

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133779.1 + phase: 0 /pseudo
(611 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
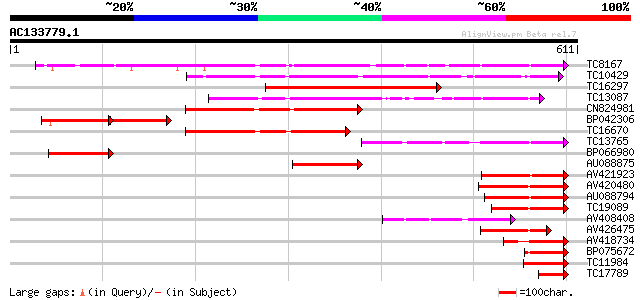
Score E
Sequences producing significant alignments: (bits) Value
TC8167 similar to PIR|JC7519|JC7519 subtilisin-like serine prote... 310 4e-85
TC10429 similar to UP|Q9ZR46 (Q9ZR46) P69C protein, partial (52%) 263 8e-71
TC16297 weakly similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type pro... 244 4e-65
TC13087 similar to UP|Q9FIF8 (Q9FIF8) Serine protease-like prote... 202 2e-52
CN824981 172 2e-43
BP042306 129 4e-41
TC16670 similar to PIR|JC7519|JC7519 subtilisin-like serine prot... 155 1e-38
TC13765 similar to UP|Q9LLL8 (Q9LLL8) Subtilisin-type serine end... 130 6e-31
BP066980 125 2e-29
AU088875 116 9e-27
AV421923 113 8e-26
AV420480 110 9e-25
AU088794 108 3e-24
TC19089 similar to UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C... 100 9e-22
AV408408 96 2e-20
AV426475 86 1e-17
AV418734 80 1e-15
BP075672 69 3e-12
TC11984 similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease, ... 59 3e-09
TC17789 similar to PIR|T05768|T05768 subtilisin-like proteinase ... 54 1e-07
>TC8167 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (65%)
Length = 2760
Score = 310 bits (794), Expect = 4e-85
Identities = 224/610 (36%), Positives = 321/610 (51%), Gaps = 35/610 (5%)
Frame = +3
Query: 28 INFISFWFLVSYLFLQC-----------YIVYLGAHVHGPT-PSSVDLETATYSHYDLLG 75
I+ +SF F++ L LQC YIV++ H P+ D TA+
Sbjct: 72 ISTLSFTFIL--LLLQCLSASCSSPKKTYIVHMNHHTKPQIYPTRRDWYTASLRSL---- 233
Query: 76 SILGSHEEAEEAIIYSYNKQINGFAAILEEEEAAQLANCTQHVLGNSLD*VQMM------ 129
S+ E +++ ++Y+Y+ NGFAA L+E++A L + VLG D + +
Sbjct: 234 SLTTDSETSDDPLLYAYDTAYNGFAASLDEQQAQTLLG-SDSVLGLYEDTLYHLHTTRTP 410
Query: 130 -----STQLGKREGLVKIPL*LTLIQVFGRNRRVLTTEE*VQFH*DGVGILLRKF----- 179
TQ G EG + L V E F+ G+ + ++
Sbjct: 411 QFLGLDTQTGLWEGHRTLELDQASRDVIIGVLDTGVWPESPSFNDAGMPEIPSRWRGECE 590
Query: 180 HVTVYLNGL-PRKLIGARFFNKAYEAFHGK-----LPSSQQTARDFVGPGTHTLSTAGGN 233
+ T + + L RKLIGAR F++ + G + RD G GTHT STA G+
Sbjct: 591 NATDFSSSLCNRKLIGARSFSRGFHMAAGNDGGFGKEREPPSPRDSDGHGTHTASTAAGS 770
Query: 234 FVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLIS 293
V NA++ G +GT +G +P++RVATYK CWS CF +D+LA +D+AI DG D++S
Sbjct: 771 HVGNASLLGYASGTARGMAPQARVATYKVCWS----DGCFASDILAGMDRAIRDGVDVLS 938
Query: 294 VSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAAS 353
+S GG + P F D I+IGAF A+ R I + SAGN GP+ S+ NVAPW+ TV A
Sbjct: 939 LSLGG--GSGP--YFRDTIAIGAFAAMERGIFVSCSAGNSGPSKASLANVAPWLMTVGAG 1106
Query: 354 TLDRDF-SSVMTINNKTLTGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLD 412
TLDRDF +S + N K G SL+ + + ++ ++ ++ C PG+LD
Sbjct: 1107TLDRDFPASALLGNKKRFAGVSLYSGKGMGAEPVGLV-----YSKGSNQSGILCLPGSLD 1271
Query: 413 PSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYY 472
