

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133341.5 - phase: 1 /pseudo
(808 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
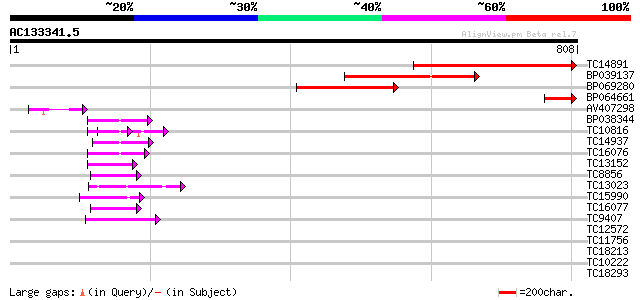
Score E
Sequences producing significant alignments: (bits) Value
TC14891 448 e-126
BP039137 264 5e-71
BP069280 181 4e-46
BP064661 90 1e-18
AV407298 82 3e-16
BP038344 61 8e-10
TC10816 homologue to UP|CAE45585 (CAE45585) Coatomer alpha subun... 48 5e-06
TC14937 similar to UP|Q9FN19 (Q9FN19) Genomic DNA, chromosome 5,... 47 2e-05
TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG... 47 2e-05
TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial... 45 6e-05
TC8856 similar to UP|Q8W403 (Q8W403) Sec13p, partial (48%) 44 8e-05
TC13023 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Fragm... 44 8e-05
TC15990 44 1e-04
TC16077 similar to PIR|T00593|T02480 sec13-related protein At2g3... 42 3e-04
TC9407 weakly similar to UP|TAF5_YEAST (P38129) Transcription in... 41 7e-04
TC12572 similar to PIR|T46032|T46032 WD-40 repeat regulatory pro... 40 0.001
TC11756 weakly similar to GB|AAH01494.1|16306637|BC001494 U5 snR... 38 0.006
TC18213 weakly similar to UP|Q9FIV2 (Q9FIV2) Similarity to pre-m... 38 0.007
TC10222 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Ar... 37 0.016
TC18293 similar to UP|PWP1_HUMAN (Q13610) Periodic tryptophan pr... 35 0.036
>TC14891
Length = 1172
Score = 448 bits (1152), Expect = e-126
Identities = 216/232 (93%), Positives = 223/232 (96%)
Frame = +3
Query: 576 ELPCTVKLRVWSHDIKNPCSPLNADRCRLIIPHAVLCSEMGAHFSPCGRFLAACVACMLP 635
ELPCTVKLRVWSHDIKNPC+PLNAD+CRL IPHAVLCSEMGAHFSPCGRFLAACVACMLP
Sbjct: 3 ELPCTVKLRVWSHDIKNPCAPLNADKCRLTIPHAVLCSEMGAHFSPCGRFLAACVACMLP 182
Query: 636 HIEADPGLQTPVHQESGIATSPTRHPISAHQVMYELRIYSLEEATFGLVLASRAIRAAHC 695
IEADPGLQ PVHQESG++ SPTRHPISAHQVMYELRIYSLEEATFG VLASRAIRAAHC
Sbjct: 183 RIEADPGLQAPVHQESGLSNSPTRHPISAHQVMYELRIYSLEEATFGSVLASRAIRAAHC 362
Query: 696 LTSIQFSPTSEHILLAYGRRHGSLLKSIVIDGETTLPIYTVLEVYRVSDMELVRVLPSAE 755
LTSIQFSPTSEHILLAYGRRHGSLLKSIVIDGETTLPIYTVLEVYRVSDMELVRVLPSAE
Sbjct: 363 LTSIQFSPTSEHILLAYGRRHGSLLKSIVIDGETTLPIYTVLEVYRVSDMELVRVLPSAE 542
Query: 756 DEVNVACFHPFPGGGLVYGTKEGKLRILHYDGARPVNGTGPSYFPEETIVGV 807
DEVNVACFHPF GGGLVYGTKEGKLRIL YDGA V+GTGPSY+PEETI+GV
Sbjct: 543 DEVNVACFHPFAGGGLVYGTKEGKLRILKYDGAHVVSGTGPSYYPEETIIGV 698
>BP039137
Length = 577
Score = 264 bits (674), Expect = 5e-71
Identities = 129/192 (67%), Positives = 150/192 (77%)
Frame = +3
Query: 478 SRDTLAPQISSSVMANTLPTSNADVAMPSSAMSGSINIPGSSVRSGLRSHFSHSRTPVSE 537
SRD+ QI S M + T+N + AM SS M +I+IP +S+RSGL++HFS S PVS+
Sbjct: 6 SRDSFPQQIGPSTMGSNPSTTNVEAAMSSSEMPATISIPAASMRSGLQNHFSQSHIPVSD 185
