

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
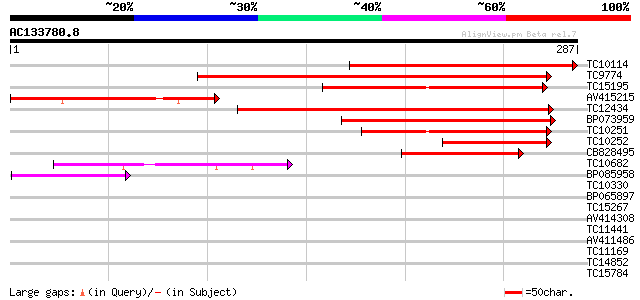
Query= AC133780.8 - phase: 0 /pseudo
(287 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC10114 similar to UP|Q9LYV6 (Q9LYV6) ABA-responsive protein-lik... 213 3e-56
TC9774 similar to UP|Q9FMW4 (Q9FMW4) Similarity to ABA-responsiv... 148 1e-36
TC15195 similar to UP|Q8S8F8 (Q8S8F8) Expressed protein, partial... 144 1e-35
AV415215 142 5e-35
TC12434 similar to UP|Q9FMW4 (Q9FMW4) Similarity to ABA-responsi... 137 3e-33
BP073959 113 4e-26
TC10251 similar to UP|Q8S8F8 (Q8S8F8) Expressed protein, partial... 100 3e-22
TC10252 similar to UP|Q8S8F8 (Q8S8F8) Expressed protein, partial... 62 1e-10
CB828495 61 2e-10
TC10682 similar to PIR|A86408|A86408 FH protein interacting prot... 57 5e-09
BP085958 43 5e-05
TC10330 similar to UP|O04344 (O04344) At2g30510 protein, partial... 36 0.007
BP065897 34 0.033
TC15267 similar to UP|EXTN_TOBAC (P13983) Extensin precursor (Ce... 34 0.033
AV414308 33 0.043
TC11441 similar to PIR|S54157|S54157 extensin-like protein - cow... 33 0.056
AV411486 33 0.073
TC11169 similar to UP|Q9SL72 (Q9SL72) At2g20090 protein, partial... 33 0.073
TC14852 similar to UP|Q9LZI2 (Q9LZI2) DTDP-glucose 4-6-dehydrata... 33 0.073
TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precurso... 32 0.096
>TC10114 similar to UP|Q9LYV6 (Q9LYV6) ABA-responsive protein-like, partial
(35%)
Length = 633
Score = 213 bits (542), Expect = 3e-56
Identities = 102/115 (88%), Positives = 106/115 (91%)
Frame = +1
Query: 173 KTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVN 232
K+FACY ST+TGPVAGTLYLS IH AFCSDRPL FTAPSGQ TWSYYKVMVPLGKIG VN
Sbjct: 1 KSFACYRSTSTGPVAGTLYLSHIHSAFCSDRPLCFTAPSGQETWSYYKVMVPLGKIGAVN 180
Query: 233 PVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGISHFVVSGVAAPST 287
PV MREN SE+YIQIVTVDGHDFWFMGFVNYDKAVKNLS+GISHFVV GVAAPST
Sbjct: 181 PVSMRENPSEKYIQIVTVDGHDFWFMGFVNYDKAVKNLSDGISHFVVPGVAAPST 345
>TC9774 similar to UP|Q9FMW4 (Q9FMW4) Similarity to ABA-responsive protein,
partial (72%)
Length = 745
Score = 148 bits (373), Expect = 1e-36
Identities = 69/179 (38%), Positives = 111/179 (61%)
Frame = +1
Query: 96 VDKPSSSPMESILHMFDSWSKKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGF 155
+ K S+L + + +KA++ A+ + +++ GP +S GK++L + + GG
Sbjct: 196 ITKSKQGKSSSVLTRMNIFGRKADSFAHGVREHVRLGPKISDTVKGKLSLGARILQVGGV 375
Query: 156 ESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVT 215
E ++KQ+F+ EKL K CYLSTT+GP+AG L++S ++FCS+R + ++P G+V
