

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
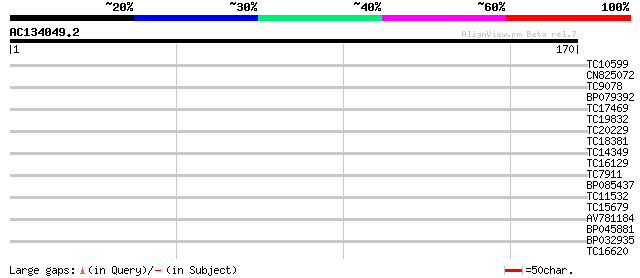
Query= AC134049.2 - phase: 0
(170 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC10599 29 0.37
CN825072 29 0.37
TC9078 similar to AAQ89615 (AAQ89615) At3g17205, partial (12%) 27 1.4
BP079392 27 1.8
TC17469 similar to UP|Q8S8Z0 (Q8S8Z0) Protein phosphatase 2C, pa... 27 2.4
TC19832 similar to GB|AAO39903.1|28372842|BT003675 At5g67590 {Ar... 26 3.2
TC20229 26 4.1
TC18381 similar to UP|Q9ASR8 (Q9ASR8) AT5g59960/mmn10_180, parti... 26 4.1
TC14349 UP|Q9M4R0 (Q9M4R0) Ubiquitin conjugating protein (Ubiqu... 25 5.4
TC16129 similar to UP|AAR14275 (AAR14275) Predicted protein, par... 25 5.4
TC7911 similar to UP|Q39800 (Q39800) Delta-tonoplast intrinsic p... 25 5.4
BP085437 25 5.4
TC11532 homologue to UP|Q8L5K5 (Q8L5K5) Ovule/fiber cell elongat... 25 5.4
TC15679 25 7.0
AV781184 25 7.0
BP045881 25 9.2
BP032935 25 9.2
TC16620 weakly similar to UP|Q9FN67 (Q9FN67) Gb|AAD56319.1, part... 25 9.2
>TC10599
Length = 600
Score = 29.3 bits (64), Expect = 0.37
Identities = 10/21 (47%), Positives = 18/21 (85%)
Frame = +2
Query: 96 LCYLEGNAPMQRVLEVYDNYI 116
LCY++GNAPM+ ++ +YD ++
Sbjct: 206 LCYIKGNAPMRSLVCLYDIFM 268
>CN825072
Length = 712
Score = 29.3 bits (64), Expect = 0.37
Identities = 17/44 (38%), Positives = 20/44 (44%), Gaps = 3/44 (6%)
Frame = +2
Query: 51 SQHATCHVLQYECRFREAVEFMEE---CSPSWNSFLSFMLTHNW 91
S HA + R R A E CSP+WNS S + TH W
Sbjct: 338 SAHARAQRRSFVVRHRVATWQWERRSICSPAWNSNRSAIRTHPW 469
>TC9078 similar to AAQ89615 (AAQ89615) At3g17205, partial (12%)
Length = 666
Score = 27.3 bits (59), Expect = 1.4
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = -3
Query: 38 AKRGFEINNQDGWSQHATCHVL 59
AK+ F+I+ WS ATCH L
Sbjct: 214 AKQRFKISKSKQWSARATCHKL 149
>BP079392
Length = 485
Score = 26.9 bits (58), Expect = 1.8
Identities = 11/17 (64%), Positives = 13/17 (75%)
Frame = -3
Query: 129 EVYLNAVALLLRLCVRD 145
E+YL+ AL L LCVRD
Sbjct: 465 EIYLSVCALFLSLCVRD 415
>TC17469 similar to UP|Q8S8Z0 (Q8S8Z0) Protein phosphatase 2C, partial
(13%)
Length = 568
Score = 26.6 bits (57), Expect = 2.4
Identities = 13/31 (41%), Positives = 21/31 (66%)
Frame = +3
Query: 31 MKEAEEAAKRGFEINNQDGWSQHATCHVLQY 61
+++AEEAAKR + Q G S + TC V+++
Sbjct: 15 IEDAEEAAKRLMQEAYQRGSSDNLTCVVVRF 107
>TC19832 similar to GB|AAO39903.1|28372842|BT003675 At5g67590 {Arabidopsis
thaliana;}, partial (75%)
Length = 530
Score = 26.2 bits (56), Expect = 3.2
Identities = 13/41 (31%), Positives = 20/41 (48%)
Frame = +1
Query: 84 SFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKELDKTD 124
+F H W +V + + P+ +V DN+ WK L K D
Sbjct: 415 AFAEKHGWEYVVK---KPHTPLLKVKSYADNFKWKGLPKVD 528
>TC20229
Length = 666
Score = 25.8 bits (55), Expect = 4.1
Identities = 9/17 (52%), Positives = 14/17 (81%)
Frame = -3
Query: 7 VLPQNEGENFIYGMLAF 23
+LPQ++GE+ +Y LAF
Sbjct: 433 LLPQDQGESLVYACLAF 383
>TC18381 similar to UP|Q9ASR8 (Q9ASR8) AT5g59960/mmn10_180, partial (53%)
Length = 758
Score = 25.8 bits (55), Expect = 4.1
Identities = 18/59 (30%), Positives = 28/59 (46%)
Frame = -3
Query: 107 RVLEVYDNYIWKELDKTDATVPEVYLNAVALLLRLCVRDELEFFGDRLKMLADRLADQV 165
R+ + NYI+ + + V E L +A L R+C L++L DRLA+ V
Sbjct: 513 RIFFLVHNYIFGSVINFQSYVGEELLQQIAPLRRICECP--------LELLVDRLAESV 361
>TC14349 UP|Q9M4R0 (Q9M4R0) Ubiquitin conjugating protein
(Ubiquitin-conjugating enzyme E2) (Ubiquitin-protein
ligase) (Ubiquitin carrier protein) , complete
Length = 897
Score = 25.4 bits (54), Expect = 5.4