Query: 538 SGNLAASINTPHDGSDIQTIMSRIQSELATSVAAAAATELPCTVKLRVWSHDIKNPCSPL 597
SGNL AS+N P DGSD QTI++RIQSE ATS AAAAATELPCTVKL+VW +DI+NPC+ L
Sbjct: 186 SGNLLASVNAPRDGSDSQTIINRIQSEHATSAAAAAATELPCTVKLKVWPYDIENPCALL 365
Query: 598 NADRCRLIIPHAVLCSEMGAHFSPCGRFLAACVACMLPHIEADPGLQTPVHQESGIATSP 657
CRL+IPH VLCSEMGAHFSPCGRFLAACVACM PH E DPG Q Q+ G+ATSP
Sbjct: 366 RG--CRLVIPHVVLCSEMGAHFSPCGRFLAACVACMHPHTEGDPGSQNLARQDPGVATSP 539
Query: 658 TRHPISAHQVMY 669
TRHPISAHQVMY
Sbjct: 540 TRHPISAHQVMY 575
>BP069280
Length = 444
Score = 181 bits (459), Expect = 4e-46
Identities = 93/147 (63%), Positives = 107/147 (72%), Gaps = 2/147 (1%)
Frame = +2
Query: 409 HRNPTDASSLNGMLHGLSRQTANHGVHPEDGH--PFVSSRDPSGWELPFLQGWMMGQSQA 466
H N T+ SLNG+L+GLSRQTAN G E G SSRDPSGWELPF QGW+MGQSQA
Sbjct: 2 HGNSTNTYSLNGVLNGLSRQTANGGFFSELGQLQQLFSSRDPSGWELPFWQGWLMGQSQA 181
Query: 467 GLPSMLPHTGVSRDTLAPQISSSVMANTLPTSNADVAMPSSAMSGSINIPGSSVRSGLRS 526
G+P MLPH G SRD L+ +I SSV A+ +SN D + SSAM GS +IPGSS+RSGLR
Sbjct: 182 GVPPMLPHIGASRDILSQEIGSSVRASNFSSSNVDAVISSSAMPGSTSIPGSSMRSGLRH 361
Query: 527 HFSHSRTPVSESGNLAASINTPHDGSD 553
FS + PVSESGNLAAS+ P DGSD
Sbjct: 362 SFSQAHVPVSESGNLAASVTIPQDGSD 442
>BP064661
Length = 490
Score = 90.1 bits (222), Expect = 1e-18
Identities = 39/46 (84%), Positives = 42/46 (90%)
Frame = -2
Query: 762 CFHPFPGGGLVYGTKEGKLRILHYDGARPVNGTGPSYFPEETIVGV 807
CFHPFPGGGL YGTKEGKLRIL YDGA V+GTGPSY+PEETI+GV
Sbjct: 489 CFHPFPGGGLFYGTKEGKLRILKYDGAHVVSGTGPSYYPEETIIGV 352
>AV407298
Length = 428
Score = 82.0 bits (201), Expect = 3e-16
Identities = 53/107 (49%), Positives = 56/107 (51%), Gaps = 22/107 (20%)
Frame = +3
Query: 27 SNVFNLLARREISPRTKHVA----------------------RDAKRGLLSWYLNYFSL* 64
SNVF+LLARREISPRTKHVA RDA+RGLL+W
Sbjct: 192 SNVFSLLARREISPRTKHVAKWHWGGASKSKSSYNSRSKNVVRDARRGLLAW-------- 347
Query: 65 TISTKM*AS*ITTSLKMILRVEADSLRHLSAKYCPLLPAPRSTIAAA 111
VEADSLRHLSAKYCPLLP PRSTIAAA
Sbjct: 348 --------------------VEADSLRHLSAKYCPLLPPPRSTIAAA 428
>BP038344
Length = 525
Score = 60.8 bits (146), Expect = 8e-10
Identities = 33/94 (35%), Positives = 48/94 (50%), Gaps = 1/94 (1%)
Frame = +2
Query: 111 AFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQE 170
A+S D + S D T++I D G C+K L GH + V F+P + SGS D+
Sbjct: 245 AWSSDSHYICSASDDRTLRIWDATGGDCVKTLRGHTHAVFCVNFNP-QSNYIVSGSFDET 421
Query: 171 VRLWDANTSECI-TSHHFYRPIASIAFHAKGEII 203
VR+W+ T CI T P+ S+ F+ G +I
Sbjct: 422 VRVWEVKTGRCIHTIIAHTMPVTSVHFNRDGTLI 523
>TC10816 homologue to UP|CAE45585 (CAE45585) Coatomer alpha subunit-like
protein, partial (20%)
Length = 979
Score = 48.1 bits (113), Expect = 5e-06
Identities = 31/106 (29%), Positives = 53/106 (49%), Gaps = 4/106 (3%)
Frame = +1
Query: 125 DHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANTSECI-- 182
