

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.7 - phase: 0 /pseudo
(833 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
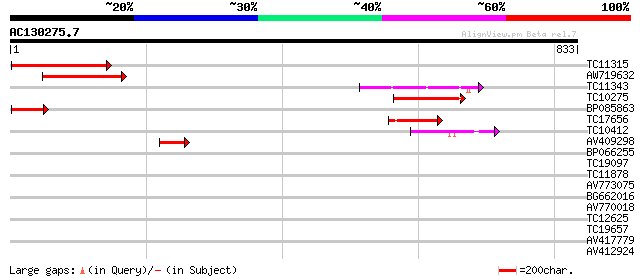
Sequences producing significant alignments: (bits) Value
TC11315 similar to UP|AHM2_ARATH (O64474) Potential cadmium/zinc... 152 2e-37
AW719632 120 1e-27
TC11343 similar to PIR|B96660|B96660 protein F2K11.18 [imported]... 99 4e-21
TC10275 similar to UP|Q7Y051 (Q7Y051) Paa2 P-type ATPase, partia... 75 6e-14
BP085863 74 1e-13
TC17656 similar to GB|BAA97361.1|8843813|AB023042 Ca2+-transport... 59 2e-09
TC10412 homologue to UP|Q9FVE8 (Q9FVE8) Plasma membrane Ca2+-ATP... 54 8e-08
AV409298 47 9e-06
BP066255 34 0.083
TC19097 30 2.0
TC11878 similar to UP|O49605 (O49605) (Cyclophilin-like protein... 29 2.7
AV773075 29 3.5
BG662016 28 6.0
AV770018 28 6.0
TC12625 similar to UP|Q84TG1 (Q84TG1) At3g09720, partial (34%) 28 6.0
TC19657 28 7.8
AV417779 28 7.8
AV412924 28 7.8
>TC11315 similar to UP|AHM2_ARATH (O64474) Potential
cadmium/zinc-transporting ATPase 2 , partial (12%)
Length = 570
Score = 152 bits (385), Expect = 2e-37
Identities = 75/147 (51%), Positives = 109/147 (74%)
Frame = +2
Query: 3 ENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIAD 62
+ +++S F+VLG+CC++E L+E ILKPL G+ VSVIVP+RT+ VVHD L+IS+ +I
Sbjct: 128 KKLQKSYFDVLGLCCSSEVPLIESILKPLEGINEVSVIVPSRTLIVVHDSLIISQLQIVK 307
Query: 63 ALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFP 122
LN ARLEA+ R G ++K+ P ++A GLLL LSFLKY+Y PL +LALG+VA+G
Sbjct: 308 VLNQARLEANVRVYGNEKHQKRWPRPYSIASGLLLLLSFLKYVYSPLKFLALGAVAVGIY 487
Query: 123 KVLFRAIASIRALTLNINILVLLAVCG 149
++ ++ AS+R L+INIL+++AV G
Sbjct: 488 PIILKSFASLRNAILDINILMIIAVIG 568
>AW719632
Length = 401
Score = 120 bits (300), Expect = 1e-27
Identities = 61/123 (49%), Positives = 87/123 (70%)
Frame = +3
Query: 49 VHDVLLISESKIADALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPP 108
V D L+IS+ +I ALN ARLEA+ R G ++K+ P ++A GLLL LSFLKY+Y P
Sbjct: 33 VGDGLIISQLQIVKALNQARLEANARVYGNEKHQKRWPSPYSIASGLLLLLSFLKYVYSP 212
Query: 109 LGWLALGSVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSI 168
L +LALG+VA G ++ ++ AS+R L+INIL+++AV GT A+ D+ + I+FLFSI
Sbjct: 213 LKFLALGAVAAGIFPIILKSFASLRNARLDINILMIIAVIGTIAMNDYLEAXTIVFLFSI 392
Query: 169 AQW 171
A+W
Sbjct: 393 AEW 401
>TC11343 similar to PIR|B96660|B96660 protein F2K11.18 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(21%)
Length = 909
Score = 98.6 bits (244), Expect = 4e-21
Identities = 62/192 (32%), Positives = 112/192 (58%), Gaps = 10/192 (5%)
