

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.5 + phase: 0
(569 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
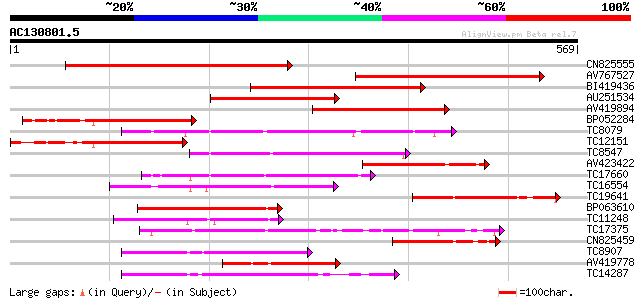
Score E
Sequences producing significant alignments: (bits) Value
CN825555 398 e-111
AV767527 349 8e-97
BI419436 267 4e-72
AU251534 253 4e-68
AV419894 224 3e-59
BP052284 183 7e-47
TC8079 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, complete 170 7e-43
TC12151 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {Ar... 159 9e-40
TC8547 homologue to UP|MMK1_MEDSA (Q07176) Mitogen-activated pro... 144 5e-35
AV423422 134 4e-32
TC17660 homologue to UP|Q9M6R8 (Q9M6R8) MAP kinase PsMAPK2, part... 134 5e-32
TC16554 homologue to UP|AAN71719 (AAN71719) Glycogen synthase ki... 130 4e-31
TC19641 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {Ar... 126 1e-29
BP063610 121 3e-28
TC11248 homologue to UP|P93323 (P93323) Cdc2MsF protein, partial... 112 1e-25
TC17375 mitogen-activated kinase kinase kinase alpha [Lotus corn... 109 1e-24
CN825459 100 5e-22
TC8907 similar to UP|Q8H0X3 (Q8H0X3) Serine/threonine protein ki... 100 5e-22
AV419778 99 2e-21
TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein k... 99 2e-21
>CN825555
Length = 714
Score = 398 bits (1023), Expect = e-111
Identities = 187/227 (82%), Positives = 211/227 (92%)
Frame = +1
Query: 57 QRSRRSKPNPRLSNPPKHLRGEQVAAGWPSWLTAVCGEALTGWIPRKADTFEKIDKIGQG 116
+R R SKPNPRLSNPP ++ GEQVAAGWPSWL+ V GEA+ G PR+ADTFEK+DKIGQG
Sbjct: 34 RRRRSSKPNPRLSNPPNNVHGEQVAAGWPSWLSKVAGEAINGLTPRRADTFEKLDKIGQG 213
Query: 117 TYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSR 176
TYSNVYKA D++TGK+VALKKVRFDNLEPES+KFMAREI+ILRRLDHPNV+KL+GLVTSR
Sbjct: 214 TYSNVYKARDTLTGKIVALKKVRFDNLEPESVKFMAREILILRRLDHPNVLKLEGLVTSR 393
Query: 177 MSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSN 236
MSCSLYLVF+YM HDLAGLA +P I+FTESQ+KCYM+QL +GLEHCHNR VLHRDIKGSN
Sbjct: 394 MSCSLYLVFEYMVHDLAGLATNPAIKFTESQVKCYMHQLFTGLEHCHNRHVLHRDIKGSN 573
Query: 237 LLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGAT 283
LLIDNEG+LKIADFGLASFFDP++ +PMTSRVVTLWYRPPELLLGAT
Sbjct: 574 LLIDNEGVLKIADFGLASFFDPDHKHPMTSRVVTLWYRPPELLLGAT 714
>AV767527
Length = 573
Score = 349 bits (895), Expect = 8e-97
Identities = 168/190 (88%), Positives = 177/190 (92%), Gaps = 1/190 (0%)
Frame = +3
Query: 348 PYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKYP 407
PYKRCIRE FK FPPSALPLID LLAIDPVER TASDALRSEFFTTEPYACDPSSLPKYP