D+ +K+ + + RCL L+GH+ V+FH P I+ S S DQ +R+W+ + CI
Sbjct: 454 DYKIKVWNYKLHRCLFTLLGHLDYIRTVQFHHESPWIV-SASDDQTIRIWNWQSRTCISV 630
Query: 183 -TSHHFYRPIASIAFHAKGEIIAVAS-GHKLYIWHYDKKGEASYSP 226
T H+ Y + FH + +++ AS + +W + SP
Sbjct: 631 LTGHNHY--VMCALFHPREDLVVSASLDQTVRVWDIGSLKRKNASP 762
Score = 43.9 bits (102), Expect = 1e-04
Identities = 22/64 (34%), Positives = 33/64 (51%)
Frame = +1
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEV 171
F + + S D T++I + ++ C+ VL GH FHP ++ S SLDQ V
Sbjct: 541 FHHESPWIVSASDDQTIRIWNWQSRTCISVLTGHNHYVMCALFHP-REDLVVSASLDQTV 717
Query: 172 RLWD 175
R+WD
Sbjct: 718 RVWD 729
>TC14937 similar to UP|Q9FN19 (Q9FN19) Genomic DNA, chromosome 5, TAC
clone:K8K14 (AT5g67320/K8K14_4), partial (19%)
Length = 1089
Score = 46.6 bits (109), Expect = 2e-05
Identities = 28/87 (32%), Positives = 46/87 (52%)
Frame = +1
Query: 118 VLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDAN 177
VLAS D TVK+ D E G+ + L GH + V F P + + LASGS D+ + +W
Sbjct: 52 VLASASFDSTVKLWDVEVGKLIYSLNGHRDGVYSVAFSP-NGEYLASGSPDKSIHIWSLK 228
Query: 178 TSECITSHHFYRPIASIAFHAKGEIIA 204
+ + ++ I + ++ +G+ IA
Sbjct: 229 EGKIVKTYTGSGGIFEVCWNKEGDKIA 309
Score = 39.3 bits (90), Expect = 0.002
Identities = 23/62 (37%), Positives = 34/62 (54%), Gaps = 2/62 (3%)
Frame = +1
Query: 155 HPLHPKILASGSLDQEVRLWDANTSECITSHHFYRP-IASIAFHAKGEIIAVASGHK-LY 212
+P +LAS S D V+LWD + I S + +R + S+AF GE +A S K ++
Sbjct: 34 NPNKKLVLASASFDSTVKLWDVEVGKLIYSLNGHRDGVYSVAFSPNGEYLASGSPDKSIH 213
Query: 213 IW 214
IW
Sbjct: 214 IW 219
Score = 28.1 bits (61), Expect = 5.8
Identities = 13/36 (36%), Positives = 21/36 (58%)
Frame = +1
Query: 109 AAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIG 144
+ AFSP+G+ LAS D ++ I + G+ +K G
Sbjct: 151 SVAFSPNGEYLASGSPDKSIHIWSLKEGKIVKTYTG 258
>TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG
{Arabidopsis thaliana;} , partial (24%)
Length = 1048
Score = 46.6 bits (109), Expect = 2e-05
Identities = 33/90 (36%), Positives = 48/90 (52%), Gaps = 2/90 (2%)
Frame = +3
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEV 171
FS DGK+LAS D V + + +T + H VRF P + LA+ S+D+ V
Sbjct: 18 FSSDGKMLASAGDDKKVVLWNMDTLQTESTPEQHKSVISDVRFRP-NSSQLATASIDKSV 194
Query: 172 RLWD-ANTSECITSHHFY-RPIASIAFHAK 199
RLWD AN + C+ ++ + I S+ FH K
Sbjct: 195 RLWDAANPTYCVQEYNGHSSAIMSLDFHPK 284
Score = 33.1 bits (74), Expect = 0.18
Identities = 19/71 (26%), Positives = 30/71 (41%), Gaps = 1/71 (1%)
Frame = +3
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGR-CLKVLIGHMRTPWVVRFHPLHPKILASGSLDQE 170
F P+ LA+ D +V++ D C++ GH + FHP I E
Sbjct: 144 FRPNSSQLATASIDKSVRLWDAANPTYCVQEYNGHSSAIMSLDFHPKKTDIFCFCDSANE 323
Query: 171 VRLWDANTSEC 181
+R W+ +S C