P+ V GKVV CDR G + +G+ AG +G+I+ N +G+ L+A+ H++ +
Sbjct: 1272PAVVRGKVVLCDR-GLNARVEKGKVVKEAGGIGMILTN-TAANGEELVADSHLLPAV--- 1436
Query: 473 DARSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPD 532
I + E D A + S LN R P+PV+A+FSSRGPN + ILKPD
Sbjct: 1437AVGRIVGDQIREYVTSDPNPTAVLSFS-GTVLNVR-PSPVVAAFSSRGPNMITKQILKPD 1610
Query: 533 VTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSP 592
V PGVNILA +S S L D+R+ FNI GTSMSCPH+ G L+K HP+WSP
Sbjct: 1611VIGPGVNILAGWSEAIGPSGLPQDSRKS-QFNIMSGTSMSCPHISGLGALLKAAHPDWSP 1787
Query: 593 AAIKSAIMTT 602
+AIKSA+MTT
Sbjct: 1788SAIKSALMTT 1817
>TC10429 similar to UP|Q9ZR46 (Q9ZR46) P69C protein, partial (52%)
Length = 1742
Score = 263 bits (671), Expect = 8e-71
Identities = 170/409 (41%), Positives = 223/409 (53%), Gaps = 3/409 (0%)
Frame = +3
Query: 191 KLIGARFFNKAYEAFHGKLPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKG 250
KLIGAR FN A EA +GK + D G GTHT STA G FV A + G GT G
Sbjct: 597 KLIGARAFNLAAEAMNGK---KAEAPIDEDGHGTHTASTAAGAFVNYAEVLGNAKGTAAG 767
Query: 251 GSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTD 310
+P + +A YK C+ DC +D+LAA+D A+ DG D+IS+S G + P F D
Sbjct: 768 MAPHAHLAIYKVCFG----EDCPESDILAALDAAVEDGVDVISISLG---LSEPPPFFND 926
Query: 311 EISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTI-NNKT 369
+IGAF A+ + I + +AGN GP S+ N APW+ TV AST+DR + + N +
Sbjct: 927 STAIGAFAAMQKGIFVSCAAGNSGPFNSSIVNAAPWILTVGASTIDRRIVATAKLGNGQE 1106
Query: 370 LTGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKI 429
G S+F F + A ++ FC G+LD S GKVV C+R G I
Sbjct: 1107FDGESVF----QPSSFTPTLLPLAYAGKNGKEESAFCANGSLDDSAFRGKVVLCERGGGI 1274
Query: 430 NSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVV--STINYYDARSITTPKGSEITP 487
IA+G+E AG +I+ N E + +L A+ H + + ++Y I S TP
Sbjct: 1275ARIAKGEEVKRAGGAAMILMND-ETNAFSLSADVHALPATHVSYAAGIEIKAYINSTATP 1451
Query: 488 EDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLL 547
ATI + G AP +ASFSSRGPN P ILKPD+ PGVNILAA+
Sbjct: 1452-----TATILFK--GTVIGNSLAPAVASFSSRGPNLPSPGILKPDIIGPGVNILAAWPFP 1610
Query: 548 ASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIK 596
S S TD++ FNI+ GTSMSCPH+ G A L+K+ HP+WSPAAIK
Sbjct: 1611LSNS---TDSK--LTFNIESGTSMSCPHLSGIAALLKSSHPHWSPAAIK 1742
Score = 32.3 bits (72), Expect = 0.23
Identities = 25/60 (41%), Positives = 31/60 (51%), Gaps = 4/60 (6%)
Frame = +3
Query: 52 HVHGPT----PSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAILEEEE 107
HV GP S DLE+ +S L L S EE + +IYSY + GFAA L +EE
Sbjct: 162 HVTGPEGKMLTESEDLESWYHS---FLPPTLKSSEE-QPRVIYSYKNVLRGFAASLTQEE 329
>TC16297 weakly similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease,
partial (10%)
Length = 574
Score = 244 bits (622), Expect = 4e-65
Identities = 126/191 (65%), Positives = 150/191 (77%), Gaps = 1/191 (0%)
Frame = +2
Query: 276 DVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGP 335
