

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
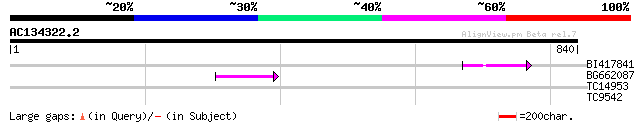
Query= AC134322.2 + phase: 0 /pseudo
(840 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BI417841 54 1e-07
BG662087 47 1e-05
TC14953 weakly similar to UP|Q9LK01 (Q9LK01) Hydrolase-like prot... 32 0.54
TC9542 homologue to UP|CAMT_MEDSA (Q40313) Caffeoyl-CoA O-methyl... 28 4.6
>BI417841
Length = 617
Score = 53.9 bits (128), Expect = 1e-07
Identities = 36/106 (33%), Positives = 53/106 (49%), Gaps = 4/106 (3%)
Frame = +1
Query: 671 LSVDGSS--NLKGSGAGVTLEGPYG--VLIEQSLRFDFKSSNNQAEYEALFAGMRLALKV 726
L DGSS N +GAG L G V + + + +NNQAEY L G++ A +
Sbjct: 142 LEFDGSSKGNPGSAGAGAVLRAEDGSKVYLREGVG---NQTNNQAEYRGLILGLKHAHEQ 312
Query: 727 VVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQKFVSF 772
+ + K DSQLV +QV G+++ ++P + L KF SF
Sbjct: 313 GYQHINVKGDSQLVCKQVEGSWKARNPNIASLCNEAKELKSKFQSF 450
>BG662087
Length = 373
Score = 47.4 bits (111), Expect = 1e-05
Identities = 33/93 (35%), Positives = 45/93 (47%)
Frame = +2
Query: 306 NGECARTTLI*IRCVPRMLIPCRA*INWYITLQGMTFLASWMLTRATTKFECT*RMKKRQ 365
NG C TT I* R + LI C+ * NW++ M+ A WM T+ TK CT +M+ ++
Sbjct: 20 NGACGWTTRI*TRHAQKTLIHCQV*TNWWMERPTMSC*A*WMPTQVITKSRCTHQMRTKR 199
Query: 366 LL*QTKLIIVTKVCHLG*GILELLFKDWWIKSS 398
* I T+ G* + WI SS
Sbjct: 200 PS*PPG*ITATRQFRSG*RMPGRHINX*WIGSS 298
>TC14953 weakly similar to UP|Q9LK01 (Q9LK01) Hydrolase-like protein,
partial (27%)
Length = 591
Score = 31.6 bits (70), Expect = 0.54
Identities = 26/105 (24%), Positives = 45/105 (42%), Gaps = 7/105 (6%)
Frame = +2
Query: 700 LRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYL 759
L F F + + + L L + K+ E K L+ + T F TK P L+
Sbjct: 35 LHFSFSWNPTSTQSHFKKSAFHLLLMLKSKD*EQKH*FLLMGMEQTSPFGTKSPPSLLKT 214
Query: 760 ESVMYLT---QKFVSFKYYMCLES----RTVEQTCWLSWLAPSDK 797
+++L Q+ + + M L S + ++ TC LSW+ + K
Sbjct: 215 TGLLFLIGLFQELLKMRACMILSSTLLWKLLQMTCSLSWIRWTSK 349
>TC9542 homologue to UP|CAMT_MEDSA (Q40313) Caffeoyl-CoA
O-methyltransferase (Trans-caffeoyl-CoA
3-O-methyltransferase) (CCoAMT) (CCoAOMT) , complete
Length = 1012
Score = 28.5 bits (62), Expect = 4.6
Identities = 20/55 (36%), Positives = 28/55 (50%)
Frame = +3
Query: 346 WMLTRATTKFECT*RMKKRQLL*QTKLIIVTKVCHLG*GILELLFKDWWIKSSRV 400
WMLTR TT + C L* KL+++TK+ + L LF W+K R+
Sbjct: 411 WMLTRRTTNWVC---------L*LRKLVLLTKL--ISEKALLSLFLMKWLKMRRI 542
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.358 0.156 0.572
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,821,093
Number of Sequences: 28460
Number of extensions: 200710
Number of successful extensions: 1892
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 1381
Number of HSP's successfully gapped in prelim test: 37
Number of HSP's that attempted gapping in prelim test: 493
Number of HSP's gapped (non-prelim): 1444
length of query: 840
length of database: 4,897,600
effective HSP length: 98
effective length of query: 742
effective length of database: 2,108,520
effective search space: 1564521840
effective search space used: 1564521840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC134322.2