Sbjct: 3 PYKRCIREIFKDFPPSALPLIDTLLAIDPVERRTASDALRSEFFTTEPYACDPSSLPKYP 182
Query: 408 PSKEMDAKRRDDEVRRQRAASKAQVDGSKKH-RTRERSMKAMPAPEANAELQSNIDRRRL 466
PSKEMDAKRRDDE+RRQRAA KAQ DGSKKH RTR+R++KA APEANAELQSNIDRRRL
Sbjct: 183 PSKEMDAKRRDDEMRRQRAAGKAQADGSKKHHRTRDRAVKAFAAPEANAELQSNIDRRRL 362
Query: 467 ITHANAKSKSEKFPPPHQDGQLGFPLGSSHHIDPDTIPADISFTSTTYTYSKEPFQAWSG 526
ITHANAKSKSEKFPPPHQDGQLGFPLG+SHHIDPDT+P D+SFTS +Y + KEPFQAWSG
Sbjct: 363 ITHANAKSKSEKFPPPHQDGQLGFPLGASHHIDPDTVPTDVSFTSVSYNFPKEPFQAWSG 542
Query: 527 PIGNAADIGV 536
PIGNAADIGV
Sbjct: 543 PIGNAADIGV 572
>BI419436
Length = 601
Score = 267 bits (682), Expect = 4e-72
Identities = 117/176 (66%), Positives = 148/176 (83%)
Frame = +1
Query: 242 EGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGEL 301
EG+LK+ADFGLA+F + P+TSRVVTLWYRPPELLLGATDYG +DLWS GC+ EL
Sbjct: 13 EGVLKVADFGLANFTSSGHKQPLTSRVVTLWYRPPELLLGATDYGPSVDLWSVGCVFAEL 192
Query: 302 LVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFP 361
LVGKP++ GRTEVEQLHKI+KLCGSP ++YWKK++LP+ATLFKP+ PY C+RE+FK P
Sbjct: 193 LVGKPVLQGRTEVEQLHKIFKLCGSPPEDYWKKTRLPHATLFKPQHPYDSCLRESFKDLP 372
Query: 362 PSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRR 417
P+++ L+ LL+++P +R TA+ AL SE+F T+PYAC+PSSLP YPPSKE+DAK R
Sbjct: 373 PASVNLLQTLLSVEPYKRGTATSALSSEYFRTKPYACEPSSLPTYPPSKEIDAKHR 540
>AU251534
Length = 391
Score = 253 bits (647), Expect = 4e-68
Identities = 118/130 (90%), Positives = 124/130 (94%)
Frame = +2
Query: 202 RFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYM 261
RFTE Q+KCYM QLLSGLEHCH+RRVLHRDIKGSNLLIDNEG+LKIADFGLAS FDPN+
Sbjct: 2 RFTEPQVKCYMRQLLSGLEHCHHRRVLHRDIKGSNLLIDNEGVLKIADFGLASVFDPNHK 181
Query: 262 NPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIY 321
+PMTSRVVTLWYRPPELLLGATDY VG+DLWSAGCILGELL GKPIMPGRTEVEQLHKIY
Sbjct: 182 HPMTSRVVTLWYRPPELLLGATDYDVGVDLWSAGCILGELLAGKPIMPGRTEVEQLHKIY 361
Query: 322 KLCGSPSDEY 331
KLCGSPSDEY
Sbjct: 362 KLCGSPSDEY 391
>AV419894
Length = 413
Score = 224 bits (571), Expect = 3e-59
Identities = 106/137 (77%), Positives = 119/137 (86%)
Frame = +1
Query: 305 KPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSA 364
+PIMPGRTEVEQLHKI+KLCGSPS+EYWKKSKLP+AT+FKP++ YKRCI ETFK FPPS+
Sbjct: 1 RPIMPGRTEVEQLHKIFKLCGSPSEEYWKKSKLPHATIFKPQQSYKRCIAETFKNFPPSS 180
Query: 365 LPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQ 424
LPLI+ LL IDP ER TA+ AL SEFFTT+PYAC+PSSLPKYPPSKEMDAK RD+E RR
Sbjct: 181 LPLIETLLTIDPDERLTATAALHSEFFTTKPYACEPSSLPKYPPSKEMDAKLRDEEARRL 360