Sbjct: 324 IRYWNITSSSC 356
Score = 28.1 bits (61), Expect = 5.8
Identities = 24/104 (23%), Positives = 44/104 (42%)
Frame = +3
Query: 104 PRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILA 163
P + + +G LAS + VKI +G C++ L + FHP + +L
Sbjct: 495 PEPVNSICWDVNGDFLASV-SPNLVKIWSLTSGECIQELSSSGNQFYSCVFHPSYSTLLV 671
Query: 164 SGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVAS 207
G + LW+ ++ +T I+++A ++A AS
Sbjct: 672 IGGF-SSLELWNMADNKSMTISAHESVISALAQSPVTGMVASAS 800
>TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial (60%)
Length = 453
Score = 44.7 bits (104), Expect = 6e-05
Identities = 24/71 (33%), Positives = 34/71 (47%), Gaps = 1/71 (1%)
Frame = +2
Query: 112 FSPDGKVLASTHGDHTVKIIDCET-GRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQE 170
FSP LA++ D TVK+ D + G L+ GH + + FHP ++ S D E
Sbjct: 110 FSPSMPRLATSSFDKTVKVWDVDNPGYSLRTFTGHSASVMSLDFHPNKEDLICSCDSDGE 289
Query: 171 VRLWDANTSEC 181
+R W N C
Sbjct: 290 IRYWSINNGSC 322
>TC8856 similar to UP|Q8W403 (Q8W403) Sec13p, partial (48%)
Length = 568
Score = 44.3 bits (103), Expect = 8e-05
Identities = 29/76 (38%), Positives = 39/76 (51%), Gaps = 4/76 (5%)
Frame = +1
Query: 116 GKVLASTHGDHTVKIIDCE--TGRCLKVLIGHMRTPWVVRF-HPLHPKILASGSLDQEVR 172
GK LA+ DHT+KII + L L GH W V + HP ++AS S D V
Sbjct: 184 GKRLATASSDHTIKIIGVSNTASQHLATLAGHQGPVWQVAWAHPKFGSMIASCSYDGRVI 363
Query: 173 LW-DANTSECITSHHF 187
+W + N +E I +H F
Sbjct: 364 IWKEGNQNEWIQAHVF 411
>TC13023 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Fragment),
partial (18%)
Length = 591
Score = 44.3 bits (103), Expect = 8e-05
Identities = 34/140 (24%), Positives = 60/140 (42%), Gaps = 2/140 (1%)
Frame = +1
Query: 113 SPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVR 172
S DG +A G+ ++KI+D L G + + P + +SG +++R
Sbjct: 187 SSDGSFIACACGE-SIKIVDSANASIRSTLQGDSESVTALALSPDDNLLFSSGH-SRQIR 360
Query: 173 LWDANTSECITSHHFYR-PIASIAFHAKGEIIAV-ASGHKLYIWHYDKKGEASYSPIFVL 230
+WD +T +C+ S + P+ ++ H G ++A + K+ +W D Y F
Sbjct: 361 VWDLSTLKCVRSWKGHEGPVMCMSCHPSGGLLATGGADRKVLVWDVD----GGYCTHFFK 528
Query: 231 KTRRSLRAVHFHPHAAPYLL 250
+ V FHP LL
Sbjct: 529 GHGGVVSCVMFHPDPEKQLL 588
>TC15990
Length = 721
Score = 43.5 bits (101), Expect = 1e-04
Identities = 26/92 (28%), Positives = 42/92 (45%)
Frame = +1
Query: 100 LLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHP 159
L P+P +TI A F+PDGK L + G T+ + ET + H+ PW +++ P
Sbjct: 91 LEPSPGTTIEATFTPDGKYLVAGSGGGTMHAWNIETKNEVACWSSHIGVPWCLKWAPRRA 270
Query: 160 KILASGSLDQEVRLWDANTSECITSHHFYRPI 191
A+ S+ + W N T+ P+
Sbjct: 271 MFAAASSV---LTFWIPNNEYGATATPITSPL 357
>TC16077 similar to PIR|T00593|T02480 sec13-related protein At2g30050 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(46%)
