

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133139.3 - phase: 0
(185 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
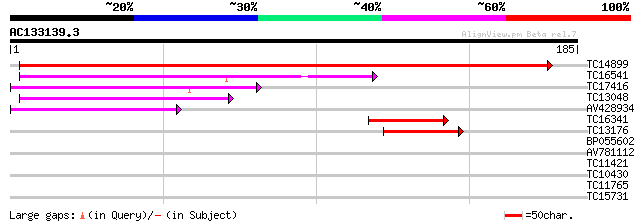
Score E
Sequences producing significant alignments: (bits) Value
TC14899 similar to UP|UMPK_ARATH (O04905) Uridylate kinase (UK)... 209 2e-55
TC16541 similar to UP|Q9SNJ4 (Q9SNJ4) ESTs AU065232(E60855), par... 65 7e-12
TC17416 similar to UP|KADD_ARATH (Q9FIJ7) Probable adenylate kin... 52 8e-08
TC13048 similar to UP|KADC_ARATH (Q9ZUU1) Probable adenylate kin... 47 2e-06
AV428934 45 8e-06
TC16341 42 6e-05
TC13176 39 5e-04
BP055602 38 0.001
AV781112 34 0.013
TC11421 33 0.030
TC10430 homologue to UP|KADB_ORYSA (Q08480) Adenylate kinase B ... 33 0.039
TC11765 similar to UP|Q8L7W7 (Q8L7W7) AT3g01820/F28J7_15, partia... 31 0.11
TC15731 similar to UP|Q07526 (Q07526) Protein kinase PSPK-5 (Pr... 25 8.1
>TC14899 similar to UP|UMPK_ARATH (O04905) Uridylate kinase (UK) (Uridine
monophosphate kinase) (UMP kinase) (UMP/CMP kinase) ,
partial (92%)
Length = 911
Score = 209 bits (532), Expect = 2e-55
Identities = 102/174 (58%), Positives = 130/174 (74%)
Frame = +2
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQC+ IV+ FG+ HL AGDLLR + S SE G MI I+EG+IVPS VT+
Sbjct: 113 GGPGSGKGTQCSNIVKHFGYTHLIAGDLLRSEIKSGSENGTMIQNLIKEGKIVPSDVTIN 292
Query: 64 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQG 123
L+ R M N KFLIDGFPR+EENR AFE +TG EP FVL+FDCPEEEM +R+L RNQG
Sbjct: 293 LLQRAMLENGNDKFLIDGFPRNEENRAAFEKVTGIEPTFVLFFDCPEEEMERRLLGRNQG 472
Query: 124 RIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVFA 177
R DDNI+TI+KR VF +LPVI++Y + ++ +I+A +E+FE V+ +F+
Sbjct: 473 REDDNIETIRKRFNVFLESSLPVINYYDAKLKVRKIDAARPVEEVFESVKAIFS 634
>TC16541 similar to UP|Q9SNJ4 (Q9SNJ4) ESTs AU065232(E60855), partial (41%)
Length = 525
Score = 65.1 bits (157), Expect = 7e-12
Identities = 33/120 (27%), Positives = 62/120 (51%), Gaps = 3/120 (2%)
Frame = +2
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
G PG GKGT +R+ G H++ GDL+R + S+ + + E + +G++V + +
Sbjct: 161 GSPGVGKGTYASRLCTLLGVPHIATGDLVRNELSSNGPLSSQLSEIVNQGKLVSDEIIIN 340
Query: 64 LILREM---QYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSR 120
L+ + + + F++DGFPR+ + E + T+ D V+ EE ++ + L R
Sbjct: 341 LLSKRLAAREANGESGFILDGFPRTMKQAEILEGV--TDIDLVVNLKLQEEVLLAKCLGR 514
>TC17416 similar to UP|KADD_ARATH (Q9FIJ7) Probable adenylate kinase 2,
chloroplast precursor (ATP-AMP transphosphorylase) ,
partial (36%)
Length = 537
Score = 51.6 bits (122), Expect = 8e-08
Identities = 30/92 (32%), Positives = 47/92 (50%), Gaps = 10/92 (10%)
Frame = +2
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVP--- 57
M+SG P SGKGTQC I +G H++AG+LLR + + S+ G + EG + P
Sbjct: 254 MISGAPASGKGTQCELITNKYGLVHVAAGNLLRAEIATGSQNGQRAKDYHGEGTVGP**K 433
Query: 58 -------SAVTVRLILREMQYGDNRKFLIDGF 82
+++ RL + + +G K LI +
Sbjct: 434 CCHDGEGTSLEARLYTQGLAFGWLSKELITSY 529
>TC13048 similar to UP|KADC_ARATH (Q9ZUU1) Probable adenylate kinase 1,
chloroplast precursor (ATP-AMP transphosphorylase) ,
partial (29%)
Length = 469
Score = 47.0 bits (110), Expect = 2e-06
Identities = 19/70 (27%), Positives = 38/70 (54%)
Frame = +3
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
G PG GKGT +R+ H++ GDL+R+ + S + E +++G++V + +
Sbjct: 240 GCPGVGKGTYASRLSNLLHVPHIATGDLVREELTSSGPLSTELSEIVKQGQLVSDEIIIS 419
Query: 64 LILREMQYGD 73
L+ + + G+