Query: 425 RAASKAQVDGSKKHRTR 441
RAA KA DG KK R R
Sbjct: 361 RAAGKANADGVKKSRPR 411
>BP052284
Length = 550
Score = 183 bits (464), Expect = 7e-47
Identities = 101/177 (57%), Positives = 124/177 (69%), Gaps = 3/177 (1%)
Frame = +1
Query: 14 GSGAQTNRIKVDEASAATTASNGEEKNVVEIENDQKKKSDDSVQRSRRSKPNPRLSNPPK 73
G GAQ R + S AS+G N V + ++KK++ + P P P
Sbjct: 43 GDGAQDQRRR--RKSERNVASDGGN-NAVSVRV-REKKTNRHTGDFPGTVPAPERRKP-- 204
Query: 74 HLRGEQVAA---GWPSWLTAVCGEALTGWIPRKADTFEKIDKIGQGTYSNVYKAIDSMTG 130
R + +A GWP+WL AV GEA+ W PR+A +FEK+ KIGQGTYSNVYKA D +TG
Sbjct: 205 --RLDPLAVTQQGWPAWLMAVAGEAIGDWTPRRASSFEKLAKIGQGTYSNVYKAKDLLTG 378
Query: 131 KVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLVFDY 187
K+VALKKVRFDNLEPES+KFMAREI++LRRLDHPNV+KL+GLVTSRMSCSLYLVF+Y
Sbjct: 379 KIVALKKVRFDNLEPESVKFMAREILVLRRLDHPNVVKLEGLVTSRMSCSLYLVFEY 549
>TC8079 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, complete
Length = 1550
Score = 170 bits (430), Expect = 7e-43
Identities = 120/351 (34%), Positives = 184/351 (52%), Gaps = 15/351 (4%)
Frame = +3
Query: 113 IGQGTYSNVYKAIDSMTGKVVALKKVR--FDNLEPESIKFMAREIIILRRLDHPNVIKLQ 170
IG+G Y V +++ T ++VA+KK+ FDN K REI +LR LDH NVI L+
Sbjct: 309 IGRGAYGIVCSLLNTETNELVAVKKIANAFDN--HMDAKRTLREIKLLRHLDHENVIALR 482
Query: 171 GLVTS---RMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRV 227
++ R +Y+ + M+ DL + S +E + ++ Q+L GL++ H+ +
Sbjct: 483 DVIPPPLRREFTDVYITTELMDTDLHQIIRSNQ-GLSEEHCQYFLYQVLRGLKYIHSANI 659
Query: 228 LHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNP-MTSRVVTLWYRPPELLLGATDYG 286
+HRD+K SNLL+++ LKI DFGLA P N MT VVT WYR PELLL ++DY
Sbjct: 660 IHRDLKPSNLLLNSNCDLKIIDFGLAR---PTVENDFMTEYVVTRWYRAPELLLNSSDYT 830
Query: 287 VGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLF--- 343
ID+WS GCI EL+ KP+ PG+ V QL + +L G+P++ K + +
Sbjct: 831 SAIDVWSVGCIFMELMNKKPLFPGKDHVHQLRLLTELLGTPTEADLGLVKNDDVRRYIRQ 1010
Query: 344 ---KPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDP 400
PR+P + + F P A+ L+DK+L IDP +R T AL + D
Sbjct: 1011LPQYPRQP----LTKVFPHVHPMAMDLVDKMLTIDPTKRITVEQALAHPYLEKLHDVADE 1178
Query: 401 SSLPKYPPSKEMDAKRRDDEVRRQ---RAASKAQVDGSKKHRTRERSMKAM 448
K P S E + ++ D+E ++ R A + + + +TRE + +
Sbjct: 1179PVCTK-PFSFEFEQQQLDEEQIKEMIYREALALNPEYA*RRKTREDKKRTL 1328
>TC12151 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {Arabidopsis
thaliana;}, partial (17%)
Length = 708
Score = 159 bits (403), Expect = 9e-40