Sbjct: 376 EKVFKQLFSIKDGEKLLKASQCYLSTTSGPIAGLLFISTHKVSFCSERSIKVSSPKGEVI 555
Query: 216 WSYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGI 274
+YKV +PL KI VN + SE+YI++VT DG DFW MGF NY K+++ L + I
Sbjct: 556 RVHYKVSIPLEKIKCVNQSQNVKKPSEKYIELVTEDGFDFWLMGFFNYQKSLRCLQQAI 732
>TC15195 similar to UP|Q8S8F8 (Q8S8F8) Expressed protein, partial (39%)
Length = 623
Score = 144 bits (364), Expect = 1e-35
Identities = 66/114 (57%), Positives = 85/114 (73%)
Frame = +3
Query: 159 YKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSY 218
++Q F T P E+L +T+ACYLST+ GPV G LYLS LAFCSD PLS+ + WSY
Sbjct: 3 FRQTFQTVPEEQLVRTYACYLSTSAGPVMGVLYLSTAKLAFCSDNPLSYQTGE-ETQWSY 179
Query: 219 YKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSE 272
YKV++PL ++ VNP I R N SE+YIQI++VD H+FWFMGFV+YD AVK++ E
Sbjct: 180 YKVVIPLHQLRAVNPSISRTNPSEKYIQIISVDNHEFWFMGFVHYDSAVKHIQE 341
>AV415215
Length = 430
Score = 142 bits (359), Expect = 5e-35
Identities = 72/113 (63%), Positives = 84/113 (73%), Gaps = 7/113 (6%)
Frame = +2
Query: 1 MNNNNNNNNQNQSSPDENPFASPTP-----PSSSSSSVPLPPQNHATTENWGTHMMGAPA 55
M NN+NN QN S+ + F++P+P PS SSS+ PP N ATTENWGTH+MG PA
Sbjct: 101 MENNSNNKTQNPSAEPSSTFSNPSPSPSPSPSPSSSAPHPPPNNPATTENWGTHIMGTPA 280
Query: 56 VPSSHPDNKKAALQTASADGQGQPQVHYY--QQDHPYVQHSPVDKPSSSPMES 106
VPSSHPDNKKAALQT SA+ QPQV YY QQ HPYVQHSP +KPS+SP+ES
Sbjct: 281 VPSSHPDNKKAALQTGSAE---QPQVQYYQQQQQHPYVQHSPAEKPSNSPLES 430
>TC12434 similar to UP|Q9FMW4 (Q9FMW4) Similarity to ABA-responsive protein,
partial (27%)
Length = 655
Score = 137 bits (344), Expect = 3e-33
Identities = 69/160 (43%), Positives = 99/160 (61%)
Frame = +2
Query: 116 KKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTF 175
KK A I ++K GP +S GK++L + I EGG +++K IF EKL K
Sbjct: 2 KKRSNFACRIHEHVKLGPKLSETVKGKLSLGARIIQEGGRGNIFKHIFGMQEGEKLLKAS 181
Query: 176 ACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVI 235
CYL TT+GP+AG L++S +AFCS+RP++ T+ +G++ YKV++P+ KI VN
Sbjct: 182 QCYLYTTSGPIAGVLFISTEKVAFCSERPITCTSAAGELVRVPYKVLIPVEKIKEVNEGQ 361
Query: 236 MRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGIS 275
+YI+IVT DG +FWFMGF+ Y+KA NL + IS
Sbjct: 362 DVNQTEPKYIEIVTGDGSEFWFMGFLRYEKAFMNLQKAIS 481
>BP073959
Length = 478
Score = 113 bits (282), Expect = 4e-26
Identities = 54/108 (50%), Positives = 74/108 (68%)
Frame = -1
Query: 169 EKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKI 228
EKL K CYLSTT+GP+AG L++S +AFCS+R + ++P G + +YKV +PL KI
Sbjct: 472 EKLLKASQCYLSTTSGPIAGLLFISTDKIAFCSERSIKISSPKG*LIRVHYKVSIPLEKI 293
Query: 229 GTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGISH 276
