

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.11 - phase: 0
(275 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
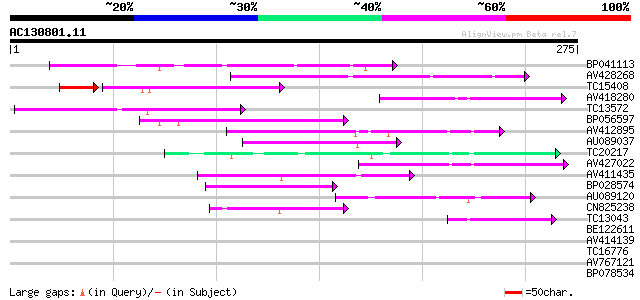
Score E
Sequences producing significant alignments: (bits) Value
BP041113 76 5e-15
AV428268 67 4e-12
TC15408 similar to UP|Q9LIC2 (Q9LIC2) Multispanning membrane pro... 57 3e-11
AV418280 62 1e-10
TC13572 62 1e-10
BP056597 60 5e-10
AV412895 58 2e-09
AU089037 52 1e-07
TC20217 49 1e-06
AV427022 46 6e-06
AV411435 45 1e-05
BP028574 44 4e-05
AU089120 40 6e-04
CN825238 39 7e-04
TC13043 similar to GB|BAB21544.1|12641847|AB022782 hydroxyprolin... 39 7e-04
BE122611 33 0.001
AV414139 38 0.002
TC16776 37 0.004
AV767121 37 0.004
BP078534 33 0.069
>BP041113
Length = 503
Score = 76.3 bits (186), Expect = 5e-15
Identities = 55/184 (29%), Positives = 80/184 (42%), Gaps = 15/184 (8%)
Frame = +1
Query: 20 FWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLRLSRGGEGWSWRRWLW-------- 71
FW W G LR+++ RLF+L+ +K + GW+ W W
Sbjct: 1 FWTEDWIGMGTLRDRYYRLFNLSKEKRCCILEC---------GGWNHGVWTWKLSWRRAL 153
Query: 72 -----AWEEIVLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAYEVLTATGNSITD 126
W E+++++ L V L + D W+WLP G Y+V AY L A + TD
Sbjct: 154 VGRELGWLELMMKD----LSGVCLSEGVEDKWLWLPG--GTYTVNSAYSFLQAPTLADTD 315
Query: 127 SAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTAL--CVTGCGSIETSDH 184
IW P V FAWR + RL T NL R ++ +D A+ C G +E+ H
Sbjct: 316 PIFATIWRTVAP-SVKAFAWRCLLGRLPTYDNLIKRQVV-VDPAMTVCKFCQGEVESVTH 489
Query: 185 LFLS 188
L +
Sbjct: 490 LLFA 501
>AV428268
Length = 429
Score = 66.6 bits (161), Expect = 4e-12
Identities = 41/145 (28%), Positives = 68/145 (46%)
Frame = +3
Query: 108 YSVRGAYEVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQL 167
YSV+ AY+ L N+ D+ L+W +VP +W+++ +R+ T+ NL RG+ +
Sbjct: 6 YSVKSAYDFLHGCANAQWDNIFKLLWSVKVPSNAIALSWKVLINRVQTKVNLNRRGV--V 179
Query: 168 DTALCVTGCGSIETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGF 227
+ +C E++DHL SCP +W ++ G V N HF+Q G
Sbjct: 180 ISNVCPLCSLDEESTDHLLFSCPIV*RIWSKISE*FG-VFSVFPNDSHGHFLQHLGSCGS 356
Query: 228 AKARRSFLQLIWLLTTWVIWSERNN 252
R+ +W+ IW RN+
Sbjct: 357 LNFRKRG-WFVWIAAVVCIWQGRNS 428
>TC15408 similar to UP|Q9LIC2 (Q9LIC2) Multispanning membrane protein-like,
partial (10%)
Length = 879
Score = 57.0 bits (136), Expect(2) = 3e-11
Identities = 34/93 (36%), Positives = 46/93 (48%), Gaps = 5/93 (5%)
