+ CAT V ++ +++ + YFPQ + +A P S
Sbjct: 221 RNQIWNPCATLVVLRLVQMNLEISYFPQSKVPSAIPKVDS 340
>AV428146
Length = 260
Score = 30.0 bits (66), Expect = 0.83
Identities = 13/30 (43%), Positives = 16/30 (53%)
Frame = +3
Query: 333 WWVELISHKPAPTPFPQTKKFRTTQSSAQL 362
W++ L H P P PFP +K T S QL
Sbjct: 57 WFLPLQPHPPPPPPFPISKPISTPPSCPQL 146
>TC12751 similar to UP|LEU3_BRANA (P29102) 3-isopropylmalate dehydrogenase,
chloroplast precursor (Beta-IPM dehydrogenase) (IMDH)
(3-IPM-DH) , partial (25%)
Length = 450
Score = 30.0 bits (66), Expect = 0.83
Identities = 23/78 (29%), Positives = 37/78 (46%), Gaps = 8/78 (10%)
Frame = +2
Query: 325 QIPRRVNPWWVELISHKPAP-TPFPQTKKFRTT----QSSAQLSDKKE---TLLNGDGFP 376
Q P+ ++L+ +P +PFP R T + SA KK TLL GDG
Sbjct: 44 QTPKMAASVQMQLLHARPMKASPFPSFSLPRPTSMRCRCSAATPSKKSYSITLLPGDGIG 223
Query: 377 VDIQRVRHDLVSISGPIH 394
++ V D++ ++G +H
Sbjct: 224 PEVVSVAKDVLLLAGSLH 277
>TC10231 similar to UP|O23110 (O23110) AP2 domain containing protein RAP2.8
(Fragment), partial (22%)
Length = 707
Score = 29.6 bits (65), Expect = 1.1
Identities = 23/64 (35%), Positives = 32/64 (49%)
Frame = +1
Query: 146 DLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKRKKLVAGDSVIFMK 205
D A + L DV G+V +F + + +L W+ FVK K L AGD+V F +
Sbjct: 25 DATAAAKGMLLNFEDVGGKVWRFRYSYWNSSQSYVL-TKGWSRFVKEKNLRAGDAVRFHR 201
Query: 206 DSTG 209
STG
Sbjct: 202 -STG 210
>TC19701 similar to UP|AAR04332 (AAR04332) Eukaryotic translation initiation
factor 4E (Fragment), partial (67%)
Length = 565
Score = 28.9 bits (63), Expect = 1.8
Identities = 23/88 (26%), Positives = 39/88 (44%), Gaps = 1/88 (1%)
Frame = +2
Query: 325 QIPRRVNPWWVELISHKPAP-TPFPQTKKFRTTQSSAQLSDKKETLLNGDGFPVDIQRVR 383
+IPR PW ++ + P+P T P T + TT +S + ++ T++ P +
Sbjct: 29 EIPRETKPWLWKIPQNPPSPTTKSPPTLRLLTTPTSRK--ERSSTMMT----PPPLPNPP 190
Query: 384 HDLVSISGPIHSHIILNSSETKFPATHN 411
+S + I S I+ S T P N
Sbjct: 191 PPPLSTNPIIPSRILGPSGSTTLPTRAN 274
>TC14668 UP|BAC98256 (BAC98256) MKIAA1813 protein (Fragment), partial (3%)
Length = 597
Score = 28.9 bits (63), Expect = 1.8
Identities = 13/25 (52%), Positives = 15/25 (60%)
Frame = +1
Query: 324 SQIPRRVNPWWVELISHKPAPTPFP 348
S +PRR+ PWW L S P PT P
Sbjct: 493 SSLPRRLPPWWGHLSS--PGPTAGP 561
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.133 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,403,373
Number of Sequences: 28460
Number of extensions: 124476
Number of successful extensions: 752
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 739
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 746
length of query: 456
length of database: 4,897,600
effective HSP length: 93
effective length of query: 363
effective length of database: 2,250,820
effective search space: 817047660
effective search space used: 817047660
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC134322.24