Length = 521
Score = 42.4 bits (98), Expect = 3e-04
Identities = 29/76 (38%), Positives = 38/76 (49%), Gaps = 4/76 (5%)
Frame = +1
Query: 116 GKVLASTHGDHTVKII--DCETGRCLKVLIGHMRTPWVVRF-HPLHPKILASGSLDQEVR 172
GK LA+ DHT+KII + L L GH W V + HP +LAS S D V
Sbjct: 160 GKRLATASSDHTIKIIGVSIAASQHLATLTGHQGPVWQVAWAHPKFGSLLASCSYDGRVI 339
Query: 173 LW-DANTSECITSHHF 187
LW + + +E +H F
Sbjct: 340 LWKEGDQNEWTQAHVF 387
>TC9407 weakly similar to UP|TAF5_YEAST (P38129) Transcription initiation
factor TFIID subunit 5 (TBP-associated factor 5)
(TBP-associated factor 90 kDa) (TAFII-90), partial (4%)
Length = 530
Score = 41.2 bits (95), Expect = 7e-04
Identities = 31/108 (28%), Positives = 46/108 (41%), Gaps = 1/108 (0%)
Frame = +2
Query: 108 IAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSL 167
++ F + LA+ DHTVKI + + K LIGH R W F + L + S
Sbjct: 26 LSPEFCEPHRYLATASADHTVKIWNVDGFTLDKTLIGHQRWVWDCVF-SVDGAYLITASS 202
Query: 168 DQEVRLWDANTSECITSHH-FYRPIASIAFHAKGEIIAVASGHKLYIW 214
D RLW +T E I + ++ A H E + H +I+
Sbjct: 203 DTTARLWSMSTGEDIKVYQGHHKATTCCALHDGAEPASS*PNHHRFIF 346
>TC12572 similar to PIR|T46032|T46032 WD-40 repeat regulatory protein tup1
homolog - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (51%)
Length = 770
Score = 40.4 bits (93), Expect = 0.001
Identities = 25/86 (29%), Positives = 43/86 (49%), Gaps = 3/86 (3%)
Frame = +3
Query: 134 ETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANTSECITS--HHFYRPI 191
++G+CLKVL H V F+ ++ S S D R+WDA+T C+ + P+
Sbjct: 3 KSGKCLKVLPAHSDPVTAVDFN-RDGSLIVSSSYDGLCRIWDASTGHCMKTLIDDENPPV 179
Query: 192 ASIAFHAKGEIIAVAS-GHKLYIWHY 216
+ + F + I V + + L +W+Y
Sbjct: 180 SFVKFSPNAKFILVGTLDNNLRLWNY 257
Score = 40.4 bits (93), Expect = 0.001
Identities = 27/113 (23%), Positives = 52/113 (45%), Gaps = 6/113 (5%)
Frame = +3
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVV--RFHPLHPKILASGSLDQ 169
FSP+ K + D+ +++ + TGR LK GH+ + + + F + K + GS D
Sbjct: 192 FSPNAKFILVGTLDNNLRLWNYSTGRFLKTYTGHVNSKYCISSTFSTTNGKYVVGGSEDH 371
Query: 170 EVRLWDANTSECITSHHFYR-PIASIAFHAKGEII---AVASGHKLYIWHYDK 218
+ LW+ T + + + + S++ H +I A+ + + IW K
Sbjct: 372 GIYLWELQTRKIVQKLEGHSDTVVSVSCHPTENMIASGALGNDKTVKIWTQQK 530
>TC11756 weakly similar to GB|AAH01494.1|16306637|BC001494 U5 snRNP-specific
40 kDa protein (hPrp8-binding) {Homo sapiens;} , partial
(43%)
Length = 719
Score = 38.1 bits (87), Expect = 0.006
Identities = 25/87 (28%), Positives = 42/87 (47%), Gaps = 2/87 (2%)
Frame = +1
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLH--PKILASGSLDQ 169
++ DG + S D TV++ D ETG+ +K ++ H+ +V P P ++ SGS D
Sbjct: 457 WTTDGTQIVSASPDKTVRVWDVETGKQVKKMVEHLS--YVNSCCPSRRGPPLVVSGSDDG 630
Query: 170 EVRLWDANTSECITSHHFYRPIASIAF 196
