

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130806.3 - phase: 0
(320 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
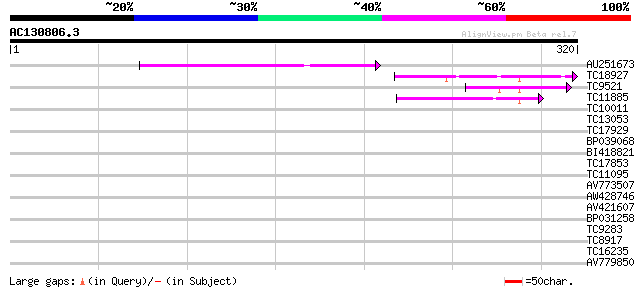
Score E
Sequences producing significant alignments: (bits) Value
AU251673 52 2e-07
TC18927 similar to PIR|AI2934|AI2934 chromate transport protein ... 45 2e-05
TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, part... 41 3e-04
TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 39 9e-04
TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, par... 36 0.008
TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 36 0.010
TC17929 32 0.11
BP039068 32 0.14
BI418821 31 0.24
TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, ... 30 0.54
TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 29 0.93
AV773507 28 1.6
AW428746 28 2.7
AV421607 27 3.5
BP031258 27 3.5
TC9283 27 4.6
TC8917 similar to UP|AAH64262 (AAH64262) MGC76273 protein, parti... 27 4.6
TC16235 27 6.0
AV779850 26 7.9
>AU251673
Length = 413
Score = 51.6 bits (122), Expect = 2e-07
Identities = 34/137 (24%), Positives = 65/137 (46%), Gaps = 1/137 (0%)
Frame = +2
Query: 74 TEDQKVRFGTHQLAEEGDDWWVSLLPTLKQDGAAVTWAVFRREFLSRYFPEDVRGKKEIE 133
++ + V + QL DW+ L TWA F EF++R+ P+ VR +
Sbjct: 8 SDTRAVELASFQLEGVARDWYNVLTRAKPVGSPPWTWADFSAEFMNRFLPQSVRDGFVRD 187
Query: 134 FLELKQG-NMSMTEYAAKFVELAKFYPHYAAETTEFLKCIKFENGLRADIKRAIGYQQIR 192
F L+Q M+++EY+A F L+++ P+ E + +F GL+ + +++ +
Sbjct: 188 FERLEQAEGMTVSEYSAHFTHLSRYVPYPLLEEE---RVKRFVRGLKEYLFKSVVGSKSS 358
Query: 193 VFSELVNSCRIYEDDTK 209
SE+++ + E K
Sbjct: 359 TLSEVLSLALLVEQRQK 409
>TC18927 similar to PIR|AI2934|AI2934 chromate transport protein chrA
[imported] - Agrobacterium tumefaciens
(strain C58, Dupont) {Agrobacterium tumefaciens;},
partial (6%)
Length = 561
Score = 45.1 bits (105), Expect = 2e-05
Identities = 32/117 (27%), Positives = 51/117 (43%), Gaps = 14/117 (11%)
Frame = -2
Query: 218 RKTKGQQSRPKPYSAPADKGKQRMVEDR------RPKKKDALAEIICFNCGEKGHKSNAC 271
R + + + KP+ P ++G RP + D +EI+C C +KGH +N C
Sbjct: 464 RFDRNKSFQKKPFQRPQNRGTSSGYSHSFGNFVPRPTQSDT-SEIVCHRCSKKGHFANRC 288
Query: 272 PSEIKRCFRCGKKGH--------KSNAEIICFNIQHEGRASHRGRSPANRSSAEVPT 320
P + C+ C K GH K A + A+++G+ P +SA V T
Sbjct: 287 PDLV--CWNCQKTGHSGKDCTNPKVEAATNAIAARRPAPAANKGKRPV--ASARVYT 129
>TC9521 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (56%)
Length = 598
Score = 40.8 bits (94), Expect = 3e-04
Identities = 21/67 (31%), Positives = 36/67 (53%), Gaps = 7/67 (10%)
Frame = +1
Query: 258 CFNCGEKGHKSNACPSEI--KRCFRCGKKGH-----KSNAEIICFNIQHEGRASHRGRSP 310
CFNCG GH + C + +C+RCG++GH K++ + + + + + R+ R RSP
Sbjct: 391 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSPKKLSRHPRSDSRSPVRSRSP 570
Query: 311 ANRSSAE 317
S +
Sbjct: 571 RRGRSRD 591
>TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (26%)
Length = 555
Score = 39.3 bits (90), Expect = 9e-04