DVLAAIDQAI DG D+ SVSAGG+ + E IFTDE+SIG+ HALA+NIL+VASAGN+GP
Sbjct: 2 DVLAAIDQAITDGVDVNSVSAGGRSIISAEEIFTDEVSIGSVHALAKNILVVASAGNDGP 181
Query: 336 TPGSVTNVAPWVFTVAASTLDRDFSSVMTI-NNKTLTGASLFVNLPPNQDFLIIISTDAK 394
G+V NVAPWVFTVAAST+DR+FSS +T+ NN+ +TGASLFVNLPPNQ F +I+STDAK
Sbjct: 182 ALGTVLNVAPWVFTVAASTIDREFSSTITLGNNQQITGASLFVNLPPNQLFSLILSTDAK 361
Query: 395 FANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEV 454
AN T+ DAQ CRPGTLDP+KV GKVV C R GKI S+AEG EALS+GA G+++
Sbjct: 362 LANATNRDAQLCRPGTLDPAKVKGKVVVCTRGGKIKSVAEGSEALSSGAKGMVLE*SKAK 541
Query: 455 DGKTLLAEPHV 465
EPHV
Sbjct: 542 WKTHFFLEPHV 574
>TC13087 similar to UP|Q9FIF8 (Q9FIF8) Serine protease-like protein, partial
(31%)
Length = 1036
Score = 202 bits (513), Expect = 2e-52
Identities = 142/368 (38%), Positives = 201/368 (54%), Gaps = 6/368 (1%)
Frame = +2
Query: 215 TARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCFG 274
T RD G G+HT STA GN V A+ +G+ GT +G P +R+A YK C+ C
Sbjct: 20 TVRDIEGHGSHTASTAAGNTVIGASFYGLAQGTARGAVPSARIAAYKVCYPNHG---CDD 190
Query: 275 ADVLAAIDQAIYDGADLISVSAGGKPNTNPEV-IFTDEISIGAFHALARNILLVASAGNE 333
A++LAA D AI DG +++SVS G TN V + +D ++IG+FHA+ R IL + +AGN
Sbjct: 191 ANILAAFDDAIADGVNILSVSLG----TNYCVDLLSDTVAIGSFHAMERGILTLQAAGNS 358
Query: 334 GPTPGSVTNVAPWVFTVAASTLDRDF-SSVMTINNKTLTGASLFVNLPPNQDFLII--IS 390
GP S+ +VAPW+ TVAAST+DR F V+ N TL G S+ + F I +
Sbjct: 359 GPASPSICSVAPWLLTVAASTIDRQFIDKVLLGNGMTLIGRSVNAVVSNGTKFPIAKWNA 538
Query: 391 TDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRN 450
+D + +D+ C+ L V GKVV C EG+++ + A GAVG I +
Sbjct: 539 SDERCKGFSDL----CQ--CLGSESVKGKVVLC--EGRMHD----EAAYLDGAVGSITED 682
Query: 451 QPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEITPE--DIKTNATIRMSPANALNGRK 508
D VS + Y+ + ++ PK I + N + + R
Sbjct: 683 LTNPDND--------VSFVTYFPSVNL-NPKDYAIVQNYTNSAKNPKAEILKSEIFKDRS 835
Query: 509 PAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQG 568
AP + SFSSRGPN + P I+KPDV+APGV+ILAA+S A+ S +D R+ +N+ G
Sbjct: 836 -APKVVSFSSRGPNPLIPEIMKPDVSAPGVDILAAWSPEAAPSEYSSDKRK-VRYNVVSG 1009
Query: 569 TSMSCPHV 576
TSM+CPHV
Sbjct: 1010TSMACPHV 1033
>CN824981
Length = 726
Score = 172 bits (435), Expect = 2e-43
Identities = 95/194 (48%), Positives = 129/194 (65%), Gaps = 3/194 (1%)
Frame = +1
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQQT--ARDFVGPGTHTLSTAGGNFVQNATIFGIGNGT 247
RKLIGARFF+K YEA G + S ++ ARD G G+HTL+TA G+ V A++FG+ +GT
Sbjct: 115 RKLIGARFFSKGYEATLGPIDVSTESRSARDDDGHGSHTLTTAAGSAVAGASLFGLASGT 294
Query: 248 IKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVI 307
+G + ++RVA YK CW + CF +D+ A ID+AI DG ++IS+S GG
Sbjct: 295 ARGMATQARVAAYKVCW----LGGCFSSDIAAGIDKAIEDGVNIISMSIGGSSAD----Y 450
Query: 308 FTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTI-N 366
F D I+IGAF A + IL+ SAGN GP+P S++N APW+ TV A T+DRDF + +T+ N
Sbjct: 451 FRDIIAIGAFTANSHGILVSTSAGNGGPSPSSLSNTAPWITTVGAGTIDRDFPAYITLGN 630
