

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.4 - phase: 0
(362 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
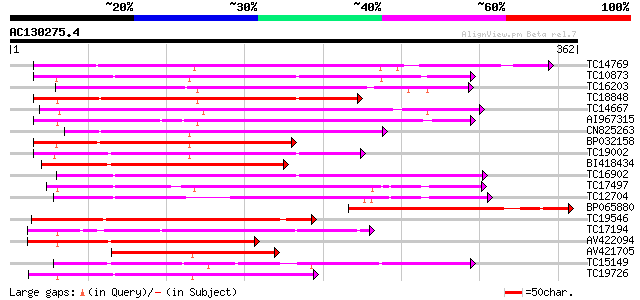
Score E
Sequences producing significant alignments: (bits) Value
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 180 3e-46
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 171 2e-43
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 165 1e-41
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 163 5e-41
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 161 2e-40
AI967315 159 5e-40
CN825263 156 4e-39
BP032158 156 6e-39
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 156 6e-39
BI418434 153 5e-38
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 151 1e-37
TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2,... 147 2e-36
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 144 2e-35
BP065880 142 9e-35
TC19546 similar to UP|Q40543 (Q40543) Protein-serine/threonine k... 141 1e-34
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 141 2e-34
AV422094 137 2e-33
AV421705 137 2e-33
TC15149 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like pro... 132 7e-32
TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase... 131 2e-31
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 180 bits (457), Expect = 3e-46
Identities = 123/345 (35%), Positives = 179/345 (51%), Gaps = 13/345 (3%)
Frame = +3
Query: 16 FTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTI 75
FT + + +AT+ Y L+G GGFG+VY+G ++G VAVKV R S++ + +F E+ +
Sbjct: 2133 FTLEYIEVATERYKTLIGEGGFGSVYRGTLNDGQEVAVKV-RSSTSTQGTREFDNELNLL 2309
Query: 76 GRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET---KVLGYEKLHEIAIGTA 132
I H NLV L G+C E + LVY +M NGSL L+ E K+L + IA+G A
Sbjct: 2310 SAIQHENLVPLLGYCNESDQQILVYPFMSNGSLQDRLYGEPAKRKILDWPTRLSIALGAA 2489
Query: 133 RGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPG 192
RG+AYLH +IH DIK NILLD + KVADFG +K +E RGT G
Sbjct: 2490 RGLAYLHTFPGRSVIHRDIKSSNILLDHSMCAKVADFGFSKYAPQEGDSYVSLEVRGTAG 2669
Query: 193 YAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQ----EWFPIWVWKKK-- 246
Y PE + ++ K DV+SFG++L EI+ R L IK ++ EW ++ K
Sbjct: 2670 YLDPEYYKTQQLSEKSDVFSFGVVLLEIVSGREPLNIKRPRTEWSLVEWATPYIRGSKVD 2849
Query: 247 ---DAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEIP 303
D G+ G + E R+++VAL C++ RP M +V+ LE +L I
Sbjct: 2850 EIVDPGIKG---------GYHAEAMWRVVEVALQCLEPFSTYRPSMVAIVRELEDALIIE 3002
Query: 304 KTFNPFQHLIDGTEFTTHSVQESNTYTTSVSS-VMVSDSSIVCAT 347
+ + ID S+ SN Y+ + V+ S +S +T
Sbjct: 3003 NNASEYMKSID-------SLGGSNRYSIVIEKRVLPSTTSTAEST 3116
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 171 bits (432), Expect = 2e-43