Identities = 91/181 (50%), Positives = 113/181 (62%), Gaps = 3/181 (1%)
Frame = +3
Query: 1 MGCVIGRQASSNKGSGAQTNRIKVDEASAATTASNGEEKNVVEIENDQKKKSDDSVQRSR 60
MGC +G A + D + A+ G KN V + + QK +
Sbjct: 234 MGCALGTPAVAG------------DRRRRSPAAAEGG-KNAVSVPDRQKDPNAGEA---- 362
Query: 61 RSKPNPRLSNPPKHLRGEQVAA---GWPSWLTAVCGEALTGWIPRKADTFEKIDKIGQGT 117
P P L + R + A GWP WL AV G+A+ W PR+A+TFEK+ KIGQGT
Sbjct: 363 ---PAPEL----RKYRLDSFTATHQGWPPWLMAVAGDAIRDWTPRRANTFEKLAKIGQGT 521
Query: 118 YSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRM 177
YSNVYKA D +TGK+VALKKVRFDNLE ES+KFMAREI++LRRLDHPNV+KL+GLVTSR+
Sbjct: 522 YSNVYKARDLVTGKIVALKKVRFDNLEAESVKFMAREILVLRRLDHPNVVKLEGLVTSRI 701
Query: 178 S 178
S
Sbjct: 702 S 704
>TC8547 homologue to UP|MMK1_MEDSA (Q07176) Mitogen-activated protein
kinase homolog MMK1 (MAP kinase MSK7) (MAP kinase ERK1)
, partial (71%)
Length = 1098
Score = 144 bits (362), Expect = 5e-35
Identities = 86/229 (37%), Positives = 124/229 (53%), Gaps = 7/229 (3%)
Frame = +1
Query: 181 LYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLID 240
+Y+ ++ M+ DL + S +E + ++ Q+L GL++ H+ VLHRD+K SNLL++
Sbjct: 64 VYIAYELMDTDLHQIIRSNQ-GLSEEHCQYFLYQILRGLKYIHSANVLHRDLKPSNLLLN 240
Query: 241 NEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGE 300
LKI DFGLA MT VVT WYR PELLL ++DY ID+WS GCI E
Sbjct: 241 ANCDLKICDFGLARVTSETDF--MTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFME 414
Query: 301 LLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKR-CIRETFKG 359
L+ KP+ PGR V QL + +L G+PS++ + PY+R +E F
Sbjct: 415 LMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFLNENAKRYIRQLPPYRRQSFQEKFPQ 594
Query: 360 FPPSALPLIDKLLAIDPVERETASDALRSEFFTT------EPYACDPSS 402
P A+ L++K+L DP +R T +AL + T+ EP P S
Sbjct: 595 VHPEAIDLVEKMLTFDPRKRITVEEALAHPYLTSLHDISDEPVCMTPFS 741
>AV423422
Length = 375
Score = 134 bits (337), Expect = 4e-32
Identities = 73/128 (57%), Positives = 96/128 (74%), Gaps = 1/128 (0%)
Frame = +1
Query: 355 ETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKYPPSKEMDA 414
ETFK FP A+ LI+ LL+IDP +R T++ AL+SEFF+T+P CDPSSLPKYPPSKE DA
Sbjct: 1 ETFKDFPAPAIELIETLLSIDPTDRGTSASALKSEFFSTKPLPCDPSSLPKYPPSKEFDA 180
Query: 415 KRRDDEVRRQRAA-SKAQVDGSKKHRTRERSMKAMPAPEANAELQSNIDRRRLITHANAK 473
K R++E RRQ AA SKAQ ++ R R +A+PAPEANAEL ++ +RR +T +++
Sbjct: 181 KVREEEARRQGAAGSKAQRHDPER---RVRESRAIPAPEANAELVLSMQKRRGLT--SSQ 345
Query: 474 SKSEKFPP 481
S+SEKF P
Sbjct: 346 SRSEKFNP 369