Frame = -1
Query: 46 YTTVANMLPLRLSRGGEG-WSW----RRWLWAWEEIVLEECRTLLLDVSLFTDISDSWIW 100
Y N L L L + EG W+W RR E +++ + LL V L D +DSW W
Sbjct: 825 YLLQKNELILNLGKRVEGVWTWSFIRRRNPSTRENLLVADLCNLLESVPLSEDTNDSWSW 646
Query: 101 LPNPIGGYSVRGAYEVLTATGNSITDSAVDLIW 133
NP G Y+V+ AY+ L A+ N TD +W
Sbjct: 645 TANPEGQYTVQTAYKFLRASSNEFTDPIFHFVW 547
Score = 26.6 bits (57), Expect(2) = 3e-11
Identities = 9/19 (47%), Positives = 13/19 (68%)
Frame = -3
Query: 25 WCGGAPLREQFPRLFDLAV 43
WCGG PL F +L+ L++
Sbjct: 874 WCGGTPLNISFCKLYHLSL 818
>AV418280
Length = 398
Score = 61.6 bits (148), Expect = 1e-10
Identities = 31/91 (34%), Positives = 52/91 (57%)
Frame = -2
Query: 180 ETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIW 239
E S HLF +C F +W+ L W+G V + ++ ++N F QF+ ++ K +R IW
Sbjct: 364 ECSGHLFFTCVFSMGVWQALHRWLGISVALPASTLAN-FAQFS-ITARNKNQRLGELAIW 191
Query: 240 LLTTWVIWSERNNRLFKNVVTEAPRLLDKIK 270
+ T W +W +RN+ +F+N + LLD I+
Sbjct: 190 IATVWSLWIQRNSIIFRNNALDHSYLLDLIQ 98
>TC13572
Length = 534
Score = 61.6 bits (148), Expect = 1e-10
Identities = 36/114 (31%), Positives = 56/114 (48%), Gaps = 2/114 (1%)
Frame = -3
Query: 3 WFGDSVRRKVGDGSDTLFWFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLRLSRGGE 62
WF + V + + G+ T FW W G LRE++ RL++L+ K+ ++ S+G
Sbjct: 379 WFEEGVHKVLRSGNHTRFWLENWTGVGILREKYYRLYNLSKLKWASIDEC--GGWSQGVW 206
Query: 63 GWS--WRRWLWAWEEIVLEECRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAY 114
W WRR L E L+ ++ V L + D W+W P+ G YSV A+
Sbjct: 205 NWDFRWRRPLAGRELDWLQALSLDIVRVPLLAGVPDKWVWKPSEDGSYSVNSAF 44
>BP056597
Length = 414
Score = 59.7 bits (143), Expect = 5e-10
Identities = 29/110 (26%), Positives = 51/110 (46%), Gaps = 9/110 (8%)
Frame = +3
Query: 64 WSWRRWLW--AWEEIVLEE-------CRTLLLDVSLFTDISDSWIWLPNPIGGYSVRGAY 114
W +W+W +W+ + E + ++ SL D+W+W+ G YSV+ AY
Sbjct: 69 WLENKWVWDFSWKNPISGEDVLKLGVMKQIVSSFSLINGKKDTWVWILEGDGQYSVKSAY 248
Query: 115 EVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGI 164
++L+ +I S +W P WR+ DR+ T+ NL+ R +
Sbjct: 249 DLLSGLDTTIGISVFSKLWKACAPSNAVALGWRVFLDRIQTKDNLSRRHV 398
>AV412895
Length = 405
Score = 57.8 bits (138), Expect = 2e-09
Identities = 43/138 (31%), Positives = 65/138 (46%), Gaps = 3/138 (2%)
Frame = +1
Query: 106 GGYSVRGAYEVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGIL 165
GG+SV ++ L D + IW PL + F WR++ R+ TR NL R ++