Identities = 107/292 (36%), Positives = 163/292 (55%), Gaps = 10/292 (3%)
Frame = +1
Query: 16 FTGQQLRIATDNY--SNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVG 73
F ++L AT N+ +NL+G GGFG VYKG + G VAVK L + E F+ EV
Sbjct: 415 FGFRELADATRNFKEANLIGEGGFGKVYKGRLTTGEAVAVKQLSHDGRQGFQE-FVMEVL 591
Query: 74 TIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLF---HETKVLGYEKLHEIAIG 130
+ +HH NLVRL G+C + + LVYEYM GSL+ +LF H+ + L + ++A+G
Sbjct: 592 MLSLLHHTNLVRLIGYCTDGDQRLLVYEYMPMGSLEDHLFELSHDKEPLNWSTRMKVAVG 771
Query: 131 TARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAK-NCNRENTHITMTGGRG 189
ARG+ YLH +I+ D+K NILLD F PK++DFGLAK +NTH++ T G
Sbjct: 772 AARGLEYLHCTADPPVIYRDLKSANILLDNEFNPKLSDFGLAKLGPVGDNTHVS-TRVMG 948
Query: 190 TPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQE----WFPIWVWKK 245
T GY APE M +T K D+YSFG++L E++ RR + ++ W + +
Sbjct: 949 TYGYCAPEYAMSGKLTLKSDIYSFGVVLLELLTGRRAIDTSRRPGEQNLVSWARPYFSDR 1128
Query: 246 KDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLE 297
+ G + + ++ + + I + C+Q +P+ RP+++ +V LE
Sbjct: 1129RRFGHMVDPLLQGRFPSR---CLHQAIAITAMCLQEQPKFRPLITDIVVALE 1275
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 165 bits (417), Expect = 1e-41
Identities = 104/277 (37%), Positives = 149/277 (53%), Gaps = 10/277 (3%)
Frame = +2
Query: 30 NLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYGF 89
N++G GG G VY+G NGT VA+K L G + + D F AE+ T+G+I H N++RL G+
Sbjct: 2204 NIIGKGGAGIVYRGSMPNGTDVAIKRLVGQGSGRNDYGFRAEIETLGKIRHRNIMRLLGY 2383
Query: 90 CFERNLIALVYEYMGNGSLDRYLFHETKV--LGYEKLHEIAIGTARGIAYLHEECQHRII 147
++ L+YEYM NGSL +L H K L +E ++IA+ ARG+ Y+H +C II
Sbjct: 2384 VSNKDTNLLLYEYMPNGSLGEWL-HGAKGGHLRWEMRYKIAVEAARGLCYMHHDCSPLII 2560
Query: 148 HYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELWMPFPITHK 207
H D+K NILLD +F VADFGLAK +M+ G+ GY APE + K
Sbjct: 2561 HRDVKSNNILLDADFEAHVADFGLAKFLYDPGASQSMSSIAGSYGYIAPEYAYTLKVDEK 2740
Query: 208 CDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAGLLGE----AMIVCGIEEK 263
DVYSFG++L E+I R+ + E + I W K L + A+++ ++ +
Sbjct: 2741 SDVYSFGVVLLELIIGRKPVG----EFGDGVDIVGWVNKTMSELSQPSDTALVLAVVDPR 2908
Query: 264 NK----EIAERMIKVALWCVQYRPELRPIMSVVVKML 296
M +A+ CV+ RP M VV ML
Sbjct: 2909 LSGYPLTSVIHMFNIAMMCVKEMGPARPTMREVVHML 3019
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 163 bits (412), Expect = 5e-41
Identities = 97/215 (45%), Positives = 132/215 (61%), Gaps = 5/215 (2%)
Frame = +2
Query: 16 FTGQQLRIATDNYS--NLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVG 73
FT ++L AT +S N LG GGFG+VY G S+G +AVK L+ + N K + +F EV
Sbjct: 266 FTYKELHAATGGFSDDNKLGEGGFGSVYWGRTSDGLQIAVKKLK-AMNSKAEMEFAVEVE 442
Query: 74 TIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV---LGYEKLHEIAIG 130