VN + SE+YI+IV+VDG DFWFMGF NY K+++ L + I H
Sbjct: 292 KCVNQSQNVKKPSEKYIEIVSVDGFDFWFMGFFNYQKSLRCLQQAIYH 149
>TC10251 similar to UP|Q8S8F8 (Q8S8F8) Expressed protein, partial (33%)
Length = 568
Score = 100 bits (249), Expect = 3e-22
Identities = 44/96 (45%), Positives = 67/96 (68%)
Frame = +3
Query: 179 LSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRE 238
LST+ GPV G LY+S +A+ SD P+S+ + + + WSYYKV++PL ++ VNP
Sbjct: 3 LSTSAGPVMGVLYVSTAKIAYSSDNPISYKSEN-KTEWSYYKVVIPLHELKAVNPSANTA 179
Query: 239 NHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGI 274
N +E+YIQ+++ D H+FWFMGF+NY+ A + L E +
Sbjct: 180 NPAEKYIQVISTDNHEFWFMGFLNYEGAQECLEEAL 287
>TC10252 similar to UP|Q8S8F8 (Q8S8F8) Expressed protein, partial (20%)
Length = 514
Score = 62.0 bits (149), Expect = 1e-10
Identities = 25/55 (45%), Positives = 39/55 (70%)
Frame = +2
Query: 220 KVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGI 274
KV++PL ++ VNP N +E+YIQ+++ D H+FWFMGF+NY+ A + L E +
Sbjct: 86 KVVIPLHELKAVNPSANTANPAEKYIQVISTDNHEFWFMGFLNYEGAQECLEEAL 250
>CB828495
Length = 546
Score = 61.2 bits (147), Expect = 2e-10
Identities = 29/62 (46%), Positives = 40/62 (63%)
Frame = +1
Query: 199 FCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFM 258
F S+R + ++P G+V +YKV +PL KI VN + SE+YI++VT DG DFW M
Sbjct: 361 FFSERSIKVSSPKGEVIRVHYKVSIPLEKIKCVNQSQNVKKPSEKYIELVTEDGFDFWLM 540
Query: 259 GF 260
GF
Sbjct: 541 GF 546
Score = 31.6 bits (70), Expect = 0.16
Identities = 12/55 (21%), Positives = 29/55 (51%)
Frame = +3
Query: 96 VDKPSSSPMESILHMFDSWSKKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAI 150
+ K S+L + + +KA++ A+ + +++ GP +S GK++L + +
Sbjct: 201 ITKSKQGKSSSVLTRMNIFGRKADSFAHGVREHVRLGPKISDTVKGKLSLGARIL 365
>TC10682 similar to PIR|A86408|A86408 FH protein interacting protein FIP1
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (36%)
Length = 577
Score = 56.6 bits (135), Expect = 5e-09
Identities = 41/139 (29%), Positives = 65/139 (46%), Gaps = 18/139 (12%)
Frame = +2
Query: 23 PTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAV--------PSSHPDNKKAALQTASAD 74
P PP++++++ P +++ T P + P S P N + L D
Sbjct: 173 PQPPTTTTTATTTTPTKPHDSDSHSTEYAPYPKLDPNDVVPPPPSQPINTPSPL-----D 337
Query: 75 GQGQPQVHYYQQDHPYVQHSPVDKPSSSP---MESILHMFDSWSKKA-EAT------ANN 124
+G + +PYV SPV +SS ++S+ W KKA EAT A N
Sbjct: 338 ARGDAATTMPTESNPYVSPSPVPTSNSSAKNTLDSVKDALGRWGKKAAEATKKAEDLAGN 517
Query: 125 IWHNLKTGPSVSSAAMGKM 143
+W +LKTGPS + AA+G++
Sbjct: 518 MWQHLKTGPSFADAAVGRI 574
>BP085958
Length = 507
Score = 43.1 bits (100), Expect = 5e-05