Sbjct: 4 GGFSVNSSFVFL*DQYLEEPDPVFN*IWMVPAPLNIKAFVWRVLLGRIQTRDNLLKRQVI 183
Query: 166 Q--LDTALCVTGCGSIETS-DHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFT 222
LD A+C CG E S HL SC +W + W+G S+ H +QF+
Sbjct: 184 HNALD-AICPL-CGLAEESGSHLLFSCAESMLIWYECFAWLGVSTAQVSD-PKVHLLQFS 354
Query: 223 CLSGFAKARRSFLQLIWL 240
+ G +KA++ IW+
Sbjct: 355 SI-GRSKAQKLGETTIWM 405
>AU089037
Length = 245
Score = 51.6 bits (122), Expect = 1e-07
Identities = 29/78 (37%), Positives = 38/78 (48%), Gaps = 1/78 (1%)
Frame = -1
Query: 114 YEVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQL-DTALC 172
Y VL + + +W P FAWRL+ DR+ TR NL R ++Q + ALC
Sbjct: 245 YAVLLGSIQVPSSDIFQKLWKVPAPSNAVSFAWRLILDRVQTRGNLRRRQVIQQSEEALC 66
Query: 173 VTGCGSIETSDHLFLSCP 190
E+S HLF SCP
Sbjct: 65 PMCSQCEESSSHLFFSCP 12
>TC20217
Length = 561
Score = 48.5 bits (114), Expect = 1e-06
Identities = 49/202 (24%), Positives = 80/202 (39%), Gaps = 10/202 (4%)
Frame = +1
Query: 76 IVLEECRTLLLDVSLFTDISDSWIWLPNPIG---------GYSVRGAYEVLTATGNSITD 126
I+L+E +++ L+ D W WLP+ G +S R Y + +
Sbjct: 1 IMLDEAKSVPLE-------PDKWCWLPSTDGILQLTLFILFFSHRCCYLQIP-----FSS 144
Query: 127 SAVDLIWHHQVPLKVSIFAWRLMRDRLSTRSNLASRGILQLDTALCVT-GCGSIETSDHL 185
S +W Q+ F R+M D + NL + + V GCG + L
Sbjct: 145 SEYGSLWQLQM*RP---FL*RVMLDWIPWLQNLWKHCVS*RNNL*GVAFGCGIL*A---L 306
Query: 186 FLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAKARRSFLQLIWLLTTWV 245
SCP ++W W+G C + N H + F G K + IWL W
Sbjct: 307 LFSCPVSLDIWRHCYRWMGVCTTLPRNP-RQHLL*FQF--GGNKKHQRGADAIWLAVIWT 477
Query: 246 IWSERNNRLFKNVVTEAPRLLD 267
+W RN +F+N + + P +++
Sbjct: 478 LWLIRNEIIFRNGMLDIPAVME 543
>AV427022
Length = 429
Score = 46.2 bits (108), Expect = 6e-06
Identities = 28/102 (27%), Positives = 50/102 (48%)
Frame = +1
Query: 170 ALCVTGCGSIETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHFVQFTCLSGFAK 229
A+C C + E+ HL SC + + W+G S+ H +QF+ + G++K
Sbjct: 34 AICPLCCLAEESGSHLLFSCSNSMLI*YECHAWLGVSTAQVSDP-KVHLLQFSSI-GWSK 207
Query: 230 ARRSFLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIKL 271
A++ IW+ W IW RN +F + +LL++I++
Sbjct: 208 AQKLGESAIWMSVLWSIWCLRNMVVFNGGELDKDQLLEQIQV 333
>AV411435
Length = 426
Score = 45.1 bits (105), Expect = 1e-05
Identities = 32/115 (27%), Positives = 47/115 (40%), Gaps = 10/115 (8%)
Frame = +3
Query: 92 TDISDSWIWLPNPIGGYSVRGAYEVLTATGNSITDSAVD----------LIWHHQVPLKV 141