+GR+ H NL+ L G+C + +VY+YM N SL +L + V L ++K +IAIG
Sbjct: 443 VLGRVRHKNLLGLRGYCVGDDQRLIVYDYMPNLSLLSHLHGQFAVEVQLNWQKRMKIAIG 622
Query: 131 TARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGT 190
+A GI YLH E IIH DIK N+LL+ +F P VADFG AK +H+T T +GT
Sbjct: 623 SAEGILYLHHEVTPHIIHRDIKASNVLLNSDFEPLVADFGFAKLIPEGVSHMT-TRVKGT 799
Query: 191 PGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRR 225
GY APE M ++ CDVYSFG+LL E++ R+
Sbjct: 800 LGYLAPEYAMWGKVSESCDVYSFGILLLELVTGRK 904
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 161 bits (407), Expect = 2e-40
Identities = 103/299 (34%), Positives = 154/299 (51%), Gaps = 15/299 (5%)
Frame = +2
Query: 20 QLRIATDNYSN--LLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGR 77
+L+ TDN+ + L+G G +G VY ++G VAVK L SS + + +F+ +V + R
Sbjct: 335 ELKEKTDNFGSKALIGEGSYGRVYYATLNDGNAVAVKKLDVSSEPETNNEFLTQVSMVSR 514
Query: 78 IHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVLGYE--------KLHEIAI 129
+ + N V L+G+C E NL L YE+ GSL L V G + + IA+
Sbjct: 515 LKNDNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWIQRVRIAV 694
Query: 130 GTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRG 189
ARG+ YLHE+ Q IIH DI+ N+L+ +++ K+ADF L+ + T G
Sbjct: 695 DAARGLEYLHEKVQPAIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAARLHSTRVLG 874
Query: 190 TPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAG 249
T GY APE M +T K DVYSFG++L E++ R+ + Q+ W +
Sbjct: 875 TFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPR---- 1042
Query: 250 LLGEAMIVCGIEEKNK-----EIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEIP 303
L E + ++ K K + ++ VA CVQY E RP MS+VVK L+ L+ P
Sbjct: 1043-LSEDKVKQCVDPKLKGEYPPKGVAKLAAVAALCVQYEAEFRPNMSIVVKALQPLLKTP 1216
>AI967315
Length = 1308
Score = 159 bits (403), Expect = 5e-40
Identities = 102/288 (35%), Positives = 166/288 (57%), Gaps = 6/288 (2%)
Frame = +1
Query: 16 FTGQQLRIATDNYS--NLLGSGGFGTVYKGIFSNGTMVAVKVL-RGSSNKKIDEQFMAEV 72
F+ ++L AT+ +S N++G GG+ VYKG +G +AVK L R +++ +++F+ E+
Sbjct: 112 FSYEELFHATNGFSSENMVGKGGYAEVYKGRLESGDEIAVKRLTRTCRDERKEKEFLTEI 291
Query: 73 GTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV--LGYEKLHEIAIG 130
GTIG + H N++ L G C + L LV+E GS+ L H+ K+ L ++ ++I +G
Sbjct: 292 GTIGHVCHSNVMPLLGCCIDNGLY-LVFELSTVGSVAS-LIHDEKMAPLDWKTRYKIVLG 465
Query: 131 TARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGT 190
TARG+ YLH+ CQ RIIH DIK NILL ++F P+++DFGLAK + TH ++ GT
Sbjct: 466 TARGLHYLHKGCQRRIIHRDIKASNILLTEDFEPQISDFGLAKWLPSQWTHHSIAPIEGT 645
Query: 191 PGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAGL 250
G+ APE +M + K DV++FG+ L E+I R+ + + W + K + L
Sbjct: 646 FGHLAPEYYMHGVVDEKTDVFAFGVFLLEVISGRKPVDGSHQSLHTWAKPILSKWEIEKL 825