Query: 367 NKTLTGASLFVNLP 380
N T TGASL+ P
Sbjct: 631 NITHTGASLYRGKP 672
>BP042306
Length = 549
Score = 129 bits (324), Expect(2) = 4e-41
Identities = 65/90 (72%), Positives = 73/90 (80%), Gaps = 12/90 (13%)
Frame = +3
Query: 35 FLVSYLFL------------QCYIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHE 82
FLVSY L +CYIVYLGAH HGPTPSSVDLETAT SHYDLLGS+LGSHE
Sbjct: 45 FLVSYFLLFISLLNSVHASKKCYIVYLGAHSHGPTPSSVDLETATSSHYDLLGSVLGSHE 224
Query: 83 EAEEAIIYSYNKQINGFAAILEEEEAAQLA 112
+A+EAI+YSYNK INGFAA+LEEEEAA++A
Sbjct: 225 KAKEAILYSYNKHINGFAALLEEEEAARIA 314
Score = 56.2 bits (134), Expect(2) = 4e-41
Identities = 35/66 (53%), Positives = 42/66 (63%)
Frame = +1
Query: 109 AQLANCTQHVLGNSLD*VQMMSTQLGKREGLVKIPL*LTLIQVFGRNRRVLTTEE*VQFH 168
A+ NCT V G+ LD +M+ Q GKR L KI *LT IQVFG N RVL T+E V F
Sbjct: 346 ARSINCTLPVHGSFLDCEEMVGIQHGKRGDLEKIQS*LTSIQVFGLNPRVLVTKELVLFL 525
Query: 169 *DGVGI 174
+GVG+
Sbjct: 526 QNGVGV 543
>TC16670 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (42%)
Length = 1183
Score = 155 bits (393), Expect = 1e-38
Identities = 82/180 (45%), Positives = 116/180 (63%), Gaps = 2/180 (1%)
Frame = +2
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQQTA--RDFVGPGTHTLSTAGGNFVQNATIFGIGNGT 247
+KLIGAR+F+K EA G + + ++ RD G GTHT STA G+ V A++FG GT
Sbjct: 668 KKLIGARYFSKGAEAMLGPIDETAESKSPRDDDGHGTHTASTAAGSVVSGASLFGYAAGT 847
Query: 248 IKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVI 307
+G +P +RVA YK CW CF +D+LAAID+AI D +++S+S GG
Sbjct: 848 ARGMAPHARVAAYKVCWK----GGCFSSDILAAIDKAIADNVNVLSLSLGG----GMADY 1003
Query: 308 FTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTINN 367
+ D ++IGAF A+ + IL+ SAGN GP+ S++NVAPW+ TV A TLDRDF +++ + N
Sbjct: 1004YRDSVAIGAFAAMEKGILVSCSAGNAGPSAYSLSNVAPWITTVGAGTLDRDFPALVELGN 1183
>TC13765 similar to UP|Q9LLL8 (Q9LLL8) Subtilisin-type serine endopeptidase
XSP1, partial (20%)
Length = 635
Score = 130 bits (327), Expect = 6e-31
Identities = 80/225 (35%), Positives = 121/225 (53%), Gaps = 2/225 (0%)
Frame = +1
Query: 380 PPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEAL 439
P +++ +I DA + + DA FC +L P+KV GK+V C K+ +
Sbjct: 1 PKRKEYSLINGIDAAKDSKSKDDAGFCYEDSLQPNKVKGKLVYC----KLGNWGTEGVVK 168
Query: 440 SAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEITPEDI--KTNATIR 497
G +G IM + D + P + +N+ S+T S +P + KT+
Sbjct: 169 KFGGIGSIMESDQYPDLAQIFMAPATI--LNHTIGESVTNYIKSTRSPSAVIYKTHEE-- 336
Query: 498 MSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDN 557
+ PAP +A+FSSRGPN +LKPD+ APG++ILA+Y+L S++ D
Sbjct: 337 ---------KCPAPFVATFSSRGPNPGSHNVLKPDIAAPGIDILASYTLRKSITGSEGDT 489
Query: 558 RRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
+ F++ GTSM+CPHV G A +K+ HPNW+PAAI+SAI+TT
Sbjct: 490 QFS-EFSLLSGTSMACPHVAGVAAYVKSFHPNWTPAAIRSAIITT 621
>BP066980
Length = 472
Score = 125 bits (314), Expect = 2e-29
Identities = 58/69 (84%), Positives = 66/69 (95%)