Sbjct: 420 LLSKRLAAGE 449
>AV428934
Length = 323
Score = 45.1 bits (105), Expect = 8e-06
Identities = 22/56 (39%), Positives = 33/56 (58%)
Frame = +1
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIV 56
+L G PGSGKGTQ I + + HL+ GD+LR A+ + + G E + +G +V
Sbjct: 145 ILIGPPGSGKGTQSPIIKDEYCLCHLATGDMLRAAVAAKTPLGIKAKEAMEKGELV 312
>TC16341
Length = 531
Score = 42.0 bits (97), Expect = 6e-05
Identities = 19/26 (73%), Positives = 23/26 (88%)
Frame = +3
Query: 118 LSRNQGRIDDNIDTIKKRLKVFEALN 143
LS +QG IDDNIDT+KK LK+FE+LN
Sbjct: 243 LSSSQGSIDDNIDTMKKCLKIFESLN 320
>TC13176
Length = 527
Score = 38.9 bits (89), Expect = 5e-04
Identities = 17/26 (65%), Positives = 22/26 (84%)
Frame = +1
Query: 123 GRIDDNIDTIKKRLKVFEALNLPVID 148
G ID+NI T+KKR+K+FE LNL V+D
Sbjct: 61 GSIDNNIVTMKKRIKIFETLNLHVLD 138
>BP055602
Length = 446
Score = 37.7 bits (86), Expect = 0.001
Identities = 17/26 (65%), Positives = 22/26 (84%)
Frame = -2
Query: 118 LSRNQGRIDDNIDTIKKRLKVFEALN 143
LS + G IDD+IDT+KK LK+FE+LN
Sbjct: 289 LSSSPGSIDDHIDTMKKCLKIFESLN 212
>AV781112
Length = 488
Score = 34.3 bits (77), Expect = 0.013
Identities = 14/21 (66%), Positives = 19/21 (89%)
Frame = -2
Query: 123 GRIDDNIDTIKKRLKVFEALN 143
G++ DNIDT+KK LK+FE+LN
Sbjct: 229 GQLTDNIDTMKKCLKIFESLN 167
>TC11421
Length = 605
Score = 33.1 bits (74), Expect = 0.030
Identities = 16/25 (64%), Positives = 19/25 (76%)
Frame = +2
Query: 123 GRIDDNIDTIKKRLKVFEALNLPVI 147
G ID NIDT+KK L +FE+ NL VI
Sbjct: 332 GSIDGNIDTMKKWLTIFESSNLYVI 406
>TC10430 homologue to UP|KADB_ORYSA (Q08480) Adenylate kinase B (ATP-AMP
transphosphorylase) , partial (27%)
Length = 392
Score = 32.7 bits (73), Expect = 0.039
Identities = 16/54 (29%), Positives = 28/54 (51%)
Frame = +3
Query: 124 RIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVFA 177
R DD +K RL+ F PVID+Y+++ + ++A E+ +V V +
Sbjct: 36 RKDDTAAVLKSRLEAFHKQTEPVIDYYSKQSLVANLHAEKPPKEVTVEVEKVLS 197
>TC11765 similar to UP|Q8L7W7 (Q8L7W7) AT3g01820/F28J7_15, partial (38%)
Length = 468
Score = 31.2 bits (69), Expect = 0.11
Identities = 15/72 (20%), Positives = 34/72 (46%)
Frame = +1
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
++ G PG+ + ++ + H+S LLR+ + S I + +G++VP +
Sbjct: 223 VMIGEPGTKRHLFAEKLSKLLEVPHISMASLLRQELNPRSSLYQQIANALDQGKLVPEEI 402
Query: 61 TVRLILREMQYG 72
L+ + ++ G
Sbjct: 403 IFALLSKRLEDG 438
>TC15731 similar to UP|Q07526 (Q07526) Protein kinase PSPK-5 (Probable
serine/threonine-protein kinase PK5) , partial (17%)
Length = 940
Score = 25.0 bits (53), Expect = 8.1
Identities = 16/47 (34%), Positives = 23/47 (48%), Gaps = 2/47 (4%)
Frame = -2
Query: 82 FPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVK--RVLSRNQGRID 126
F SE+ I F I F+ +F CP+E +K R + N G +D
Sbjct: 273 FNLSEDTHILFHRIYIF--GFLFFFSCPKERSIKLRRFRATN*GPVD 139
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.142 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,408,844
Number of Sequences: 28460
Number of extensions: 24147
Number of successful extensions: 93
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 92
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 92
length of query: 185
length of database: 4,897,600
effective HSP length: 85
effective length of query: 100
effective length of database: 2,478,500
effective search space: 247850000
effective search space used: 247850000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC133139.3