Query: 251 LGEAMIVC-GIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLE 297
+ + C + + N R+ A C++ RP MS V++++E
Sbjct: 826 VDPRLEGCYDVTQFN-----RVAFAASLCIRASSTWRPTMSEVLEVME 954
>CN825263
Length = 663
Score = 156 bits (395), Expect = 4e-39
Identities = 87/209 (41%), Positives = 124/209 (58%), Gaps = 3/209 (1%)
Frame = +1
Query: 36 GFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNL 95
GFG VYKGI ++G VAVK+L+ +++ +F+AEV + R+HH NLV+L G C E+
Sbjct: 1 GFGLVYKGILNDGRDVAVKILK-RDDQRGGREFLAEVEMLSRLHHRNLVKLIGICIEKQT 177
Query: 96 IALVYEYMGNGSLDRYLF---HETKVLGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIK 152
L+YE + NGS++ +L ET L + +IA+G ARG+AYLHE+ +IH D K
Sbjct: 178 RCLIYELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFK 357
Query: 153 PGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELWMPFPITHKCDVYS 212
NILL+ +F PKV+DFGLA+ E T GT GY APE M + K DVYS
Sbjct: 358 SSNILLECDFTPKVSDFGLARTALDEGNKHISTHVMGTFGYLAPEYAMTGHLLVKSDVYS 537
Query: 213 FGMLLFEIIGRRRNLAIKNTESQEWFPIW 241
+G++L E++ + + + QE W
Sbjct: 538 YGVVLLELLTGTKPVDLSQPPGQENLVTW 624
>BP032158
Length = 555
Score = 156 bits (394), Expect = 6e-39
Identities = 84/173 (48%), Positives = 118/173 (67%), Gaps = 5/173 (2%)
Frame = +2
Query: 16 FTGQQLRIATDNY--SNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVG 73
F+ +Q++ AT+N+ +N +G GGFG VYKG+ S G ++AVK L S +K+ + +F+ E+G
Sbjct: 14 FSLRQIKAATNNFDPANKIGEGGFGPVYKGVLSEGDVIAVKQL-SSKSKQGNREFINEIG 190
Query: 74 TIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLF--HETKV-LGYEKLHEIAIG 130
I + H NLV+LYG C E N + LVYEYM N SL R LF E K+ L + +I +G
Sbjct: 191 MISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGNEEQKLNLNWRTRMKICVG 370
Query: 131 TARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHIT 183
A+G+AYLHEE + +I+H DIK N+LLDK+ K++DFGLAK ENTHI+
Sbjct: 371 IAKGLAYLHEESRLKIVHRDIKATNVLLDKDLKAKISDFGLAKLDEEENTHIS 529
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 156 bits (394), Expect = 6e-39
Identities = 96/217 (44%), Positives = 130/217 (59%), Gaps = 5/217 (2%)
Frame = +1
Query: 16 FTGQQLRIATDN--YSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVG 73
F+ ++L AT+N Y N LG GGFG+VY G +G+ +AVK L+ SNK D +F EV
Sbjct: 28 FSLKELHSATNNFNYDNKLGEGGFGSVYWGQLWDGSQIAVKRLKVWSNKA-DMEFAVEVE 204
Query: 74 TIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLF--HETK-VLGYEKLHEIAIG 130
+ R+ H NL+ L G+C E +VY+YM N SL +L H ++ +L + + IAIG
Sbjct: 205 ILARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLLDWNRRMNIAIG 384
Query: 131 TARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGT 190
+A GI YLH + IIH DIK N+LLD +F +VADFG AK TH+T T +GT
Sbjct: 385 SAEGIVYLHHQATPHIIHRDIKASNVLLDSDFQARVADFGFAKLIPDGATHVT-TRVKGT 561
