

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130802.9 + phase: 0 /pseudo
(937 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
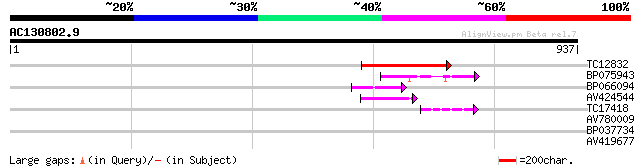
Sequences producing significant alignments: (bits) Value
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 130 7e-31
BP075943 76 3e-14
BP066094 71 9e-13
AV424544 55 7e-08
TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partia... 44 9e-05
AV780009 39 0.003
BP037734 31 1.0
AV419677 30 1.4
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 130 bits (328), Expect = 7e-31
Identities = 66/149 (44%), Positives = 92/149 (61%)
Frame = +2
Query: 582 FDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASSSKDEIMKLKERLNGEF 641
FD F++ G+ R D C Y + ++ + LLLYVDD+L+ +KD + +LK +L EF
Sbjct: 2 FDSFIMSLGYNRLSSDHCTYHKRFDDNDFIILLLYVDDMLVVGPNKDRVQELKAQLAREF 181
Query: 642 EMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSNSKTVSTPLGHHTKLSI 701
+MKDLGPA ++LG+ I R+R ++LSQ YL+K RF M + +STPL + KLS
Sbjct: 182 DMKDLGPANKILGMQIHRDRKDRRIWLSQKNYLQKVLRRFNMQDCNPISTPLPVNYKLSS 361
Query: 702 QQCPQSEDEKQLMEGTPYASGVGSIMYGM 730
P SE E+ M PYAS VGS+MY M
Sbjct: 362 SMIPSSEAERMEMSRVPYASAVGSLMYAM 448
>BP075943
Length = 547
Score = 75.9 bits (185), Expect = 3e-14
Identities = 62/178 (34%), Positives = 90/178 (49%), Gaps = 14/178 (7%)
Frame = -3
Query: 613 LLLYVDDILMASSSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKR-------NRDKGE 665
LLLYVDD+L+A + EI +K L+ +F +KDLG AK LG++I R N+ K
Sbjct: 521 LLLYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIARSTSGIVLNQRKYA 342
Query: 666 L-FLSQLGYLKKGGERFRMSNSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTP------ 718
L +S G+ G + R + T S LG +T GTP
Sbjct: 341 LQLISDSGHF---GFQPRFYSHGTNSQTLGTNT------------------GTPLTDIGS 225
Query: 719 YASGVGSIMYGMVCSRPDLAYAVSIVSRFMANPGIVHWQALKWVLRYLNGSLKGGLKY 776
Y VG ++Y + +RPD+ +AV+ +S+F++ P +H Q L L+YL GS GL Y
Sbjct: 224 YRRIVGRLLY-LNTTRPDITFAVNQLSQFLSAPTDIHEQQLTGFLKYL*GSPGSGLFY 54
>BP066094
Length = 532
Score = 70.9 bits (172), Expect = 9e-13
Identities = 37/91 (40%), Positives = 54/91 (58%)
Frame = +2
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQ+PR WY R FLL+ VR D+ ++ K + IL + +YVDDI+ S
Sbjct: 260 KKSLYGLKQAPRAWYERLSSFLLENEXVRGKVDTTLF-CKTYKDDILIVQIYVDDIIFGS 436
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGI 655
++ + E + EFEM+ +G K LGI
Sbjct: 437 ANPSLCKEFSEMMQAEFEMRMMGELKYFLGI 529
>AV424544
Length = 276
Score = 54.7 bits (130), Expect = 7e-08
Identities = 30/93 (32%), Positives = 55/93 (58%)
Frame = +3
Query: 581 RFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASSSKDEIMKLKERLNGE 640
+ +L G+++S +D ++ K + +L+YVDD+++A + +EI +K +L+ +
Sbjct: 6 KLSSYLHILGYIQSAHDHSLFT-KFRDASFTVILVYVDDLILAGNDLNEIQCVKNKLDIQ 182
Query: 641 FEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGY 673
F +KDLG K LG+++ R+ LFLSQ Y
Sbjct: 183 FRIKDLGTLKYFLGLEVARS--SCGLFLSQRKY 275
>TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partial (20%)
Length = 739
Score = 44.3 bits (103), Expect = 9e-05
Identities = 32/96 (33%), Positives = 54/96 (55%)
Frame = -1
Query: 679 ERFRMSNSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLA 738
++F+M+NSK +ST +G I+ ++ ++ ++ T Y S +GS+ Y + R D+
Sbjct: 712 KKFKMTNSKYISTTIGGK---EIEAGRRNGGKR--VDSTYYKSLIGSVRY-LNTVRSDIV 551
Query: 739 YAVSIVSRFMANPGIVHWQALKWVLRYLNGSLKGGL 774
V + SRFM P H Q + LRY+ G+LK G+
Sbjct: 550 CGVGLRSRFM-EP*DCH*QGAQRSLRYIKGTLKDGI 446
>AV780009
Length = 529
Score = 39.3 bits (90), Expect = 0.003
Identities = 21/54 (38%), Positives = 33/54 (60%)
Frame = -1
Query: 757 QALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLVGFCVY 810
QA VLRY+ G+ GL ++ A L+ Y D+D+AG DTR+S+ G+ ++
Sbjct: 529 QAATRVLRYVKGAPAQGLFFS--ADSPLKLQAYSDSDWAGCPDTRRSVTGYSIF 374
>BP037734
Length = 490
Score = 30.8 bits (68), Expect = 1.0
Identities = 11/15 (73%), Positives = 13/15 (86%)
Frame = +2
Query: 568 LYGLKQSPREWYRRF 582
LYGLKQ PR+W+ RF
Sbjct: 194 LYGLKQLPRDWFERF 238
>AV419677
Length = 411
Score = 30.4 bits (67), Expect = 1.4
Identities = 13/50 (26%), Positives = 24/50 (48%)
Frame = +3
Query: 724 GSIMYGMVCSRPDLAYAVSIVSRFMANPGIVHWQALKWVLRYLNGSLKGG 773
G + VC + L ++V + R ++ +Q W+L Y N S++ G
Sbjct: 69 GCLRLFYVCQKESLGFSVKVNQRSCLPNYLIRFQNFWWILHYTNNSMR*G 218
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.356 0.161 0.595
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,867,052
Number of Sequences: 28460
Number of extensions: 226867
Number of successful extensions: 1950
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1401
Number of HSP's successfully gapped in prelim test: 45
Number of HSP's that attempted gapping in prelim test: 506
Number of HSP's gapped (non-prelim): 1492
length of query: 937
length of database: 4,897,600
effective HSP length: 99
effective length of query: 838
effective length of database: 2,080,060
effective search space: 1743090280
effective search space used: 1743090280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC130802.9


