

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.12 + phase: 0
(124 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
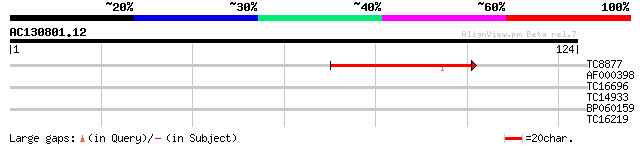
Sequences producing significant alignments: (bits) Value
TC8877 similar to UP|O65917 (O65917) Dehydroquinate dehydratase/... 40 1e-04
AF000398 31 0.071
TC16696 similar to UP|Q9AR81 (Q9AR81) Germin-like protein precur... 25 3.9
TC14933 homologue to UP|TCMO_GLYEC (Q96423) Trans-cinnamate 4-mo... 25 3.9
BP060159 24 6.6
TC16219 24 8.7
>TC8877 similar to UP|O65917 (O65917) Dehydroquinate
dehydratase/shikimate:NADP oxidoreductase
(3-dehydroquinate dehydratase) , partial (15%)
Length = 661
Score = 40.0 bits (92), Expect = 1e-04
Identities = 21/37 (56%), Positives = 23/37 (61%), Gaps = 5/37 (13%)
Frame = +3
Query: 71 GAILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
G ILAN T IGM P D PV++ Y LVFDAVY
Sbjct: 6 GMILANTTSIGMQPKVDETPVSKHALKFYSLVFDAVY 116
>AF000398
Length = 119
Score = 30.8 bits (68), Expect = 0.071
Identities = 16/22 (72%), Positives = 18/22 (81%), Gaps = 2/22 (9%)
Frame = +1
Query: 9 PLDGRLFV--RAGGAGKALAFG 28
PL G+LFV AGGAGKALA+G
Sbjct: 52 PLAGKLFVVIGAGGAGKALAYG 117
>TC16696 similar to UP|Q9AR81 (Q9AR81) Germin-like protein precursor,
partial (96%)
Length = 834
Score = 25.0 bits (53), Expect = 3.9
Identities = 12/25 (48%), Positives = 15/25 (60%)
Frame = +1
Query: 96 LVFDAVYLHFYIKLGGSKAVLFPSF 120
+VF +HF I GGS A+ F SF
Sbjct: 406 MVFPQGLVHFQIHDGGSSALAFVSF 480
>TC14933 homologue to UP|TCMO_GLYEC (Q96423) Trans-cinnamate 4-monooxygenase
(Cinnamic acid 4-hydroxylase) (CA4H) (C4H) (P450C4H)
(Cytochrome P450 73) , partial (57%)
Length = 917
Score = 25.0 bits (53), Expect = 3.9
Identities = 12/34 (35%), Positives = 20/34 (58%)
Frame = +1
Query: 23 KALAFGAKTRGARIVIFDIDFDKSRSLACAVFGE 56
+ + FG++TR V+FDI K + + V+GE
Sbjct: 337 QGVEFGSRTRN---VVFDIFTGKGQDMVFTVYGE 429
>BP060159
Length = 254
Score = 24.3 bits (51), Expect = 6.6
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = -1
Query: 98 FDAVYLHFYIKLGGSKAVLFPSFWC 122
FD + L Y+K S+ ++FP F C
Sbjct: 74 FDEIILTLYLKKSASECIIFP-FLC 3
>TC16219
Length = 430
Score = 23.9 bits (50), Expect = 8.7
Identities = 11/31 (35%), Positives = 19/31 (60%)
Frame = -1
Query: 62 NLVNFQPVKGAILANATPIGMHPDTDRIPVA 92
NL NF P + +++T I ++P+T + P A
Sbjct: 277 NLHNFHPPSSSSSSSSTHILLNPETTKNPNA 185
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.143 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,937,371
Number of Sequences: 28460
Number of extensions: 19463
Number of successful extensions: 118
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 117
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 117
length of query: 124
length of database: 4,897,600
effective HSP length: 79
effective length of query: 45
effective length of database: 2,649,260
effective search space: 119216700
effective search space used: 119216700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 50 (23.9 bits)
Medicago: description of AC130801.12