Query: 191 PGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNL 227
GY APE M CDV+SFG+LL E+ ++ L
Sbjct: 562 LGYLAPEYAMLGKANECCDVFSFGILLLELASGKKPL 672
>BI418434
Length = 478
Score = 153 bits (386), Expect = 5e-38
Identities = 82/158 (51%), Positives = 107/158 (66%)
Frame = +2
Query: 21 LRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHH 80
L++AT +S +G GGFGTV++G S+ ++VAVK L +++F AEV TIG I H
Sbjct: 2 LQLATRGFSEKVGHGGFGTVFQGELSDSSLVAVKRLERPGGG--EKEFRAEVSTIGNIQH 175
Query: 81 FNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVLGYEKLHEIAIGTARGIAYLHE 140
NLVRL GFC E + LVYEYM NG+L YL + L ++ ++AIGTA+GIAYLHE
Sbjct: 176 VNLVRLRGFCSENSHRLLVYEYMHNGALSAYLRKDGPCLSWDVRFQVAIGTAKGIAYLHE 355
Query: 141 ECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRE 178
E + I+H DIKP NILLD +F KV+DFGLAK R+
Sbjct: 356 ELRCCILHCDIKPENILLDSDFTAKVSDFGLAKLIGRD 469
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 151 bits (382), Expect = 1e-37
Identities = 100/277 (36%), Positives = 153/277 (55%), Gaps = 2/277 (0%)
Frame = +3
Query: 31 LLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYGFC 90
LLG GGFG VY G GT VA+K S + + E F E+ + ++ H +LV L G+C
Sbjct: 12 LLGVGGFGKVYYGEVDGGTKVAIKRGNPLSEQGVHE-FQTEIEMLSKLRHRHLVSLIGYC 188
Query: 91 FERNLIALVYEYMGNGSLDRYLFHETKV-LGYEKLHEIAIGTARGIAYLHEECQHRIIHY 149
E + LVY++M G+L +L+ K L +++ EI IG ARG+ YLH ++ IIH
Sbjct: 189 EENTEMILVYDHMAYGTLREHLYKTQKPPLPWKQRLEICIGAARGLHYLHTGAKYTIIHR 368
Query: 150 DIKPGNILLDKNFYPKVADFGLAKNC-NRENTHITMTGGRGTPGYAAPELWMPFPITHKC 208
D+K NILLD+ + KV+DFGL+K +NTH++ T +G+ GY PE + +T K
Sbjct: 369 DVKTTNILLDEKWVAKVSDFGLSKTGPTLDNTHVS-TVVKGSFGYLDPEYFRRQQLTDKS 545
Query: 209 DVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIA 268
DVYSFG++LFEI+ R L + Q W + G+L + + + E
Sbjct: 546 DVYSFGVVLFEILCARPALNPSLAKEQVSLAEWASHCYNKGILDQILDPYLKGKIAPECF 725
Query: 269 ERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEIPKT 305
++ + A+ CV + RP M V+ LE +L++ ++
Sbjct: 726 KKFAETAMKCVSDQGIERPSMGDVLWNLEFALQLQES 836
>TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2, complete
Length = 2946
Score = 147 bits (372), Expect = 2e-36
Identities = 106/291 (36%), Positives = 156/291 (53%), Gaps = 10/291 (3%)
Frame = +1
Query: 24 ATDNYS--NLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHF 81
AT+++S N LG GGFG VYKG+ +NG +AVK L +S + ++E F E+ I R+ H
Sbjct: 1660 ATNHFSLSNKLGEGGFGPVYKGLLANGQEIAVKRLSNTSGQGMEE-FKNEIKLIARLQHR 1836
Query: 82 NLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET--KVLGYEKLHEIAIGTARGIAYLH 139
NLV+L+G ++ + + M + L T K++ + K +I G ARG+ YLH
Sbjct: 1837 NLVKLFGCSVHQDENSHANKKM------KILLDSTRSKLVDWNKRLQIIDGIARGLLYLH 1998
Query: 140 EECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELW 199