Frame = +2
Query: 514 LVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLP 573
++GV ++ D + A E + L + +RS+M+TGD+ A + Q I+ V AE P
Sbjct: 161 VIGVLAVSDPLKPNAREVVSILNSMNIRSIMVTGDNWGTANSIARQA--GIETVIAEAQP 334
Query: 574 HEKAKLIENFKKEG-PIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDI 632
KA ++ + G +AM+GDGIND+PAL AD+GI++G +G+ +A E S+ +LM +++
Sbjct: 335 QTKATKVKELQTSGYTVAMVGDGINDSPALVAADVGIAIG-AGTDIAIEASDIVLMRSNL 511
Query: 633 RKVPEAIRLARKTTRKLVENVIISVGFKCAILALAIAG---YPLV------WLAVLTDVG 683
+ A+ LA+KT ++ N + ++G+ +LA+ IA YP + W+A +
Sbjct: 512 EDIIIALDLAKKTFSRIRLNYLWALGYN--LLAIPIAAGILYPFIRFRLHPWIAGVAMAA 685
Query: 684 TCLLVILNSMLI 695
+ + V+ +S+L+
Sbjct: 686 SSVRVVCSSLLL 721
>TC10275 similar to UP|Q7Y051 (Q7Y051) Paa2 P-type ATPase, partial (16%)
Length = 724
Score = 74.7 bits (182), Expect = 6e-14
Identities = 43/107 (40%), Positives = 69/107 (64%), Gaps = 2/107 (1%)
Frame = +1
Query: 565 DIVHAELLPHEKAKLIENFKKEGP-IAMIGDGINDAPALATADIGISM-GISGSALANET 622
D V A L P +K++ I + K G +AM+GDGINDAPALA AD+GI++ + A++
Sbjct: 1 DFVKASLAPQQKSEFISSLKAAGHHVAMVGDGINDAPALAVADVGIALQNEAQENAASDA 180
Query: 623 SNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFKCAILALAIA 669
++ IL+ N I +V +AI LA+ T K+ +N+ +V + ++A+ IA
Sbjct: 181 ASIILLGNKISQVVDAIDLAQSTMAKVYQNLSWAVAYN--VIAIPIA 315
>BP085863
Length = 246
Score = 73.6 bits (179), Expect = 1e-13
Identities = 33/55 (60%), Positives = 46/55 (83%)
Frame = +1
Query: 3 ENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISE 57
+ +++S F+VLG+CC++E L+E ILKPL G+K VSVIVP+RTV VVHD L+IS+
Sbjct: 76 KKLQKSYFDVLGLCCSSEVPLIEGILKPLEGIKEVSVIVPSRTVIVVHDGLIISQ 240
>TC17656 similar to GB|BAA97361.1|8843813|AB023042 Ca2+-transporting
ATPase-like protein {Arabidopsis thaliana;} , partial
(25%)
Length = 821
Score = 59.3 bits (142), Expect = 2e-09
Identities = 29/80 (36%), Positives = 55/80 (68%), Gaps = 1/80 (1%)
Frame = +3
Query: 557 QSQLKNAIDIVHAELLPHEKAKLIENFKKEGPI-AMIGDGINDAPALATADIGISMGISG 615
+ ++ +AI ++ P++K L++ +++G + A+ GDG NDAPAL ADIG++MGI+G
Sbjct: 33 RDEIADAISVM-GRSSPNDKLLLVQALRRKGHVVAVTGDGTNDAPALHEADIGLAMGIAG 209
Query: 616 SALANETSNAILMSNDIRKV 635
+ +A E+S+ I++ ++ V
Sbjct: 210 TEVAKESSDIIILDDNFASV 269
>TC10412 homologue to UP|Q9FVE8 (Q9FVE8) Plasma membrane Ca2+-ATPase,
partial (33%)
Length = 1061
Score = 54.3 bits (129), Expect = 8e-08
Identities = 38/142 (26%), Positives = 72/142 (49%), Gaps = 11/142 (7%)
Frame = +3
Query: 589 IAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARK---T 645