>TC17660 homologue to UP|Q9M6R8 (Q9M6R8) MAP kinase PsMAPK2, partial (64%)
Length = 717
Score = 134 bits (336), Expect = 5e-32
Identities = 93/240 (38%), Positives = 132/240 (54%), Gaps = 5/240 (2%)
Frame = +2
Query: 133 VALKKVR--FDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCS---LYLVFDY 187
VA+KK++ F+N ++++ + RE+ +LR L H NVI L+ ++ S +YLV++
Sbjct: 8 VAIKKIQNAFEN-RVDALRTL-RELKLLRHLHHDNVIALKDIMMPVHRSSFKDVYLVYEL 181
Query: 188 MEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKI 247
M+ DL + S + + ++ QLL GL++ H+ +LHRD+K NLLI+ LKI
Sbjct: 182 MDTDLHQIIKSSQ-SLSNDHCQYFLFQLLRGLKYLHSANILHRDLKPGNLLINANCDLKI 358
Query: 248 ADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPI 307
DFGLA + + MT VVT WYR PELLL +YG ID+WS GCI ELL KPI
Sbjct: 359 CDFGLARI-NCSKNQFMTEYVVTRWYRAPELLLCCDNYGTSIDVWSVGCIFAELLGRKPI 535
Query: 308 MPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSALPL 367
PG + QL I + GS +E + P A + PY I F P+A PL
Sbjct: 536 FPGSECLNQLKLIINILGSQREEDIEFIDNPKAKKYIKSLPYS--IGAPFSRLYPNAHPL 709
>TC16554 homologue to UP|AAN71719 (AAN71719) Glycogen synthase kinase 3 beta
protein kinase DWARF12 , partial (71%)
Length = 927
Score = 130 bits (328), Expect = 4e-31
Identities = 82/238 (34%), Positives = 130/238 (54%), Gaps = 8/238 (3%)
Frame = +1
Query: 101 PRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRR 160
P++ ++ +G G++ V++A +G+ VA+KKV D ++ RE+ ++R
Sbjct: 220 PKQTISYMAERVVGTGSFGIVFQAKCLESGEAVAIKKVLQDR------RYKNRELQLMRV 381
Query: 161 LDHPNVIKLQGLVTSRMSCS---LYLVFDYMEHDLAGLA---ASPVIRFTESQIKCYMNQ 214
+DHPNV+ L+ S S L LV +Y+ + + ++ R +K YM Q
Sbjct: 382 MDHPNVVSLKHCFFSTTSTDELFLNLVMEYVPESMYRVIKHYSNANQRMPIIYVKLYMYQ 561
Query: 215 LLSGLEHCHN-RRVLHRDIKGSNLLIDN-EGILKIADFGLASFFDPNYMNPMTSRVVTLW 272
+ GL + H V HRD+K N+L+D +K+ DFG A N S + + +
Sbjct: 562 IFRGLAYIHTVPGVCHRDLKPQNILVDPLTHQVKLCDFGSAKMLVKGEAN--ISYICSRF 735
Query: 273 YRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDE 330
YR PEL+ GAT+Y ID+WSAGC+L ELL+G+P+ PG V+QL I K+ G+P+ E
Sbjct: 736 YRAPELIFGATEYSTSIDIWSAGCVLAELLLGQPLFPGENAVDQLVHIIKVLGTPTRE 909
>TC19641 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {Arabidopsis
thaliana;}, partial (15%)
Length = 823
Score = 126 bits (316), Expect = 1e-29
Identities = 75/154 (48%), Positives = 99/154 (63%), Gaps = 6/154 (3%)
Frame = +3
Query: 405 KYPPSKEMDAKRRDDEVRRQRAASKAQ--VDGSKKHRTRERSMKAMPAPEANAELQSNID 462
KYPPSKE+D K RD+ RR++A S VDG+K+ RTRER A+PAPEANAE+Q+N+D
Sbjct: 3 KYPPSKELDVKLRDEAARRRKALSGKNNAVDGTKRVRTRERGC-AIPAPEANAEIQTNLD 179