Identities = 20/86 (23%), Positives = 38/86 (43%), Gaps = 3/86 (3%)
Frame = +2
Query: 219 KTKGQQSRPKPYSAPADKGKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNACPSEIKRC 278
+++ + P +D+ R RR ++ + +C NC GH + CP+ + C
Sbjct: 227 RSRSRSRSPMDRKIRSDRFSYRDAPYRRDSRRGFSRDNLCKNCKRPGHFARECPN-VAIC 403
Query: 279 FRCGKKGH---KSNAEIICFNIQHEG 301
CG GH + + +C+N + G
Sbjct: 404 HNCGLPGHIASECTTKSLCWNCKEPG 481
Score = 29.3 bits (64), Expect = 0.93
Identities = 8/18 (44%), Positives = 15/18 (82%)
Frame = +2
Query: 257 ICFNCGEKGHKSNACPSE 274
+C+NC E GH +++CP++
Sbjct: 455 LCWNCKEPGHMASSCPND 508
>TC10011 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (62%)
Length = 684
Score = 36.2 bits (82), Expect = 0.008
Identities = 14/31 (45%), Positives = 19/31 (61%), Gaps = 2/31 (6%)
Frame = +2
Query: 258 CFNCGEKGHKSNACPSE--IKRCFRCGKKGH 286
CFNCG GH + C + +C+RCG +GH
Sbjct: 371 CFNCGLDGHWARDCKAGDWKNKCYRCGDRGH 463
>TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (9%)
Length = 450
Score = 35.8 bits (81), Expect = 0.010
Identities = 12/31 (38%), Positives = 17/31 (54%)
Frame = +3
Query: 256 IICFNCGEKGHKSNACPSEIKRCFRCGKKGH 286
++C NC + GH S C + C CG +GH
Sbjct: 3 VVCRNCQQLGHMSRDCMGPLMICHNCGGRGH 95
Score = 29.3 bits (64), Expect = 0.93
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = +3
Query: 256 IICFNCGEKGHKSNACPS 273
+IC NCG +GH + CPS
Sbjct: 63 MICHNCGGRGHLAYECPS 116
>TC17929
Length = 791
Score = 32.3 bits (72), Expect = 0.11
Identities = 12/32 (37%), Positives = 16/32 (49%)
Frame = +2
Query: 243 EDRRPKKKDALAEIICFNCGEKGHKSNACPSE 274
E+ + D+ C+ CGE GHK CP E
Sbjct: 17 EETSNRPNDSKFRQTCYRCGESGHKMRNCPKE 112
>BP039068
Length = 467
Score = 32.0 bits (71), Expect = 0.14
Identities = 11/19 (57%), Positives = 13/19 (67%)
Frame = +3
Query: 255 EIICFNCGEKGHKSNACPS 273
+ +C NC E GH SN CPS
Sbjct: 303 QTVCMNCQETGHASNDCPS 359
>BI418821
Length = 614
Score = 31.2 bits (69), Expect = 0.24
Identities = 12/36 (33%), Positives = 16/36 (44%), Gaps = 7/36 (19%)
Frame = +2
Query: 258 CFNCGEKGHKSNACPSEIKR-------CFRCGKKGH 286
C+NCG+ GH + C C+ CG GH
Sbjct: 476 CYNCGDAGHLARDCNRSNNNSGGGGAGCYNCGDTGH 583
Score = 30.8 bits (68), Expect = 0.32
Identities = 12/37 (32%), Positives = 16/37 (42%), Gaps = 8/37 (21%)
Frame = +2
Query: 258 CFNCGEKGHKSNACPSEIKR--------CFRCGKKGH 286
C+NCG+ GH + C C+ CG GH
Sbjct: 392 CYNCGDTGHLARDCHRSNNNGGGGGGAACYNCGDAGH 502
>TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, partial
(58%)
Length = 881
Score = 30.0 bits (66), Expect = 0.54
Identities = 10/17 (58%), Positives = 12/17 (69%)
Frame = +2
Query: 258 CFNCGEKGHKSNACPSE 274
CF CG+ GH + CPSE
Sbjct: 419 CFKCGKPGHFARECPSE 469
>TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (89%)
Length = 912
Score = 29.3 bits (64), Expect = 0.93
Identities = 20/63 (31%), Positives = 28/63 (43%), Gaps = 2/63 (3%)
Frame = +1
Query: 254 AEIICFNCGEKGHKSNACPSEIKRCFRCGKKGHKSNAEIICFNIQHEGRASH--RGRSPA 311