T D W P G YSV+ Y + AT ++++ A IW QVP KV
Sbjct: 27 TGSEDRLFWPFIPSGEYSVKSGYRAVKATQFNLSNLASSSSHPSSFLWHTIWGAQVPKKV 206
Query: 142 SIFAWRLMRDRLSTRSNLASRGILQLDTALCVTGCGSIETSDHLFLSCPFFAELW 196
F WR + + + NL R + + D C E +H L+C + +W
Sbjct: 207 RSFLWRAANNAIPVKRNLKRRNMGRDD--FCPICNKGPEDINHALLACEWTRAVW 365
>BP028574
Length = 454
Score = 43.5 bits (101), Expect = 4e-05
Identities = 19/64 (29%), Positives = 32/64 (49%)
Frame = +3
Query: 96 DSWIWLPNPIGGYSVRGAYEVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRLST 155
DSW+W+ + G ++V+ AYE L I D+ +W + P AWR+ +R+
Sbjct: 261 DSWVWIKDGSGSFTVKSAYEELHGEIQEIEDNFFRKLWQSKAPSNSIALAWRVGLNRVPC 440
Query: 156 RSNL 159
+ L
Sbjct: 441 MAEL 452
>AU089120
Length = 586
Score = 39.7 bits (91), Expect = 6e-04
Identities = 32/98 (32%), Positives = 44/98 (44%), Gaps = 1/98 (1%)
Frame = +2
Query: 159 LASRGILQLDTALCVTGCGSIETSDHLFLSCPFFAELWEQLRLWIGCCVGVDSNIISNHF 218
L R I+Q ++ C+ S E+ HLF C LW + W+ V V HF
Sbjct: 8 LKRRRIIQTLSSFCLH---SEESIPHLFFRCFQSWNLWSSVHRWLVFTV-VTPEEGKMHF 175
Query: 219 VQF-TCLSGFAKARRSFLQLIWLLTTWVIWSERNNRLF 255
+ F SG + ++ L IWL T W IW RN +F
Sbjct: 176 IMF*DYYSG--RDIKNGLGFIWLSTIWFIWHLRNKVVF 283
>CN825238
Length = 692
Score = 39.3 bits (90), Expect = 7e-04
Identities = 25/73 (34%), Positives = 34/73 (46%), Gaps = 6/73 (8%)
Frame = +3
Query: 98 WIWLPNPIGGYSVRGAYEVLTATGNSITDSAV------DLIWHHQVPLKVSIFAWRLMRD 151
W W N G +SVR AY + T S+ +W VP K+ FA+R+ R+
Sbjct: 420 WRWSRN--GAFSVRSAYYNIQVAKEMQTSSSAASSFPWKKLWRTPVPNKMKHFAYRVARN 593
Query: 152 RLSTRSNLASRGI 164
+ R NL RGI
Sbjct: 594 IIPCRENLQRRGI 632
>TC13043 similar to GB|BAB21544.1|12641847|AB022782 hydroxyproline-rich
glycoprotein {Arabidopsis thaliana;} , partial (6%)
Length = 678
Score = 39.3 bits (90), Expect = 7e-04
Identities = 17/53 (32%), Positives = 29/53 (54%)
Frame = +3
Query: 213 IISNHFVQFTCLSGFAKARRSFLQLIWLLTTWVIWSERNNRLFKNVVTEAPRL 265
+I+ +QF L G + + +++WL W +W+ RNN +F+ V E RL
Sbjct: 519 VIAMFMLQFE-LMGLNRQQNKLWKVVWLAVIWTVWNGRNNLVFREVEMEIERL 674
>BE122611
Length = 357
Score = 33.1 bits (74), Expect(2) = 0.001
Identities = 22/79 (27%), Positives = 36/79 (44%), Gaps = 2/79 (2%)
Frame = -3
Query: 95 SDSWIWLPNPIGGYSVRGAYEVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLMRDRL- 153
+D +W G YS + Y +L G + L+W P KV F W + R+ L
Sbjct: 334 TDRILWTEESNGCYSAKSTYNLLIEDGGN-ESRFWSLVWKANAPEKVRFFLWLVGRNALP 158