+LWD I + I ++ F
Sbjct: 631 TAKLWDMRQRGSIQTFPDKYQITAVGF 711
Score = 31.2 bits (69), Expect = 0.68
Identities = 15/43 (34%), Positives = 27/43 (61%)
Frame = +1
Query: 139 LKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANTSEC 181
+ +L GH + ++F+P ++ASGS D+E+ LW+ + EC
Sbjct: 283 IMLLTGHQSVIYTMKFNPAGT-VIASGSHDREIFLWNVH-GEC 405
>TC18213 weakly similar to UP|Q9FIV2 (Q9FIV2) Similarity to pre-mRNA
splicing factor, partial (25%)
Length = 662
Score = 37.7 bits (86), Expect = 0.007
Identities = 22/68 (32%), Positives = 36/68 (52%)
Frame = +1
Query: 108 IAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSL 167
I FS DGK LAS D +V + D + + +K + + V FHP+ P ++AS S
Sbjct: 103 IKCNFSRDGKKLASGSSDGSVYLYDYHSSKVVKKIKAFDQACIDVSFHPVIPNVIASCSW 282
Query: 168 DQEVRLWD 175
D + +++
Sbjct: 283 DGGILVFE 306
>TC10222 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Arabidopsis
thaliana;}, partial (55%)
Length = 557
Score = 36.6 bits (83), Expect = 0.016
Identities = 25/74 (33%), Positives = 35/74 (46%), Gaps = 11/74 (14%)
Frame = +1
Query: 145 HMRTPWVVRFHPL---HPKILASGSLDQEVRLWDA--------NTSECITSHHFYRPIAS 193
H + W V + P P +L +GSLD+ VRLW + NT C+ +AS
Sbjct: 49 HDDSVWAVTWAPATANRPPLLLTGSLDETVRLWRSDELVLERTNTGHCL-------GVAS 207
Query: 194 IAFHAKGEIIAVAS 207
+A H G I A +S
Sbjct: 208 VAAHPLGSIAASSS 249
>TC18293 similar to UP|PWP1_HUMAN (Q13610) Periodic tryptophan protein 1
homolog (Keratinocyte protein IEF SSP 9502), partial
(7%)
Length = 804
Score = 35.4 bits (80), Expect = 0.036
Identities = 36/133 (27%), Positives = 54/133 (40%), Gaps = 5/133 (3%)
Frame = +2
Query: 118 VLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDAN 177
+LAS D VKI D +C + H V ++ P++L +GS D V L D
Sbjct: 413 ILASASADKRVKIWDVVAEKCDITMEHHTDKVQAVAWNHFQPQVLLTGSFDHTVALKDGR 592
Query: 178 TSECITSHHFYR-----PIASIAFHAKGEIIAVASGHKLYIWHYDKKGEASYSPIFVLKT 232
+ SH YR + S+A+ E V S + +D + A +P L +
Sbjct: 593 ----MPSHSGYRWSVNSDVESLAWDPHTEHSFVVSLDDGTVKCFDIR-TAQSNPTSELSS 757
Query: 233 RRSLRAVHFHPHA 245
S +H H A
Sbjct: 758 AASTFTLHAHDKA 796
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.134 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,198,547
Number of Sequences: 28460
Number of extensions: 230001
Number of successful extensions: 1356
Number of sequences better than 10.0: 85
Number of HSP's better than 10.0 without gapping: 1299
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1336
length of query: 808
length of database: 4,897,600
effective HSP length: 98
effective length of query: 710
effective length of database: 2,108,520
effective search space: 1497049200
effective search space used: 1497049200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC133341.5