+++ C+ CGE GH + C R G +S + GR S+ RGRSP
Sbjct: 367 SDMKCYECGEPGHFARECR------MRGGSGRRRSRSPPRFRRSPSYGRRSYSPRGRSPR 528
Query: 312 NRS 314
RS
Sbjct: 529 RRS 537
>AV773507
Length = 496
Score = 28.5 bits (62), Expect = 1.6
Identities = 9/22 (40%), Positives = 13/22 (58%)
Frame = +2
Query: 251 DALAEIICFNCGEKGHKSNACP 272
D + + CF CG GH++ CP
Sbjct: 329 DRVGQDDCFECGRPGHRARDCP 394
>AW428746
Length = 398
Score = 27.7 bits (60), Expect = 2.7
Identities = 20/71 (28%), Positives = 30/71 (42%), Gaps = 1/71 (1%)
Frame = -3
Query: 113 FRREFLSRYFPEDVRGKK-EIEFLELKQGNMSMTEYAAKFVELAKFYPHYAAETTEFLKC 171
F R + +Y P++ + EFL L N+ EY V FY HY +C
Sbjct: 231 FFRILVLKYLPKNFKSSSPNFEFLFL---NVIFHEYHLTHVSKGNFYIHYI*LPQGIHQC 61
Query: 172 IKFENGLRADI 182
I + +R +I
Sbjct: 60 ILMKTNMRVNI 28
>AV421607
Length = 245
Score = 27.3 bits (59), Expect = 3.5
Identities = 9/16 (56%), Positives = 10/16 (62%)
Frame = +3
Query: 258 CFNCGEKGHKSNACPS 273
C+ CG GH S CPS
Sbjct: 54 CYKCGRPGHWSRDCPS 101
>BP031258
Length = 526
Score = 27.3 bits (59), Expect = 3.5
Identities = 14/28 (50%), Positives = 17/28 (60%)
Frame = -2
Query: 278 CFRCGKKGHKSNAEIICFNIQHEGRASH 305
C R KGH+ N+ I N+Q EGR SH
Sbjct: 507 CTRHPSKGHRHNSSPILCNLQ-EGRFSH 427
>TC9283
Length = 1307
Score = 26.9 bits (58), Expect = 4.6
Identities = 11/42 (26%), Positives = 24/42 (56%)
Frame = +2
Query: 64 IERVFRVMQYTEDQKVRFGTHQLAEEGDDWWVSLLPTLKQDG 105
+ER+ R M E ++ +G H L+ ++ +++ LK++G
Sbjct: 905 VERLCRAMGGAEKVEIEYGNHSLSNRVEEAVQAIIDFLKREG 1030
>TC8917 similar to UP|AAH64262 (AAH64262) MGC76273 protein, partial (3%)
Length = 676
Score = 26.9 bits (58), Expect = 4.6
Identities = 11/46 (23%), Positives = 23/46 (49%)
Frame = +2
Query: 207 DTKAHYKIVNERKTKGQQSRPKPYSAPADKGKQRMVEDRRPKKKDA 252
+ K E K + ++ PKP P ++ + V++ +PK ++A
Sbjct: 341 EPKEEKATTEEVKVETKEVNPKPEEEPKEEEPKAQVQEEKPKTEEA 478
>TC16235
Length = 411
Score = 26.6 bits (57), Expect = 6.0
Identities = 10/16 (62%), Positives = 11/16 (68%)
Frame = +1
Query: 266 HKSNACPSEIKRCFRC 281
H+SN CP IK C RC
Sbjct: 169 HRSNRCPRLIKICRRC 216
>AV779850
Length = 525
Score = 26.2 bits (56), Expect = 7.9
Identities = 14/29 (48%), Positives = 14/29 (48%)
Frame = -3
Query: 225 SRPKPYSAPADKGKQRMVEDRRPKKKDAL 253
S PKP SAP D K R PK AL
Sbjct: 388 SSPKPLSAPGDLKKSRRRSLNGPKSATAL 302
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.133 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,406,102
Number of Sequences: 28460
Number of extensions: 71548
Number of successful extensions: 373
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 343
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 366
length of query: 320
length of database: 4,897,600
effective HSP length: 90
effective length of query: 230
effective length of database: 2,336,200
effective search space: 537326000
effective search space used: 537326000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC130806.3


