

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130804.6 + phase: 0 /pseudo
(99 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
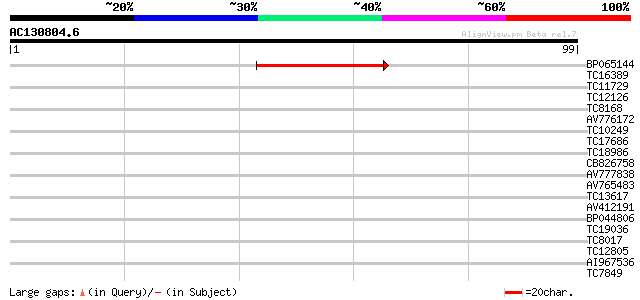
Sequences producing significant alignments: (bits) Value
BP065144 40 5e-05
TC16389 weakly similar to GB|AAO11631.1|27363424|BT002715 At2g39... 27 0.74
TC11729 27 0.74
TC12126 similar to PIR|S49166|S49166 cysteine proteinase precur... 26 0.97
TC8168 similar to UP|O04720 (O04720) Cysteine proteinase inhibit... 26 1.3
AV776172 25 1.7
TC10249 similar to UP|Q84LL9 (Q84LL9) Salt tolerance protein 3, ... 25 2.2
TC17686 similar to UP|CYSP_VIGMU (P12412) Vignain precursor (Be... 25 2.8
TC18986 24 3.7
CB826758 24 3.7
AV777838 24 4.8
AV765483 24 4.8
TC13617 similar to UP|Q9FKY1 (Q9FKY1) Similarity to ribonucleopr... 23 6.3
AV412191 23 6.3
BP044806 23 6.3
TC19036 23 6.3
TC8017 weakly similar to UP|Q7UAE4 (Q7UAE4) Invasion plasmid ant... 23 6.3
TC12805 23 6.3
AI967536 23 6.3
TC7849 similar to GB|AAN18126.1|23308313|BT000557 At2g10940/F15K... 23 8.2
>BP065144
Length = 498
Score = 40.4 bits (93), Expect = 5e-05
Identities = 18/23 (78%), Positives = 21/23 (91%)
Frame = +2
Query: 44 IDGVTPRAKMNKQRTRRFRTAKD 66
ID V PRAKMN+QR+RRFR+AKD
Sbjct: 347 IDWVAPRAKMNQQRSRRFRSAKD 415
>TC16389 weakly similar to GB|AAO11631.1|27363424|BT002715
At2g39190/T16B24.17 {Arabidopsis thaliana;}, partial
(22%)
Length = 704
Score = 26.6 bits (57), Expect = 0.74
Identities = 11/29 (37%), Positives = 18/29 (61%)
Frame = -3
Query: 34 LIAPPPTLPDIDGVTPRAKMNKQRTRRFR 62
+IAPPPTL + T + NK++ ++ R
Sbjct: 549 VIAPPPTLFSVRAATASSSRNKKKQKKPR 463
>TC11729
Length = 503
Score = 26.6 bits (57), Expect = 0.74
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = -1
Query: 23 HETLHAKFNRWLIAPPPTLPDIDGVT 48
H T H+KFN ++ PPP L ++ +T
Sbjct: 275 HITCHSKFNGFI--PPPPLGNVSSIT 204
>TC12126 similar to PIR|S49166|S49166 cysteine proteinase precursor -
spring vetch {Vicia sativa;} , partial (71%)
Length = 839
Score = 26.2 bits (56), Expect = 0.97
Identities = 11/33 (33%), Positives = 19/33 (57%)
Frame = +1
Query: 56 QRTRRFRTAKDNEMREAEEERLRKEFEMEANKF 88
++ +RF K N M E +L K ++++ NKF
Sbjct: 178 EKHKRFNVFKANVMHVHETNKLDKPYKLKLNKF 276
>TC8168 similar to UP|O04720 (O04720) Cysteine proteinase inhibitor,
partial (55%)
Length = 808
Score = 25.8 bits (55), Expect = 1.3
Identities = 19/70 (27%), Positives = 31/70 (44%), Gaps = 1/70 (1%)
Frame = -3
Query: 11 CHLLANTLKQKIHETLHAKF-NRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEM 69
C LL ++ ++ ++ +F N W PP+L ID + + R R R K
Sbjct: 407 CLLLCSSTAKRARASISREF*NPWESRMPPSLVAIDT*SYSEQTADDRGRVKRAQKRKRK 228
Query: 70 REAEEERLRK 79
+ EEE L +