Identities = 25/61 (40%), Positives = 33/61 (53%), Gaps = 1/61 (1%)
Frame = +2
Query: 2 NNNNNNNNQNQSSPDENPFASPTPPSS-SSSSVPLPPQNHATTENWGTHMMGAPAVPSSH 60
NNNNN ++N SSP +PF SP SS SSS P+PP +T + + PS +
Sbjct: 107 NNNNNRRSENGSSP*SSPFYSPPSSSSFQSSSPPIPPPCSTSTAHAPPQTNRTSSSPS*N 286
Query: 61 P 61
P
Sbjct: 287 P 289
>TC10330 similar to UP|O04344 (O04344) At2g30510 protein, partial (18%)
Length = 607
Score = 36.2 bits (82), Expect = 0.007
Identities = 19/61 (31%), Positives = 32/61 (52%)
Frame = -1
Query: 13 SSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTAS 72
S+ ++ P A+P PPSSSSS+ P++ + T + P + HP + AA ++
Sbjct: 256 STTEQRPSANPKPPSSSSSAPIAAPRSPSPTSHPSEAAPSPPRLRRRHPPHSSAATGRST 77
Query: 73 A 73
A
Sbjct: 76 A 74
>BP065897
Length = 499
Score = 33.9 bits (76), Expect = 0.033
Identities = 19/55 (34%), Positives = 28/55 (50%), Gaps = 13/55 (23%)
Frame = -2
Query: 1 MNNNNNNNNQNQSSPDENPFASPT-------------PPSSSSSSVPLPPQNHAT 42
+ +NNNNNNQN+++ +E A+PT PP+ + V QNH T
Sbjct: 267 ITHNNNNNNQNKNTTEEKSRANPTCTCTWLLCYXSRNPPTDFNPIVNKNLQNHRT 103
>TC15267 similar to UP|EXTN_TOBAC (P13983) Extensin precursor (Cell wall
hydroxyproline-rich glycoprotein), partial (6%)
Length = 1003
Score = 33.9 bits (76), Expect = 0.033
Identities = 27/99 (27%), Positives = 37/99 (37%), Gaps = 3/99 (3%)
Frame = +1
Query: 13 SSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTAS 72
SS +P+ SP+PPSS++ P PP +H T N P+ PS+ + S
Sbjct: 166 SSSSLSPWLSPSPPSSTT---PPPPPSHRTNNN-NLSTQPTPSAPSATSLASRTPASPPS 333
Query: 73 ADGQGQPQVHYYQQDHPYV---QHSPVDKPSSSPMESIL 108
P H P SP P S +L
Sbjct: 334 PPPTTTPPTHNPSSPSPSAPPWTSSPPSPPRSESRPQVL 450
Score = 28.1 bits (61), Expect = 1.8
Identities = 23/82 (28%), Positives = 31/82 (37%), Gaps = 4/82 (4%)
Frame = +1
Query: 2 NNNNNNNN----QNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVP 57
++ NNNN S+P AS TP S S PP T N + AP
Sbjct: 235 SHRTNNNNLSTQPTPSAPSATSLASRTPASPPSP----PPTTTPPTHNPSSPSPSAPPWT 402
Query: 58 SSHPDNKKAALQTASADGQGQP 79
SS P ++ + +G P
Sbjct: 403 SSPPSPPRSESRPQVLTARGLP 468
>AV414308
Length = 389
Score = 33.5 bits (75), Expect = 0.043
Identities = 18/62 (29%), Positives = 27/62 (43%)
Frame = +3
Query: 2 NNNNNNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHP 61
+ +N + Q+ P P ASP+ P S PP +HA + +W T PS P
Sbjct: 132 STTSNKESLLQNPPSPPPEASPSSPGPGSP----PPPHHAPSSSWSTATATTSPGPSKPP 299
Query: 62 DN 63
+
Sbjct: 300 QS 305
>TC11441 similar to PIR|S54157|S54157 extensin-like protein - cowpea
(fragment) {Vigna unguiculata;} , partial (6%)
Length = 541
Score = 33.1 bits (74), Expect = 0.056