Frame = +3
Query: 43 QCYIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAI 102
+CYIVYLGAH HGPTPSSVDLETAT SHYDLLGS+LGSHE+A+EAI+YSYNK INGFAA+
Sbjct: 162 KCYIVYLGAHSHGPTPSSVDLETATSSHYDLLGSVLGSHEKAKEAILYSYNKHINGFAAL 341
Query: 103 LEEEEAAQL 111
LEEEEAA++
Sbjct: 342 LEEEEAARI 368
>AU088875
Length = 322
Score = 116 bits (291), Expect = 9e-27
Identities = 57/77 (74%), Positives = 69/77 (89%), Gaps = 1/77 (1%)
Frame = +3
Query: 305 EVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMT 364
E IFTDE+SIG+FHALA+NIL+VASAGN+GP G+V NVAPWVFTVAAST+D +FSS +T
Sbjct: 3 EEIFTDEVSIGSFHALAKNILVVASAGNDGPALGTVLNVAPWVFTVAASTIDXEFSSTIT 182
Query: 365 I-NNKTLTGASLFVNLP 380
+ NN+ +TGASLFVNLP
Sbjct: 183 LGNNQQITGASLFVNLP 233
>AV421923
Length = 463
Score = 113 bits (283), Expect = 8e-26
Identities = 59/94 (62%), Positives = 72/94 (75%)
Frame = +2
Query: 509 PAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQG 568
PAPV+A+FSSRGP+ V ++KPDVTAPGV+ILAA+ S S L D RR FNI G
Sbjct: 50 PAPVVAAFSSRGPSIVGLDVIKPDVTAPGVHILAAWPSKTSPSMLKNDKRRVL-FNIISG 226
Query: 569 TSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
TSMSCPHV G A LI+++H +WSPAA+KSA+MTT
Sbjct: 227 TSMSCPHVSGVAALIRSVHRDWSPAAVKSALMTT 328
>AV420480
Length = 412
Score = 110 bits (274), Expect = 9e-25
Identities = 56/97 (57%), Positives = 68/97 (69%)
Frame = +3
Query: 506 GRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNI 565
G AP +A+FS+RGP+ P ILKPDV APGVNI+AA+ ++L D RR F++
Sbjct: 18 GNSRAPAVATFSARGPSFTNPSILKPDVVAPGVNIIAAWPQNLGPTSLPQDLRR-VNFSV 194
Query: 566 QQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
GTSMSCPHV G A L+ + HP WSPAAIKSAIMTT
Sbjct: 195 MSGTSMSCPHVSGIAALVHSAHPKWSPAAIKSAIMTT 305
>AU088794
Length = 433
Score = 108 bits (269), Expect = 3e-24
Identities = 56/91 (61%), Positives = 65/91 (70%)
Frame = +2
Query: 512 VMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSM 571
V+ASFS+RGPN P ILKPDV PG+NILAA+ S + +D RR FNI GTSM
Sbjct: 2 VVASFSARGPNPESPEILKPDVIXPGLNILAAWPDXXGPSGVPSDVRRT-EFNILSGTSM 178
Query: 572 SCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
+CPHV G A L+K HP WSPAAIKSA+MTT
Sbjct: 179 ACPHVSGLAALLKAAHPXWSPAAIKSALMTT 271
>TC19089 similar to UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C1, partial
(16%)
Length = 600
Score = 100 bits (248), Expect = 9e-22
Identities = 46/83 (55%), Positives = 58/83 (69%)
Frame = +1
Query: 520 GPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGT 579
GPN + P ILKPD+ APGV +LAA+S L +S + D R+ P+N GTSM+CPH
Sbjct: 16 GPNPITPNILKPDIAAPGVTVLAAWSPLNPISEVEGDKRK-LPYNAISGTSMACPHAAAA 192
Query: 580 AGLIKTLHPNWSPAAIKSAIMTT 602
A +K+ HPNWSPA IKSA+MTT
Sbjct: 193 AAYVKSFHPNWSPAMIKSALMTT 261
>AV408408
Length = 429
Score = 95.9 bits (237), Expect = 2e-20
Identities = 56/144 (38%), Positives = 82/144 (56%)
Frame = +3
Query: 402 DAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLA 461