Sbjct: 227 KGMEEEDLMR 198
>AV776172
Length = 433
Score = 25.4 bits (54), Expect = 1.7
Identities = 17/55 (30%), Positives = 23/55 (40%)
Frame = +2
Query: 25 TLHAKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRK 79
TLHA WL PP LP G P+ + +Q R R E+ + R+
Sbjct: 254 TLHAPAGVWLRREPPLLP---GSLPQPREARQ-GRELRRVPGQELHQLRNGSTRR 406
>TC10249 similar to UP|Q84LL9 (Q84LL9) Salt tolerance protein 3, partial
(35%)
Length = 1096
Score = 25.0 bits (53), Expect = 2.2
Identities = 15/35 (42%), Positives = 19/35 (53%), Gaps = 8/35 (22%)
Frame = +1
Query: 61 FRTAKDNEM--------REAEEERLRKEFEMEANK 87
F+TAK+ + REAEE R RKE E E +
Sbjct: 127 FQTAKEKAIEEERLAREREAEERRKRKEQEYERKR 231
>TC17686 similar to UP|CYSP_VIGMU (P12412) Vignain precursor (Bean
endopeptidase) (Cysteine proteinase)
(Sulfhydryl-endopeptidase) (SH-EP) , partial (54%)
Length = 624
Score = 24.6 bits (52), Expect = 2.8
Identities = 10/33 (30%), Positives = 18/33 (54%)
Frame = +3
Query: 56 QRTRRFRTAKDNEMREAEEERLRKEFEMEANKF 88
++ +RF K N M + +L +++E NKF
Sbjct: 162 EKRKRFNVFKANVMHVHDTNKLDMPYKLELNKF 260
>TC18986
Length = 678
Score = 24.3 bits (51), Expect = 3.7
Identities = 10/21 (47%), Positives = 12/21 (56%)
Frame = +2
Query: 7 NKFACHLLANTLKQKIHETLH 27
NKF CHL +T KQ+ H
Sbjct: 383 NKFECHLRQSTWKQQALSPTH 445
>CB826758
Length = 537
Score = 24.3 bits (51), Expect = 3.7
Identities = 20/50 (40%), Positives = 24/50 (48%), Gaps = 2/50 (4%)
Frame = +2
Query: 50 RAKMNKQRTRRFRTAKDNEMREAEEERLRK--EFEMEANKFFLNKNVKCQ 97
R + K R AK E AEE LRK E E+E K L +N+K Q
Sbjct: 86 RGRAEKDAIEAIRRAKAAESLYAEELNLRKMAEEELETEKEEL-ENLKSQ 232
>AV777838
Length = 565
Score = 23.9 bits (50), Expect = 4.8
Identities = 13/41 (31%), Positives = 18/41 (43%), Gaps = 13/41 (31%)
Frame = +1
Query: 35 IAPPPTLPDIDGV-------------TPRAKMNKQRTRRFR 62
+ PPP+ PD+ +PR + N QR RFR
Sbjct: 442 LGPPPSKPDLRRTAHDLDKPTPPLCSSPRPQQNPQRNFRFR 564
>AV765483
Length = 483
Score = 23.9 bits (50), Expect = 4.8
Identities = 9/19 (47%), Positives = 13/19 (68%)
Frame = +2
Query: 16 NTLKQKIHETLHAKFNRWL 34
NT+ KI E + +KFN +L
Sbjct: 59 NTIDNKIDERIRSKFNNYL 115
>TC13617 similar to UP|Q9FKY1 (Q9FKY1) Similarity to ribonucleoprotein F,
partial (35%)
Length = 474
Score = 23.5 bits (49), Expect = 6.3
Identities = 22/80 (27%), Positives = 34/80 (42%), Gaps = 4/80 (5%)
Frame = -1
Query: 10 ACHLLANTLKQKIHETLHAKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRT----AK 65
A H+L L+ K++ K N+ L PPTL D + N Q ++ F+ A
Sbjct: 231 ATHVLPIPLQSKLNLHCSWKDNKGLS*EPPTLVDQQHI-----HNGQSSKEFQNVNICAV 67
Query: 66 DNEMREAEEERLRKEFEMEA 85
+ E E + R+ E A
Sbjct: 66 EREPPETNHWKGRRRLEARA 7
>AV412191
Length = 416
Score = 23.5 bits (49), Expect = 6.3
Identities = 12/38 (31%), Positives = 17/38 (44%)
