

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130807.10 - phase: 0 /pseudo
(530 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
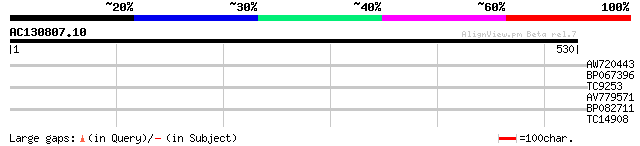
Score E
Sequences producing significant alignments: (bits) Value
AW720443 31 0.43
BP067396 28 4.8
TC9253 weakly similar to UP|Q9XI93 (Q9XI93) F7A19.2 protein (F16... 27 6.3
AV779571 27 8.2
BP082711 27 8.2
TC14908 weakly similar to PIR|A86218|A86218 protein T27G7.18 [im... 27 8.2
>AW720443
Length = 406
Score = 31.2 bits (69), Expect = 0.43
Identities = 19/33 (57%), Positives = 23/33 (69%)
Frame = +3
Query: 322 LVSLIVLL*LLL*ILVLLIVSLLLNVHISWVWF 354
L S ++LL LL*+LVLLIVS LL+ I W F
Sbjct: 204 LDSRMMLLSTLL*LLVLLIVSPLLSQSIQWTGF 302
>BP067396
Length = 506
Score = 27.7 bits (60), Expect = 4.8
Identities = 19/61 (31%), Positives = 29/61 (47%), Gaps = 11/61 (18%)
Frame = -2
Query: 241 SVAIKATRVMFVPKMRR-----------SASGVVRRVTH*LIVRGETLFAITTMKRAISV 289
++ ++ATR F PK+R+ SA G+ R++TH* F I M + V
Sbjct: 463 TIVLRATRNCFTPKLRKEVILVFLLAAQSA*GITRKLTH*------GCFLIVAMSTTLHV 302
Query: 290 L 290
L
Sbjct: 301 L 299
>TC9253 weakly similar to UP|Q9XI93 (Q9XI93) F7A19.2 protein (F16A14.14)
(AT1G13930/F16A14.27), partial (46%)
Length = 745
Score = 27.3 bits (59), Expect = 6.3
Identities = 18/46 (39%), Positives = 29/46 (62%), Gaps = 1/46 (2%)
Frame = +2
Query: 313 LMRTDLSEVLVSLIV-LL*LLL*ILVLLIVSLLLNVHISWVWFYLI 357
L+R ++ ++SL+V LL LLL L VS+LLN+ + VW ++
Sbjct: 278 LIRLRITSTIISLVVLLLLLLLPNQKNLKVSMLLNLLVEVVWVEIL 415
>AV779571
Length = 538
Score = 26.9 bits (58), Expect = 8.2
Identities = 20/58 (34%), Positives = 37/58 (63%)
Frame = -2
Query: 321 VLVSLIVLL*LLL*ILVLLIVSLLLNVHISWVWFYLI*KEKWLLKLQLRV**LLLLFV 378
+++ ++L LLL +L+LLIV ++L++ I V + LLK++LR+ LL+L +
Sbjct: 321 LIIYHLLLSLLLLLLLLLLIVIMILSLTIE*VLML*V-----LLKMRLRLQQLLILLI 163
>BP082711
Length = 379
Score = 26.9 bits (58), Expect = 8.2
Identities = 16/37 (43%), Positives = 24/37 (64%)
Frame = -1
Query: 309 VRRLLMRTDLSEVLVSLIVLL*LLL*ILVLLIVSLLL 345
V LMR+ E + ++LL LLL +L+LL++ LLL
Sbjct: 286 VEMWLMRSSFGES*LMFLLLLLLLLLLLLLLLLLLLL 176
>TC14908 weakly similar to PIR|A86218|A86218 protein T27G7.18 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(68%)
Length = 1219
Score = 26.9 bits (58), Expect = 8.2
Identities = 15/37 (40%), Positives = 22/37 (58%)
Frame = +2
Query: 93 CSGENF*TGTSRKMSEVRKRLNFWS*SRVTCLLLSML 129
CS F T TSRKM + R+ + S+V+C LL ++
Sbjct: 59 CSASTF-THTSRKMEQGRRSSSCLLCSKVSCFLLLLI 166
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.370 0.167 0.608
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,002,731
Number of Sequences: 28460
Number of extensions: 107762
Number of successful extensions: 1633
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 874
Number of HSP's successfully gapped in prelim test: 35
Number of HSP's that attempted gapping in prelim test: 660
Number of HSP's gapped (non-prelim): 956
length of query: 530
length of database: 4,897,600
effective HSP length: 95
effective length of query: 435
effective length of database: 2,193,900
effective search space: 954346500
effective search space used: 954346500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC130807.10


