

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133571.7 + phase: 0
(79 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
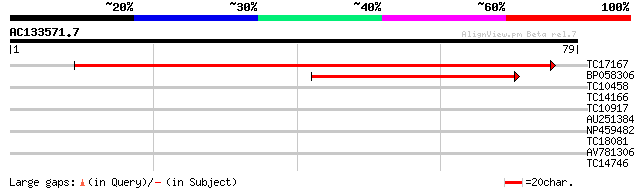
TC17167 similar to UP|SDF2_ARATH (Q93ZE8) Stromal cell-derived f... 132 1e-32
BP058306 53 1e-08
TC10458 26 1.2
TC14166 weakly similar to UP|Q9U929 (Q9U929) Splicing factor pTS... 25 2.6
TC10917 similar to UP|Q7M4A3 (Q7M4A3) Spermatid-specific protein... 24 4.4
AU251384 24 4.4
NP459482 RAB11J [Lotus japonicus] 23 7.6
TC18081 similar to UP|O81249 (O81249) Globulin-1 (Fragment), par... 23 7.6
AV781306 23 7.6
TC14746 weakly similar to GB|AAG21976.1|10716947|AF263377 SKP1 i... 23 9.9
>TC17167 similar to UP|SDF2_ARATH (Q93ZE8) Stromal cell-derived factor
2-like protein precursor (SDF2-like protein), partial
(47%)
Length = 588
Score = 132 bits (332), Expect = 1e-32
Identities = 61/67 (91%), Positives = 63/67 (93%)
Frame = +1
Query: 10 EGSGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY 69
EGSGKTWKQDQ+ RLQHIDT GY +SHDKKYSRIAGGQQEVC VREKRADNVWLAAEGVY
Sbjct: 109 EGSGKTWKQDQKIRLQHIDTGGYLHSHDKKYSRIAGGQQEVCAVREKRADNVWLAAEGVY 288
Query: 70 LPVTESK 76
LPVTESK
Sbjct: 289 LPVTESK 309
>BP058306
Length = 357
Score = 52.8 bits (125), Expect = 1e-08
Identities = 24/29 (82%), Positives = 25/29 (85%)
Frame = -1
Query: 43 IAGGQQEVCGVREKRADNVWLAAEGVYLP 71
I+GG EVC VREKRAD VWLAAEGVYLP
Sbjct: 357 ISGGPPEVCPVREKRADQVWLAAEGVYLP 271
>TC10458
Length = 526
Score = 26.2 bits (56), Expect = 1.2
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = +1
Query: 59 DNVWLAAEGVYLPVTESK 76
+NVWL GVY+ +T SK
Sbjct: 118 ENVWLVEGGVYIQMTASK 171
>TC14166 weakly similar to UP|Q9U929 (Q9U929) Splicing factor pTSR1, partial
(7%)
Length = 1473
Score = 25.0 bits (53), Expect = 2.6
Identities = 11/30 (36%), Positives = 16/30 (52%)
Frame = +3
Query: 45 GGQQEVCGVREKRADNVWLAAEGVYLPVTE 74
GGQ +V V RA+ A G+ +PV +
Sbjct: 1134 GGQNKVAAVEAGRAEETQAMASGLAIPVID 1223
>TC10917 similar to UP|Q7M4A3 (Q7M4A3) Spermatid-specific protein T2
precursor, partial (21%)
Length = 589
Score = 24.3 bits (51), Expect = 4.4
Identities = 13/42 (30%), Positives = 18/42 (41%)
Frame = -3
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGV 53
+ K WK F LQ + G+F AGGQ + G+
Sbjct: 194 TSKNWKASLSFLLQETNVFGFFG---------AGGQFRIAGI 96
>AU251384
Length = 364
Score = 24.3 bits (51), Expect = 4.4
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = +2
Query: 35 SHDKKYSRIAGGQQEVCGVREKRADNVWLAAE 66
SH++ +S G QQ V G+ R DN+ + E
Sbjct: 263 SHERVFSTKEGVQQIVLGLYIIRGDNISVVGE 358
>NP459482 RAB11J [Lotus japonicus]
Length = 672
Score = 23.5 bits (49), Expect = 7.6
Identities = 12/45 (26%), Positives = 21/45 (46%)
Frame = +3
Query: 24 LQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGV 68
LQH + GY S + A + E CG +++ + W +G+
Sbjct: 525 LQHSKSEGYDVSRAHQT*SSAD*KWEECGYTRRKSRSRWAVKKGL 659
>TC18081 similar to UP|O81249 (O81249) Globulin-1 (Fragment), partial (5%)
Length = 662
Score = 23.5 bits (49), Expect = 7.6
Identities = 13/33 (39%), Positives = 16/33 (48%)
Frame = -3
Query: 45 GGQQEVCGVREKRADNVWLAAEGVYLPVTESKQ 77
G QQ+ G EK A +W V TES+Q
Sbjct: 231 GNQQQQRGEPEKLAATIWQKQGTVATSKTESEQ 133
>AV781306
Length = 525
Score = 23.5 bits (49), Expect = 7.6
Identities = 10/30 (33%), Positives = 17/30 (56%), Gaps = 3/30 (10%)
Frame = +2
Query: 37 DKKYSRIAGGQQEVCGVREKRADN---VWL 63
D YS+++GG + C +R D+ +WL
Sbjct: 431 DATYSKLSGGMAKKCKMRSYIGDSETFIWL 520
>TC14746 weakly similar to GB|AAG21976.1|10716947|AF263377 SKP1
interacting partner 1 {Arabidopsis thaliana;}, partial
(62%)
Length = 719
Score = 23.1 bits (48), Expect = 9.9
Identities = 10/26 (38%), Positives = 18/26 (68%)
Frame = +2
Query: 11 GSGKTWKQDQRFRLQHIDTSGYFYSH 36
G+GK ++ +RFR++ +D SG + H
Sbjct: 65 GNGKRGRRRKRFRVERVD-SGVPHQH 139
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.131 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,461,677
Number of Sequences: 28460
Number of extensions: 15155
Number of successful extensions: 45
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 45
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 45
length of query: 79
length of database: 4,897,600
effective HSP length: 55
effective length of query: 24
effective length of database: 3,332,300
effective search space: 79975200
effective search space used: 79975200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 48 (23.1 bits)
Medicago: description of AC133571.7