Query: 463 RRRLITHANAKSKSEKFPPPHQDGQLGFPLGSSHHIDPDTIPADISFTSTTYTYSKEPFQ 522
R R++T ANAKSKSEKFPPPHQDG +G+PL +S+ + SF S+T + SK
Sbjct: 180 RMRVVTRANAKSKSEKFPPPHQDGAVGYPLDASNKGAVSFGATETSF-SSTISNSKP--- 347
Query: 523 AWSGPIGNAADIGVSKRKKYTAGE----AWDLLK 552
SG +G+ A K AG +W L++
Sbjct: 348 --SGSVGSYASSSYRGWKTNKAGSHSAPSWKLMR 443
>BP063610
Length = 461
Score = 121 bits (304), Expect = 3e-28
Identities = 65/145 (44%), Positives = 92/145 (62%)
Frame = -3
Query: 129 TGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYM 188
TG++VALKKV + + REI IL HP ++ ++ +V S+++V +YM
Sbjct: 459 TGEIVALKKVXMEKEKEGFPLTSLREINILLSFHHPYIVDVKEVVVGSSLDSIFMVMEYM 280
Query: 189 EHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIA 248
EHDL GL S F++S++KC M QLL G+++ H+ VLHRD+K SNLL++N G LKI
Sbjct: 279 EHDLKGLMESMKQPFSQSEVKCLMIQLLEGVKYLHDNWVLHRDLKTSNLLLNNRGELKIC 100
Query: 249 DFGLASFFDPNYMNPMTSRVVTLWY 273
DFGLA + + + P T VVTLWY
Sbjct: 99 DFGLARXYG-SPLKPYTHLVVTLWY 28
>TC11248 homologue to UP|P93323 (P93323) Cdc2MsF protein, partial (72%)
Length = 747
Score = 112 bits (281), Expect = 1e-25
Identities = 72/180 (40%), Positives = 99/180 (55%), Gaps = 10/180 (5%)
Frame = +1
Query: 105 DTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDH- 163
+ FEK++K+G+GTY VY+A + TGK+VALKK R + RE+ ILR L
Sbjct: 43 EAFEKLEKVGEGTYGKVYRAREKATGKIVALKKTRLHEDDEGVPPTTLREVSILRMLSRD 222
Query: 164 PNVIKLQGLVTSRM---SCSLYLVFDYMEHDLAGLAASPVIRFT-----ESQIKCYMNQL 215
P+V++L + + LYLVF+YM+ DL + R T +K M QL
Sbjct: 223 PHVVRLMDVKQGQSKEGKTVLYLVFEYMDTDLKKFIRT--FRQTGQNVPPKTVKSLMYQL 396
Query: 216 LSGLEHCHNRRVLHRDIKGSNLLIDNE-GILKIADFGLASFFDPNYMNPMTSRVVTLWYR 274
G+ CH +LHRD+K NLL+D E +LKIAD GLA F + T ++TLWYR
Sbjct: 397 RKGVAFCHGHGILHRDLKPHNLLMDRETNMLKIADLGLARAFTVP-IKKYTHEILTLWYR 573
Score = 37.7 bits (86), Expect = 0.005
Identities = 18/46 (39%), Positives = 25/46 (54%), Gaps = 5/46 (10%)
Frame = +2
Query: 262 NPMTSRVVTLWYRPP-----ELLLGATDYGVGIDLWSAGCILGELL 302
+P R + Y P E+LLGAT Y + +D+WS CI EL+
Sbjct: 521 SPCPLRSTRMRYSPSGTELAEVLLGATHYSMAVDMWSVACIFAELV 658
>TC17375 mitogen-activated kinase kinase kinase alpha [Lotus corniculatus
var. japonicus]
Length = 1422
Score = 109 bits (272), Expect = 1e-24
Identities = 100/378 (26%), Positives = 172/378 (45%), Gaps = 12/378 (3%)
Frame = +3
Query: 131 KVVALKKVRF---DNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLVFDY 187
++ A+K+V+ D E +K + +EI +L + HPN+++ G S S+YL +Y