Query: 154 -STRSNLASRGILQLDTAL 171
+ R + + +LQL +A+
Sbjct: 157 SNARRHHITLLVLQLVSAV 101
Score = 24.3 bits (51), Expect(2) = 0.001
Identities = 10/30 (33%), Positives = 16/30 (53%)
Frame = -2
Query: 169 TALCVTGCGSIETSDHLFLSCPFFAELWEQ 198
+A C+ E ++H+F CP A +W Q
Sbjct: 122 SAACIRCGAPSEDANHVFRWCPGSAWMWTQ 33
>AV414139
Length = 309
Score = 38.1 bits (87), Expect = 0.002
Identities = 28/101 (27%), Positives = 38/101 (36%), Gaps = 11/101 (10%)
Frame = -2
Query: 60 GGEGWSWRRWLWAWE-----EIVLEECRT------LLLDVSLFTDISDSWIWLPNPIGGY 108
GG G W +W WE E+ + E R L V L D W W + Y
Sbjct: 305 GGMG-QWHNGVWEWEFRWRRELSVREQRQEADLIQALAGVCLQQGTEDRWKWTLDTEDIY 129
Query: 109 SVRGAYEVLTATGNSITDSAVDLIWHHQVPLKVSIFAWRLM 149
+VR AY +L + +W P + F WRL+
Sbjct: 128 TVRSAY*LLLDPIADSDSEVYEKVWVGIAPSNANAFVWRLL 6
>TC16776
Length = 606
Score = 37.0 bits (84), Expect = 0.004
Identities = 18/55 (32%), Positives = 29/55 (52%)
Frame = +1
Query: 216 NHFVQFTCLSGFAKARRSFLQLIWLLTTWVIWSERNNRLFKNVVTEAPRLLDKIK 270
+HF+Q L +K L LIW+ T +IW+ RN F+ V E + +D ++
Sbjct: 61 SHFLQHESLLPGSKKVAQTLTLIWVATVGLIWNSRNGAEFREEVVEICKSVDTVQ 225
>AV767121
Length = 525
Score = 37.0 bits (84), Expect = 0.004
Identities = 14/37 (37%), Positives = 20/37 (53%)
Frame = +2
Query: 3 WFGDSVRRKVGDGSDTLFWFHWWCGGAPLREQFPRLF 39
WF +++ VG+G T FW W G L +F RL+
Sbjct: 413 WFSSGLKKLVGEGDQTKFWSEDWLGSGLLSARFRRLY 523
>BP078534
Length = 410
Score = 32.7 bits (73), Expect = 0.069
Identities = 21/59 (35%), Positives = 26/59 (43%), Gaps = 4/59 (6%)
Frame = -3
Query: 21 WFHWWCGGAPLREQFPRLFDLAVDKYTTVANMLPLRLSRGGEGWSW----RRWLWAWEE 75
W W L + FPRLF L +K VA++ S G W W RR + WEE
Sbjct: 216 WLDPWIRERALCDMFPRLFTLCENKGALVADV----GSYHGGSWHWNLSFRREFFQWEE 52
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.326 0.140 0.483
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,836,432
Number of Sequences: 28460
Number of extensions: 119812
Number of successful extensions: 1503
Number of sequences better than 10.0: 93
Number of HSP's better than 10.0 without gapping: 1415
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1481
length of query: 275
length of database: 4,897,600
effective HSP length: 89
effective length of query: 186
effective length of database: 2,364,660
effective search space: 439826760
effective search space used: 439826760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC130801.11