Identities = 21/65 (32%), Positives = 29/65 (44%), Gaps = 3/65 (4%)
Frame = +2
Query: 6 NNNNQNQSSPDENPFASPTPPSSSSSSVPLP---PQNHATTENWGTHMMGAPAVPSSHPD 62
+ +N N S+ ++ ASP P SSSSS P P Q + T+ +PA HP
Sbjct: 17 DRSNSNSSNTNQTTTASPPSPQSSSSSPPSPAPSKQEGQSPPPADTNTTPSPASDHDHPS 196
Query: 63 NKKAA 67
A
Sbjct: 197PPNGA 211
>AV411486
Length = 419
Score = 32.7 bits (73), Expect = 0.073
Identities = 14/34 (41%), Positives = 20/34 (58%)
Frame = +3
Query: 13 SSPDENPFASPTPPSSSSSSVPLPPQNHATTENW 46
+ PD+NPF +P ++PLPP+ H TT W
Sbjct: 180 TKPDKNPFTLLSP--MEGFTLPLPPRPHTTTLQW 275
>TC11169 similar to UP|Q9SL72 (Q9SL72) At2g20090 protein, partial (14%)
Length = 650
Score = 32.7 bits (73), Expect = 0.073
Identities = 22/59 (37%), Positives = 27/59 (45%), Gaps = 9/59 (15%)
Frame = +3
Query: 2 NNNNNNNNQNQSSPDENPFASPTP-------PSSSSSSVPLP--PQNHATTENWGTHMM 51
N N N+ P PF PTP P SSSSS+P P P+N E+W +M
Sbjct: 96 NININSMRPPSPQPQAPPFL-PTPSNFLTPYPPSSSSSLPFPSWPENQEVPESWSQLLM 269
>TC14852 similar to UP|Q9LZI2 (Q9LZI2) DTDP-glucose 4-6-dehydratase homolog
D18 (At3g62830/F26K9_260), partial (70%)
Length = 1470
Score = 32.7 bits (73), Expect = 0.073
Identities = 17/31 (54%), Positives = 22/31 (70%)
Frame = +1
Query: 7 NNNQNQSSPDENPFASPTPPSSSSSSVPLPP 37
+++ + SSP ASP PPSSS SS+PLPP
Sbjct: 544 SSDSSSSSP-----ASP*PPSSSPSSLPLPP 621
>TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precursor (Atperox
P31) (ATP41) , partial (38%)
Length = 475
Score = 32.3 bits (72), Expect = 0.096
Identities = 26/103 (25%), Positives = 46/103 (44%), Gaps = 2/103 (1%)
Frame = +1
Query: 3 NNNNNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPD 62
+N+ ++ S +++P P PP S+SSS A+T T +P +PS+ P+
Sbjct: 139 HNSPKSSATPSPTNKSPPPPPAPPRSASSSTT------ASTPTAATPPSSSPQLPSTKPN 300
Query: 63 NKKAALQTASADGQGQPQVHYYQQDHP--YVQHSPVDKPSSSP 103
A T+++ P + P + +P P+SSP
Sbjct: 301 ----ATPTSTSPSPATPSTSSSELKPPSNSLAPTPSPAPTSSP 417
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.127 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,959,870
Number of Sequences: 28460
Number of extensions: 102415
Number of successful extensions: 1988
Number of sequences better than 10.0: 269
Number of HSP's better than 10.0 without gapping: 1432
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1752
length of query: 287
length of database: 4,897,600
effective HSP length: 89
effective length of query: 198
effective length of database: 2,364,660
effective search space: 468202680
effective search space used: 468202680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC133780.8