Sbjct: 3 QMCAIKEVKVFSDDKTSKECLKQLNQEINLLNQFSHPNIVQYYGSELGEESLSVYL--EY 176
Query: 188 MEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKI 247
+ F E I+ Y Q++SGL + H+R +HRDIKG+N+L+D G +K+
Sbjct: 177 VSGGSIHKLLQEYGAFKEPVIQNYTRQIVSGLAYLHSRNTVHRDIKGANILVDPNGEIKL 356
Query: 248 ADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPI 307
ADFG++ N M S + ++ PE+++ YG+ +D+ S GC + E+ KP
Sbjct: 357 ADFGMSKHI--NSAASMLSFKGSPYWMAPEVVMNTNGYGLPVDISSLGCTILEMATSKPP 530
Query: 308 MPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSALPL 367
++ E + I+K+ G+ D ++P + K+C++ P A P
Sbjct: 531 W---SQFEGVAAIFKI-GNSKD----MPEIPEHLSDDAKNFIKQCLQR-----DPLARPT 671
Query: 368 IDKLLAIDPVERETASDALRSEFFTTE--PYACDPSSLPKYPPSKEMDAKRRDDEVRRQR 425
LL P R+ ++ + + T + PY D S + PP E + R
Sbjct: 672 AQSLLN-HPFIRDQSATKVANASITRDAFPYMSDGS---RTPPVLEPHSNRSSITTLDVD 839
Query: 426 AASK---AQVDGSKKHRTRERSMKAMPAPEANAELQSNIDRRRLITHANAKSKSEKFPPP 482
A+K A V + R R++ ++P +++ L R+ TH S F PP
Sbjct: 840 YATKPALAAVRTLRNPRDSTRTITSLPVSPSSSPL-----RQHRPTH-----NSPFFSPP 989
Query: 483 HQD----GQLGFPLGSSH 496
H GQ + + H
Sbjct: 990 HPSYAMMGQSSYTSNNMH 1043
>CN825459
Length = 696
Score = 100 bits (250), Expect = 5e-22
Identities = 54/108 (50%), Positives = 75/108 (69%)
Frame = +1
Query: 385 ALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQVDGSKKHRTRERS 444
AL++EFFT +P CDPS+LPKYPPSKE DAK R++E RR+RA++K S RE +
Sbjct: 7 ALQNEFFTAKPLPCDPSTLPKYPPSKEFDAKLREEEARRRRASNKGHGQESVGRNFRESN 186
Query: 445 MKAMPAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQDGQLGFPL 492
+ +PA +ANAELQ+++++R+ +K SEKF P +DG GFPL
Sbjct: 187 V--VPASDANAELQASMEKRQ----EQSKCISEKF-NPEEDGDYGFPL 309
>TC8907 similar to UP|Q8H0X3 (Q8H0X3) Serine/threonine protein kinase-like
protein, partial (44%)
Length = 941
Score = 100 bits (250), Expect = 5e-22
Identities = 62/194 (31%), Positives = 110/194 (55%), Gaps = 2/194 (1%)
Frame = +3
Query: 113 IGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPES-IKFMAREIIILRRLDHPNVIKLQG 171
+GQG ++ VY + T + VA+K ++ + L+ E +K + RE+ ++R + HP++++L+
Sbjct: 360 LGQGNFAKVYHGRNMETNESVAIKVIKKERLKKERLVKQIKREVSVMRLVRHPHIVELKE 539
Query: 172 LVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRD 231
++ ++ +++V +Y++ A + TE + Y QL+S ++ CH+R V HRD
Sbjct: 540 VMATKTK--IFMVVEYVKGGEL-FAKLTKGKMTEVAARKYFQQLISAVDFCHSRGVTHRD 710
Query: 232 IKGSNLLIDNEGILKIADFGLASFFDPNYMNPM-TSRVVTLWYRPPELLLGATDYGVGID 290
+K NLL+D+ LK++DFGL+S + + M + T Y PE+L G D