++ + RIIH D+K NILLD PK++DFGLA+ + GT GY PE
Sbjct: 1999 QDSRLRIIHRDLKTSNILLDDEMNPKISDFGLARIFIGDQVEARTKRVMGTYGYMPPEYA 2178
Query: 200 MPFPITHKCDVYSFGMLLFEIIGRRRNLAIK------NTESQEWFPIWVWKKKDAGLLGE 253
+ + K DV+SFG+++ EII ++ N S W +W+ +++ L+ E
Sbjct: 2179 VHGSFSIKSDVFSFGVIVLEIISGKKIGRFYDPHHHLNLLSHAW-RLWI-EERPLELVDE 2352
Query: 254 AMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEIPK 304
+ I + R I VAL CVQ RPE RP M +V ML G E+PK
Sbjct: 2353 LLDDPVIPTE----ILRYIHVALLCVQRRPENRPDMLSIVLMLNGEKELPK 2493
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 144 bits (364), Expect = 2e-35
Identities = 102/309 (33%), Positives = 150/309 (48%), Gaps = 29/309 (9%)
Frame = +1
Query: 29 SNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYG 88
SN LG GGFG VYKG+ +NG +AVK L +S + ++E F EV I R+ H NLV+L G
Sbjct: 1462 SNKLGEGGFGPVYKGVLANGQEIAVKRLSNTSGQGMEE-FKNEVKLIARLQHRNLVKLLG 1638
Query: 89 FCFERNLIALVYEYMGNGSLDRYLFHETKVLGYEKLHEIAIGTARGIAYLHEECQHRIIH 148
+ + L+YE+M N SLD ++F + + RIIH
Sbjct: 1639 CSIHHDEMLLIYEFMHNRSLDYFIF---------------------------DSRLRIIH 1737
Query: 149 YDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELWMPFPITHKC 208
D+K NILLD PK++DFGLA+ + GT GY +PE + + K
Sbjct: 1738 RDLKTSNILLDSEMNPKISDFGLARIFTGDQVEAKTKRVMGTYGYMSPEYAVHGSFSVKS 1917
Query: 209 DVYSFGMLLFEIIGRRR------------------NLAI-----------KNTESQEWFP 239
DV+SFG+++ EII ++ N A+ +N ++++ +
Sbjct: 1918 DVFSFGVIVLEIISGKKIGRFCDPHHHRNLLSHSSNFAVFLIKALRICMFENVKNRKAWR 2097
Query: 240 IWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGS 299
+W+ +++ L+ E + I + R I +AL CVQ RPE RP M VV ML G
Sbjct: 2098 LWI-EERPLELVDELLDGLAIPTE----ILRYIHIALLCVQQRPEYRPDMLSVVLMLNGE 2262
Query: 300 LEIPKTFNP 308
E+PK P
Sbjct: 2263 KELPKPSLP 2289
>BP065880
Length = 509
Score = 142 bits (358), Expect = 9e-35
Identities = 81/144 (56%), Positives = 91/144 (62%)
Frame = -1
Query: 217 LFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVAL 276
LFEII RRRN ESQEWFP W W+K D+G L E I C EEKN EI ERM KVAL
Sbjct: 509 LFEIIVRRRNFDTSLPESQEWFPRWXWEKFDSGKLEELRIACRNEEKNMEIVERMSKVAL 330
Query: 277 WCVQYRPELRPIMSVVVKMLEGSLEIPKTFNPFQHLIDGTEFTTHSVQESNTYTTSVSSV 336
WCVQYR ELRP+MS VV+MLEG +EIPK NPFQ ++ GT T S
Sbjct: 329 WCVQYRSELRPLMSGVVRMLEGVVEIPKPLNPFQTMMVGT---------LPNPTNHPSDT 177
Query: 337 MVSDSSIVCATPIMRKYEIELASS 360
VS S CATPI+ ++ELASS
Sbjct: 176 TVS-SGCGCATPILTILDVELASS 108
>TC19546 similar to UP|Q40543 (Q40543) Protein-serine/threonine kinase ,
partial (46%)
Length = 731
Score = 141 bits (356), Expect = 1e-34
Identities = 78/182 (42%), Positives = 111/182 (60%)
Frame = +1
Query: 15 RFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGT 74
+++ + ++ AT N++ +LG G FGTVYK G +VAVKVL +S K+ + +F EV
Sbjct: 196 KYSYKDIQKATQNFTTILGQGSFGTVYKATMPTGEVVAVKVLAPNS-KQGEHEFQTEVHL 372
