

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.16 - phase: 0
(690 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
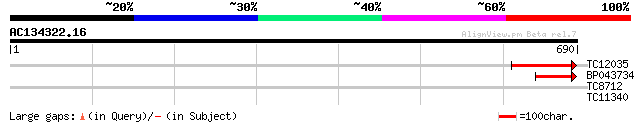
Score E
Sequences producing significant alignments: (bits) Value
TC12035 135 2e-32
BP043734 94 7e-20
TC8712 homologue to UP|O81413 (O81413) Ferric leghemoglobin redu... 28 3.7
TC11340 27 8.3
>TC12035
Length = 515
Score = 135 bits (340), Expect = 2e-32
Identities = 69/79 (87%), Positives = 75/79 (94%)
Frame = +1
Query: 611 IEEENEKKTEKVMQALLNSLFGNVKVGENGKLIINIDGNVAELNKESGEVESENEGLKER 670
+EEENEKK EKVM ALL SLFG+VKVG NG+LIIN+DGNVAELN+ESGEVESENEGLKER
Sbjct: 1 LEEENEKKAEKVMHALLVSLFGDVKVGVNGQLIINLDGNVAELNQESGEVESENEGLKER 180
Query: 671 VRTAFRRIQSSVKPIPLSA 689
V+TAFRRIQSSVKPIPLSA
Sbjct: 181 VKTAFRRIQSSVKPIPLSA 237
>BP043734
Length = 458
Score = 94.0 bits (232), Expect = 7e-20
Identities = 48/49 (97%), Positives = 48/49 (97%)
Frame = -1
Query: 641 KLIINIDGNVAELNKESGEVESENEGLKERVRTAFRRIQSSVKPIPLSA 689
KLIINIDGNVAELNKESGEVESENEGLKERV TAFRRIQSSVKPIPLSA
Sbjct: 458 KLIINIDGNVAELNKESGEVESENEGLKERVNTAFRRIQSSVKPIPLSA 312
>TC8712 homologue to UP|O81413 (O81413) Ferric leghemoglobin reductase-2
precursor , partial (60%)
Length = 1230
Score = 28.5 bits (62), Expect = 3.7
Identities = 19/60 (31%), Positives = 27/60 (44%)
Frame = -2
Query: 239 TISQGGRVLIPAYALGRAQELLLILDEYWANHPELQNIPIYYASPLAKKCLTVYETYTLS 298
++S +L AL A LLL + N P Y++PLA C +V+ TY S
Sbjct: 728 SVSFSANILTSPSALSIALALLLAMKG---------NFPTRYSTPLAFTCSSVFPTYATS 576
>TC11340
Length = 543
Score = 27.3 bits (59), Expect = 8.3
Identities = 17/46 (36%), Positives = 20/46 (42%), Gaps = 11/46 (23%)
Frame = -1
Query: 10 NGGTINRETEDQL-----------IVTPLGAGNEVGRSCVYMTYKG 44
NGG + TE L V PL A R CVY+T+KG
Sbjct: 252 NGGNSYKRTEWSLDEQHNTCLPSASVIPLSATTPFSRVCVYVTFKG 115
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.134 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,111,248
Number of Sequences: 28460
Number of extensions: 126611
Number of successful extensions: 619
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 618
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 619
length of query: 690
length of database: 4,897,600
effective HSP length: 97
effective length of query: 593
effective length of database: 2,136,980
effective search space: 1267229140
effective search space used: 1267229140
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC134322.16