Frame = +1
Query: 15 ANTLKQKIHETLHAKFNRWLIAPPPTLPDIDGVTPRAK 52
ANT + T + +RW PPP+ + G P K
Sbjct: 298 ANTPSSHVKATNVSHHHRWSQQPPPSSREHHGEAP*TK 411
>BP044806
Length = 538
Score = 23.5 bits (49), Expect = 6.3
Identities = 22/93 (23%), Positives = 34/93 (35%), Gaps = 1/93 (1%)
Frame = +1
Query: 4 FLSNKFACHLLAN-TLKQKIHETLHAKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFR 62
F+S+ F L +N K H L K PP P I +TP N +
Sbjct: 196 FISSSFIFFLYSNQNPKA*PHHHLQLK--------PPPPPAIGTLTPPPTENAYASADPS 351
Query: 63 TAKDNEMREAEEERLRKEFEMEANKFFLNKNVK 95
NE A+ + +F + ++ N+K
Sbjct: 352 NPPRNENASADPSNTKHQFHPQIHQIHRTPNIK 450
>TC19036
Length = 613
Score = 23.5 bits (49), Expect = 6.3
Identities = 8/10 (80%), Positives = 10/10 (100%)
Frame = +3
Query: 35 IAPPPTLPDI 44
IAPPP+LPD+
Sbjct: 168 IAPPPSLPDL 197
>TC8017 weakly similar to UP|Q7UAE4 (Q7UAE4) Invasion plasmid antigen,
partial (4%)
Length = 678
Score = 23.5 bits (49), Expect = 6.3
Identities = 10/36 (27%), Positives = 19/36 (52%)
Frame = -2
Query: 47 VTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFE 82
+T +MN + TRR + K ++ + +L +E E
Sbjct: 494 ITCNVRMNNKSTRRPKLMKAHQNEKGRRNKLLQEME 387
>TC12805
Length = 702
Score = 23.5 bits (49), Expect = 6.3
Identities = 12/43 (27%), Positives = 17/43 (38%)
Frame = +1
Query: 24 ETLHAKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKD 66
+ +H K +W PP P V K+ RR R K+
Sbjct: 52 DEIHKKIEKWQEPPPAKQPKPLPVPDSEPKKKRGGRRLRKMKE 180
>AI967536
Length = 431
Score = 23.5 bits (49), Expect = 6.3
Identities = 13/49 (26%), Positives = 20/49 (40%)
Frame = +2
Query: 37 PPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEA 85
PPP LP+ VT R T R+ + + + +R E + A
Sbjct: 47 PPPNLPETTTVTRRHHRRAPNTGPPRSPR*KQWKRGGAASIRSEHGLRA 193
>TC7849 similar to GB|AAN18126.1|23308313|BT000557 At2g10940/F15K19.1
{Arabidopsis thaliana;}, partial (55%)
Length = 1157
Score = 23.1 bits (48), Expect = 8.2
Identities = 11/19 (57%), Positives = 12/19 (62%), Gaps = 3/19 (15%)
Frame = +2
Query: 34 LIAPP---PTLPDIDGVTP 49
+ PP PTLP I GVTP
Sbjct: 338 ITVPPVVKPTLPPIPGVTP 394
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,683,927
Number of Sequences: 28460
Number of extensions: 17660
Number of successful extensions: 124
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 124
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 124
length of query: 99
length of database: 4,897,600
effective HSP length: 75
effective length of query: 24
effective length of database: 2,763,100
effective search space: 66314400
effective search space used: 66314400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 47 (22.7 bits)
Medicago: description of AC130804.6