Query: 75 IGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVLGYEKLHEIAIGTARG 134
+GR+HH NLV L GFC ++ LVY++M NGSL L+ E K L +++ +IA+ + G
Sbjct: 373 LGRLHHRNLVNLVGFCVDKGQRILVYQFMSNGSLANLLYGEEKELSWDERLQIAMDISHG 552
Query: 135 IAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYA 194
I YLHE +IH D+K NILLD + VADFGL+K E +G +GT GY
Sbjct: 553 IEYLHEGAVPPVIHRDLKSANILLDDSMRAMVADFGLSK---EEIFDGRNSGLKGTYGYM 723
Query: 195 AP 196
P
Sbjct: 724 DP 729
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 141 bits (355), Expect = 2e-34
Identities = 91/225 (40%), Positives = 128/225 (56%), Gaps = 3/225 (1%)
Frame = +1
Query: 12 KPIRFTGQQLRIATDNYS--NLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFM 69
K + F+ Q+L AT+N+S N +G GGFG VY G A+K + + + +F+
Sbjct: 1039 KSMEFSYQELAKATNNFSLDNKIGQGGFGAVYYAEL-RGKKTAIKKM----DVQASTEFL 1203
Query: 70 AEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV-LGYEKLHEIA 128
E+ + +HH NLVRL G+C E +L LVYE++ NG+L +YL K L + +IA
Sbjct: 1204 CELKVLTHVHHLNLVRLIGYCVEGSLF-LVYEHIDNGNLGQYLHGSGKEPLPWSSRVQIA 1380
Query: 129 IGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGR 188
+ ARG+ Y+HE IH D+K NIL+DKN KVADFGL K N+ + T
Sbjct: 1381 LDAARGLEYIHEHTVPVYIHRDVKSANILIDKNLRGKVADFGLTKLIEVGNSTL-QTRLV 1557
Query: 189 GTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTE 233
GT GY PE I+ K DVY+FG++LFE+I +N +K E
Sbjct: 1558 GTFGYMPPEYAQYGDISPKIDVYAFGVVLFELIS-AKNAVLKTGE 1689
>AV422094
Length = 462
Score = 137 bits (346), Expect = 2e-33
Identities = 72/150 (48%), Positives = 101/150 (67%), Gaps = 2/150 (1%)
Frame = +2
Query: 12 KPIRFTGQQLRIATDNYS--NLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFM 69
KP F+ +L+ AT++++ N LG GGFG VYKGI ++GT++AVK L S++ QF+
Sbjct: 2 KPYTFSYSELKNATNDFNIDNKLGEGGFGPVYKGILNDGTVIAVKQLSLGSHQG-KSQFI 178
Query: 70 AEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVLGYEKLHEIAI 129
AE+ TI + H NLV+LYG C E + LVYEY+ N SLD+ LF + L + ++I +
Sbjct: 179 AEIATISAVQHRNLVKLYGCCIEGSKRLLVYEYLENKSLDQALFGKALSLNWSTRYDICL 358
Query: 130 GTARGIAYLHEECQHRIIHYDIKPGNILLD 159
G ARG+ YLHEE + RI+H D+K NILLD
Sbjct: 359 GVARGLTYLHEESRLRIVHRDVKASNILLD 448
>AV421705
Length = 341
Score = 137 bits (346), Expect = 2e-33
Identities = 63/109 (57%), Positives = 87/109 (79%), Gaps = 2/109 (1%)
Frame = +3
Query: 66 EQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHE--TKVLGYEK 123
+ F++EV TIGRIHH N+VRL G+C + ALVYE+M NGSLD+Y+F + + L Y++
Sbjct: 15 QDFISEVATIGRIHHINVVRLVGYCVDGKKRALVYEFMPNGSLDKYIFSKEGSAWLSYDQ 194
Query: 124 LHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLA 172
++EI++G ARG+AYLH+ C +I+H+DIKP NILLDK+F PKV+DFGLA
Sbjct: 195 IYEISLGIARGVAYLHQGCDMQILHFDIKPHNILLDKDFIPKVSDFGLA 341