Identities = 12/37 (32%), Positives = 16/37 (42%)
Frame = +3
Query: 60 QYECRFREAVEFMEECSPSWNSFLSFMLTHNWWHVAL 96
Q EC R +V +EEC W+ H + V L
Sbjct: 534 QLECSVRTSVSTIEECGKLWSRVGQLTKAHLYMLVGL 644
>TC16129 similar to UP|AAR14275 (AAR14275) Predicted protein, partial (19%)
Length = 599
Score = 25.4 bits (54), Expect = 5.4
Identities = 10/40 (25%), Positives = 19/40 (47%)
Frame = +2
Query: 79 WNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWK 118
WN F+L WW + +++G M+ + + + WK
Sbjct: 359 WNVHHEFLL---WWSIK*LWMQGT*RMRMTINILECLSWK 469
>TC7911 similar to UP|Q39800 (Q39800) Delta-tonoplast intrinsic protein,
complete
Length = 1018
Score = 25.4 bits (54), Expect = 5.4
Identities = 15/53 (28%), Positives = 21/53 (39%), Gaps = 3/53 (5%)
Frame = +3
Query: 50 WSQHATCHVLQYECRFREAVEFMEECSPSWNSFLSFML---THNWWHVALCYL 99
W QH +C + + E + CS W L T +WW + CYL
Sbjct: 594 WCQHLSCRAILWWVN--EPSTLLWPCSCQW*LP*QLDLLGWTSHWWWLGWCYL 746
>BP085437
Length = 456
Score = 25.4 bits (54), Expect = 5.4
Identities = 10/40 (25%), Positives = 19/40 (47%)
Frame = -1
Query: 79 WNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWK 118
WN F+L WW + +++G M+ + + + WK
Sbjct: 231 WNVHHEFLL---WWSIK*LWMQGT*RMRMTINILECLSWK 121
>TC11532 homologue to UP|Q8L5K5 (Q8L5K5) Ovule/fiber cell elongation protein
Ghfe1, partial (24%)
Length = 415
Score = 25.4 bits (54), Expect = 5.4
Identities = 13/39 (33%), Positives = 22/39 (56%), Gaps = 3/39 (7%)
Frame = +3
Query: 15 NFIYGMLAFP---LLELGQMKEAEEAAKRGFEINNQDGW 50
+F+ GM+ P ++ G+ EEA K+G +IN+ W
Sbjct: 66 DFVEGMVYTPTEAVMMTGRYASKEEAKKKGNKINSVGWW 182
>TC15679
Length = 609
Score = 25.0 bits (53), Expect = 7.0
Identities = 10/27 (37%), Positives = 13/27 (48%)
Frame = +3
Query: 91 WWHVALCYLEGNAPMQRVLEVYDNYIW 117
WW V L + GNA + VY + W
Sbjct: 252 WWEVMLDSVHGNAKTTGFVSVYYSLSW 332
>AV781184
Length = 462
Score = 25.0 bits (53), Expect = 7.0
Identities = 10/38 (26%), Positives = 14/38 (36%)
Frame = +1
Query: 49 GWSQHATCHVLQYECRFREAVEFMEECSPSWNSFLSFM 86
GW + V C V +EEC W + F+
Sbjct: 175 GWGRKPISEVYSLACNISHIVRELEECLDRWQFRVPFI 288
>BP045881
Length = 420
Score = 24.6 bits (52), Expect = 9.2
Identities = 14/39 (35%), Positives = 19/39 (47%), Gaps = 3/39 (7%)
Frame = -1
Query: 24 PLLELGQMKEAEEAAKRGFEINNQDGWSQH---ATCHVL 59
PL LG+ KE E K+G+ + GW H CH +
Sbjct: 225 PLRHLGEGKEVPEPWKKGWLLR---GWLLHTMPGDCHTM 118
>BP032935
Length = 534
Score = 24.6 bits (52), Expect = 9.2
Identities = 7/14 (50%), Positives = 10/14 (71%)
Frame = +2
Query: 89 HNWWHVALCYLEGN 102
H W ++ CYL+GN
Sbjct: 14 HGWCNMXYCYLQGN 55
>TC16620 weakly similar to UP|Q9FN67 (Q9FN67) Gb|AAD56319.1, partial (17%)
Length = 518
Score = 24.6 bits (52), Expect = 9.2
Identities = 8/29 (27%), Positives = 17/29 (58%)
Frame = -2
Query: 82 FLSFMLTHNWWHVALCYLEGNAPMQRVLE 110
+L ++ WWHV+ C E + M+ +++
Sbjct: 109 WLEWVRACKWWHVSCCSCEESEMMKMLMK 23
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.137 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,271,400
Number of Sequences: 28460
Number of extensions: 45894
Number of successful extensions: 255
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 255
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 255
length of query: 170
length of database: 4,897,600
effective HSP length: 84
effective length of query: 86
effective length of database: 2,506,960
effective search space: 215598560
effective search space used: 215598560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC134049.2