Sbjct: 711 LKPENLLLDDNEDLKVSDFGLSSLPEQRRSDGMLLTPCGTPAYVAPEVLKKKGYDGSKAD 890
Query: 291 LWSAGCILGELLVG 304
+WS G IL LL G
Sbjct: 891 IWSCGVILYALLCG 932
>AV419778
Length = 411
Score = 99.0 bits (245), Expect = 2e-21
Identities = 48/119 (40%), Positives = 72/119 (60%)
Frame = +1
Query: 214 QLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWY 273
Q+ GL + H R HRD+K NLL+ + I+K++DFGLA + P T V T WY
Sbjct: 16 QVFQGLAYMHQRGYFHRDLKPENLLVTKD-IIKVSDFGLAREISSH--PPYTEYVSTRWY 186
Query: 274 RPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYW 332
R PE+LL + Y +D+W+ G I+ EL +P+ PG +E ++++KI + GSP+ E W
Sbjct: 187 RAPEVLLQSYLYSSKVDMWAMGAIMAELFTLRPLFPGTSEADEIYKICSVIGSPTTESW 363
>TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial
(75%)
Length = 1999
Score = 98.6 bits (244), Expect = 2e-21
Identities = 78/283 (27%), Positives = 136/283 (47%), Gaps = 4/283 (1%)
Frame = +3
Query: 113 IGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPES-IKFMAREIIILRRLDHPNVIKLQG 171
+GQG ++ VY + T + VA+K ++ + L+ + +K + RE+ ++R + HP++++L+
Sbjct: 450 LGQGNFAKVYHGRNLATNENVAIKVIKKEKLKKDRLVKQIKREVSVMRLVRHPHIVELKE 629
Query: 172 LVTSRMSCSLYLVFDYMEHDLAGLAASPVIR--FTESQIKCYMNQLLSGLEHCHNRRVLH 229
++ ++ ++LV +Y++ G + V + E + Y QL+S ++ CH+R V H
Sbjct: 630 VMATKGK--IFLVMEYVK---GGELFTKVNKGKLNEDDARKYFQQLISAVDFCHSRGVTH 794
Query: 230 RDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPM-TSRVVTLWYRPPELLLGATDYGVG 288
RD+K NLL+D LK++DFGL++ + + M + T Y PE+L G
Sbjct: 795 RDLKPENLLLDENEDLKVSDFGLSALPEQRRDDGMLVTPCGTPAYVAPEVLKKKGYDGSK 974
Query: 289 IDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREP 348
D+WS G IL LL G G E + +IY+ FK
Sbjct: 975 ADIWSCGVILYALLSGYLPFQG----ENVMRIYR------------------KAFKAEYE 1088
Query: 349 YKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFF 391
+ I P A LI LL DP +R + + + +F
Sbjct: 1089FPEWI-------SPQAKNLISNLLVADPEKRYSIPEIISDPWF 1196
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,037,518
Number of Sequences: 28460
Number of extensions: 145366
Number of successful extensions: 1081
Number of sequences better than 10.0: 196
Number of HSP's better than 10.0 without gapping: 1007
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1017
length of query: 569
length of database: 4,897,600
effective HSP length: 95
effective length of query: 474
effective length of database: 2,193,900
effective search space: 1039908600
effective search space used: 1039908600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC130801.5