+ C GTL P KV GK+V CDR G + +GQ AG +G+++ N +G+ L+A
Sbjct: 18 NGNLCMMGTLSPDKVAGKIVMCDR-GMNARVQKGQVVKGAGGLGMVLSNTA-ANGEELVA 191
Query: 462 EPHVVSTINYYDARSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGP 521
+ H++ T + K P+ AT+++ G +P+PV+A+FSSRGP
Sbjct: 192 DAHLLPTSAVGEKAGNAIKKYLFSDPK-----ATVKILFEGTKVGIQPSPVVAAFSSRGP 356
Query: 522 NKVQPYILKPDVTAPGVNILAAYS 545
N + P ILKPD+ APGVNILA +S
Sbjct: 357 NSITPQILKPDLIAPGVNILAGWS 428
>AV426475
Length = 300
Score = 86.3 bits (212), Expect = 1e-17
Identities = 44/77 (57%), Positives = 54/77 (69%)
Frame = +2
Query: 508 KPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQ 567
KP+PV+A+FSSRGPN + P ILKPD+ APGVNILA ++ + L D R FNI
Sbjct: 71 KPSPVVAAFSSRGPNGLTPKILKPDLIAPGVNILAGWTGAIGPTGLPVDTRH-VSFNIIS 247
Query: 568 GTSMSCPHVVGTAGLIK 584
GTSMSCPHV G A ++K
Sbjct: 248 GTSMSCPHVSGLAAILK 298
>AV418734
Length = 287
Score = 80.1 bits (196), Expect = 1e-15
Identities = 42/70 (60%), Positives = 49/70 (70%)
Frame = +1
Query: 533 VTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSP 592
VTAPG+NILAA+S A G FNI GTSM+CPHV G A L+K +HP+WSP
Sbjct: 4 VTAPGLNILAAWSPAA-----------GNMFNIVSGTSMACPHVTGIATLVKAVHPSWSP 150
Query: 593 AAIKSAIMTT 602
+AIKSAIMTT
Sbjct: 151 SAIKSAIMTT 180
>BP075672
Length = 437
Score = 68.6 bits (166), Expect = 3e-12
Identities = 33/48 (68%), Positives = 37/48 (76%)
Frame = -2
Query: 555 TDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
+D+RR P N GTSM+CPHV G GL+ TLHP WSPAAIKSAIMTT
Sbjct: 436 SDDRR-VPLNSLTGTSMACPHVSGIVGLLHTLHPEWSPAAIKSAIMTT 296
>TC11984 similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease, partial
(30%)
Length = 966
Score = 58.5 bits (140), Expect = 3e-09
Identities = 28/49 (57%), Positives = 34/49 (69%)
Frame = +1
Query: 554 VTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
V ++ +NI GTSMSCPHV G AG IK+ H WS +AIKSAIMT+
Sbjct: 31 VPKGKKPSQYNIISGTSMSCPHVSGLAGSIKSRHTTWSASAIKSAIMTS 177
>TC17789 similar to PIR|T05768|T05768 subtilisin-like proteinase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (14%)
Length = 940
Score = 53.5 bits (127), Expect = 1e-07
Identities = 20/32 (62%), Positives = 28/32 (87%)
Frame = +2
Query: 571 MSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
M+CPH+ G A L+K+ HP+WSPAA++SA+MTT
Sbjct: 2 MACPHISGAAALLKSAHPDWSPAAVRSAMMTT 97
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,606,098
Number of Sequences: 28460
Number of extensions: 125691
Number of successful extensions: 680
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 642
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 647
length of query: 611
length of database: 4,897,600
effective HSP length: 96
effective length of query: 515
effective length of database: 2,165,440
effective search space: 1115201600
effective search space used: 1115201600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC133779.1