+A+ GDG NDAPAL ADIG++MGI+G+ +A E+++ I++ ++ + + R
Sbjct: 657 VAVTGDGTNDAPALHEADIGLAMGIAGTEVAKESADVIILDDNFSTIVTVAKWGRSVYIN 836
Query: 646 TRKLVE-----NVI-ISVGFKCAIL--ALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQ 697
+K V+ NV+ + V F A L + L+W+ ++ D L ++ +
Sbjct: 837 IQKFVQFQLTVNVVALIVNFTSACLTGTAPLTAVQLLWVNMIMDT-------LGALALAT 995
Query: 698 ENHKYEKRESTKGSKYGKFLED 719
E K + + + + G F+ +
Sbjct: 996 EPPKDDLMKRSPVGRKGNFISN 1061
>AV409298
Length = 332
Score = 47.4 bits (111), Expect = 9e-06
Identities = 24/44 (54%), Positives = 28/44 (63%)
Frame = +3
Query: 221 DAIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMN 264
D IP DGIV G+ VDE TGE PVTK + V AG+IN+N
Sbjct: 3 DRIPADGIVRAGRSTVDESSFTGEPLPVTKVAGCEVAAGSINLN 134
>BP066255
Length = 540
Score = 34.3 bits (77), Expect = 0.083
Identities = 19/73 (26%), Positives = 37/73 (50%)
Frame = +2
Query: 335 DMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAF 394
+M +F +A+ +++ P L L+ +++ A+ K L++ ET+ T+
Sbjct: 272 EMLEYFAIAVTIVVVAVPEGLPLAVTLSLAFAMKKMMNDKALVRHLAACETMGSATTICS 451
Query: 395 DKTGTITRGEFSV 407
DKTGT+T +V
Sbjct: 452 DKTGTLTTNHMTV 490
>TC19097
Length = 405
Score = 29.6 bits (65), Expect = 2.0
Identities = 14/29 (48%), Positives = 18/29 (61%), Gaps = 2/29 (6%)
Frame = +2
Query: 339 WF--HLALVVLLSGCPCALILSTPVAIFC 365
WF LAL VL CP AL+L+ P ++ C
Sbjct: 257 WF*IFLALSVLCDSCPDALVLNFPCSLLC 343
>TC11878 similar to UP|O49605 (O49605) (Cyclophilin-like protein)
(Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase) , partial (47%)
Length = 504
Score = 29.3 bits (64), Expect = 2.7
Identities = 17/59 (28%), Positives = 32/59 (53%), Gaps = 4/59 (6%)
Frame = -2
Query: 455 NVENFQNFPGEGIFGTIDGRDIYIGNKRV---GVRA-ICKRDNCEVQFQRPEISTKKNN 509
N+ N + +F +I ++ GN+++ G+R+ IC RDN ++ PEI +K +
Sbjct: 215 NIHALNNLAKDNMF-SIQPTGLHCGNEKL*AMGIRS*ICHRDNTSMRMFNPEILIRKGS 42
>AV773075
Length = 472
Score = 28.9 bits (63), Expect = 3.5
Identities = 18/54 (33%), Positives = 28/54 (51%), Gaps = 11/54 (20%)
Frame = -2
Query: 224 PLDGIVVEGKCEVDEKM---LTGESFPVTKESDS--------LVWAGTINMNAY 266
P +GIVV+ C V+ K+ L+ + P++ SD L+W GT +N Y
Sbjct: 408 P*NGIVVKIFCWVEPKVKPHLSSYNLPISTSSDVHICLQHIWLIWCGTKELNVY 247
>BG662016
Length = 459
Score = 28.1 bits (61), Expect = 6.0
Identities = 14/34 (41%), Positives = 23/34 (67%)
Frame = +1
Query: 428 IESKSSHPVAGALVDYARLHSIKPVPENVENFQN 461
+ S +SH A ++D ARL+ ++ +PEN+EN N
Sbjct: 244 LPSSTSHLTALTVLD-ARLNCLRALPENLENLIN 342
>AV770018
Length = 599
Score = 28.1 bits (61), Expect = 6.0
Identities = 19/80 (23%), Positives = 31/80 (38%)
Frame = +2
