

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.9 + phase: 0
(487 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
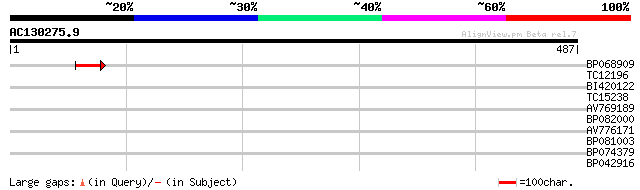
Score E
Sequences producing significant alignments: (bits) Value
BP068909 42 3e-04
TC12196 weakly similar to UP|Q39061 (Q39061) RNA-binding protein... 29 2.0
BI420122 28 2.6
TC15238 28 2.6
AV769189 28 4.4
BP082000 27 5.7
AV776171 27 9.8
BP081003 27 9.8
BP074379 27 9.8
BP042916 27 9.8
>BP068909
Length = 279
Score = 41.6 bits (96), Expect = 3e-04
Identities = 14/26 (53%), Positives = 22/26 (83%)
Frame = +3
Query: 57 EPKAYIPSNISIGPYHYGSQHLQEME 82
+ KAY+P +S+GPYH+G +HL++ME
Sbjct: 117 DDKAYVPQIVSLGPYHHGKEHLRQME 194
>TC12196 weakly similar to UP|Q39061 (Q39061) RNA-binding protein cp33
(AT3g52380/F22O6_240), partial (19%)
Length = 552
Score = 28.9 bits (63), Expect = 2.0
Identities = 11/26 (42%), Positives = 18/26 (68%)
Frame = +3
Query: 50 PPNMLNVEPKAYIPSNISIGPYHYGS 75
PP LN++PK + PS++S+ +H S
Sbjct: 102 PPKPLNLKPKLFSPSSLSLYRFHLPS 179
>BI420122
Length = 590
Score = 28.5 bits (62), Expect = 2.6
Identities = 17/44 (38%), Positives = 24/44 (53%), Gaps = 5/44 (11%)
Frame = -2
Query: 169 IRREMIMLENQ-LPMFVLTKLFELT----DKNNSLHPQMSFYNL 207
+R ++ +Q L FVL L +L D+NNS HP+M Y L
Sbjct: 478 VRTSSQLIHHQCLAFFVLIHLLQLGS*IFDRNNSKHPEMQLYPL 347
>TC15238
Length = 665
Score = 28.5 bits (62), Expect = 2.6
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = -3
Query: 243 YNIRPKLIGEEPRGCQSQMIH 263
YN+RPK+ + P+ Q QM+H
Sbjct: 597 YNLRPKMNSDNPKNKQRQMMH 535
>AV769189
Length = 410
Score = 27.7 bits (60), Expect = 4.4
Identities = 13/41 (31%), Positives = 19/41 (45%)
Frame = +3
Query: 274 KIKACESRELMDISFGKKWGIMVKELTIPPLYIGDHRGTVF 314
K+ AC SR G+ G K L++ +Y H G +F
Sbjct: 234 KVSACSSRTSTSPIVGRSSGSCFKHLSMISMYTA*HSGAIF 356
>BP082000
Length = 362
Score = 27.3 bits (59), Expect = 5.7
Identities = 13/41 (31%), Positives = 24/41 (57%)
Frame = +3
Query: 7 KLIKRKFHGTAVKSSKFWRGLVEKELQSIKPEEKNESVCIY 47
KL+ ++HG + S F+ LVE ++++ + N S CI+
Sbjct: 162 KLL*GQYHGKSWPKSHFYMHLVEYQVRASPKKSFNRSHCIH 284
>AV776171
Length = 184
Score = 26.6 bits (57), Expect = 9.8
Identities = 9/27 (33%), Positives = 16/27 (58%)
Frame = -1
Query: 413 YLTSWAVGLSTLGALLALYFTFIQAIC 439
+L W++ L L +L Y+ F+ A+C
Sbjct: 163 WLIHWSIYLYLLSCILTSYYVFVVALC 83
>BP081003
Length = 205
Score = 26.6 bits (57), Expect = 9.8
Identities = 18/39 (46%), Positives = 22/39 (56%)
Frame = -1
Query: 387 LYNVVNEANEYLGCWRARFRASLVHNYLTSWAVGLSTLG 425
LY VV A E L CWRA + S+ N+L WA+ LG
Sbjct: 118 LYFVVL-AMENLLCWRALYSFSVG*NHLLFWALHGLCLG 5
>BP074379
Length = 465
Score = 26.6 bits (57), Expect = 9.8
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = +1
Query: 10 KRKFHGTAVKSSKFWRG 26
K +FHGT VK ++ W G
Sbjct: 127 KMQFHGTQVKENQLWEG 177
>BP042916
Length = 490
Score = 26.6 bits (57), Expect = 9.8
Identities = 16/42 (38%), Positives = 21/42 (49%), Gaps = 1/42 (2%)
Frame = +3
Query: 380 PDMNESYLYNVVNEANEYLG-CWRARFRASLVHNYLTSWAVG 420
P++ + YLYN E+ EYL WR R + TS A G
Sbjct: 66 PEILKKYLYNKTVESLEYLSPWWRLSMRNPALIKLNTSSAAG 191
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,786,915
Number of Sequences: 28460
Number of extensions: 119615
Number of successful extensions: 526
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 524
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 526
length of query: 487
length of database: 4,897,600
effective HSP length: 94
effective length of query: 393
effective length of database: 2,222,360
effective search space: 873387480
effective search space used: 873387480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC130275.9