>TC15149 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like protein kinase
protein (AT3g17840/MEB5_6), partial (45%)
Length = 1489
Score = 132 bits (333), Expect = 7e-32
Identities = 97/280 (34%), Positives = 148/280 (52%), Gaps = 11/280 (3%)
Frame = +3
Query: 29 SNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYG 88
+ +LG G FGT YK + G +VAVK L+ + +++F ++ T+G + H +LV L
Sbjct: 285 AEVLGKGTFGTAYKAVLETGLVVAVKRLKDVTIS--EKEFKDKIETVGAMDHQSLVPLRA 458
Query: 89 FCFERNLIALVYEYMGNGSLDRYLFHETKVLGYEKLH-----EIAIGTARGIAYLHEECQ 143
+ F R+ LVY+YM GSL L H K G L+ IA+G ARGI YLH +
Sbjct: 459 YYFSRDEKLLVYDYMPMGSLSA-LLHGNKGAGRTPLNWEIRSGIALGAARGIEYLHSQGP 635
Query: 144 HRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTP----GYAAPELW 199
+ + H +IK NILL K++ KV+DFGLA + G TP GY APE+
Sbjct: 636 N-VSHGNIKASNILLTKSYEAKVSDFGLAH----------LVGPSSTPNRVAGYRAPEVT 782
Query: 200 MPFPITHKCDVYSFGMLLFEII-GRRRNLAIKNTESQEWFPIWVWKKKDAGLLGEAMIVC 258
P ++ K DVYSFG+LL E++ G+ A+ N E + P WV E +
Sbjct: 783 DPRKVSQKADVYSFGVLLLELLTGKAPTHALLNEEGVD-LPRWVQSLVREEWTSEVFDLE 959
Query: 259 GIEEKN-KEIAERMIKVALWCVQYRPELRPIMSVVVKMLE 297
+ +N +E +++++A+ C P+ RP ++ V + +E
Sbjct: 960 LLRYQNVEEEMVQLLQLAVDCAAPYPDRRPSIAEVTRSIE 1079
>TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase, partial
(30%)
Length = 728
Score = 131 bits (330), Expect = 2e-31
Identities = 76/194 (39%), Positives = 111/194 (57%), Gaps = 9/194 (4%)
Frame = +3
Query: 13 PIRFTGQQLRIATDNYS--NLLGSGGFGTVYKGIFSNGTM-VAVKVLRGSSNKKIDEQFM 69
P F +L+ AT+N+ + LG GG+G VY+G+ + VAVK+ K D+ F+
Sbjct: 123 PREFNYVELKKATNNFDEKHKLGQGGYGVVYRGMLPKEKLEVAVKMFSRDKMKSTDD-FL 299
Query: 70 AEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHE-----TKVLGYEKL 124
+E+ I R+ H +LVRL G+C + ++ LVY+YM NGSLD ++F E T L +
Sbjct: 300 SELIIINRLRHKHLVRLQGWCHKNGVLLLVYDYMPNGSLDSHIFCEEGGTITTPLSWPLR 479
Query: 125 HEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHIT- 183
++I G A + YLH E +++H D+K NI+LD F K+ DFGLA+ E T
Sbjct: 480 YKIISGVASALNYLHNEYDQKVVHRDLKASNIMLDSEFNAKLGDFGLARALENEKISYTE 659
Query: 184 MTGGRGTPGYAAPE 197
+ G GT GY APE
Sbjct: 660 LEGVHGTMGYIAPE 701
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,088,773
Number of Sequences: 28460
Number of extensions: 85971
Number of successful extensions: 878
Number of sequences better than 10.0: 293
Number of HSP's better than 10.0 without gapping: 707
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 714
length of query: 362
length of database: 4,897,600
effective HSP length: 91
effective length of query: 271
effective length of database: 2,307,740
effective search space: 625397540
effective search space used: 625397540
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC130275.4