Query: 3 ENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIAD 62
+ I + GM C + + +E L+ + GVK V + V D + KI +
Sbjct: 350 QEISVCQVRIKGMACTSCSESIENALQMVDGVKRAIVGLALEEAKVHFDPSVTGADKIIE 529
Query: 63 ALNTARLEASFRPQGETNNE 82
A+ A A G N+
Sbjct: 530 AVEDAGFGAELISSGNDMNK 589
Score = 27.7 bits (60), Expect = 7.8
Identities = 16/60 (26%), Positives = 29/60 (47%)
Frame = +2
Query: 5 IKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADAL 64
IK F + + CA+ +E LK L G+K V+V + + LI+ +I +++
Sbjct: 128 IKTVTFRITDIKCASCVNSIESALKTLAGIKTVAVSPLDGRAAIKFEPNLITVKRIKESI 307
>TC12625 similar to UP|Q84TG1 (Q84TG1) At3g09720, partial (34%)
Length = 549
Score = 28.1 bits (61), Expect = 6.0
Identities = 16/64 (25%), Positives = 31/64 (48%), Gaps = 12/64 (18%)
Frame = +2
Query: 533 EELKLLGVRSVMLTGDSSQVAMYVQSQLKNA------------IDIVHAELLPHEKAKLI 580
EE KLL +R + V +++QS+ + +D++H++L E+ I
Sbjct: 263 EEGKLLAIRQSFAESLNPPVLVFLQSKERAKELYGELAFDNIRVDVIHSDLSQEERENAI 442
Query: 581 ENFK 584
+NF+
Sbjct: 443 DNFR 454
>TC19657
Length = 632
Score = 27.7 bits (60), Expect = 7.8
Identities = 15/42 (35%), Positives = 23/42 (54%)
Frame = +3
Query: 54 LISESKIADALNTARLEASFRPQGETNNEKKCPDILTMACGL 95
L+S +K +A+NTA+ +AS + + EKK T GL
Sbjct: 267 LVSCAKAVEAINTAKSDASLDSTSQDSVEKKALQSATFPNGL 392
>AV417779
Length = 401
Score = 27.7 bits (60), Expect = 7.8
Identities = 20/49 (40%), Positives = 27/49 (54%)
Frame = -3
Query: 112 LALGSVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGG 160
LALG + P+VL RA+A + +L L L+ A+ T AL D GG
Sbjct: 282 LALGLQFVQHPRVLERALAHLSSLLLE---LLNRALLDTPALVDQMAGG 145
>AV412924
Length = 401
Score = 27.7 bits (60), Expect = 7.8
Identities = 16/44 (36%), Positives = 22/44 (49%)
Frame = -2
Query: 204 VDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGESFP 247
VDV D + A DA+P+D V+ G D ++ GE FP
Sbjct: 193 VDVGDATGGGV----AADAVPVDAAVIAGP*VEDAEVRFGEFFP 74
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.137 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,097,070
Number of Sequences: 28460
Number of extensions: 164414
Number of successful extensions: 805
Number of sequences better than 10.0: 37
Number of HSP's better than 10.0 without gapping: 799
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 804
length of query: 833
length of database: 4,897,600
effective HSP length: 98
effective length of query: 735
effective length of database: 2,108,520
effective search space: 1549762200
effective search space used: 1549762200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC130275.7


