

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
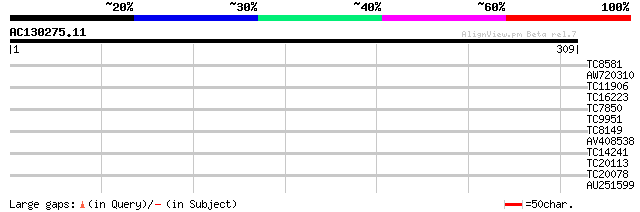
Query= AC130275.11 - phase: 0
(309 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8581 similar to UP|Q94II4 (Q94II4) NAM-like protein, partial (9%) 28 1.5
AW720310 28 2.6
TC11906 similar to UP|O82058 (O82058) Glucose-6-phosphate isomer... 27 4.4
TC16223 weakly similar to PIR|T04762|T04762 chitinase homolog T1... 27 4.4
TC7850 similar to GB|AAG24884.1|10946379|AF303941 ribulose-1,5-b... 27 5.7
TC9951 homologue to UP|TBB3_SOYBN (P28551) Tubulin beta chain (F... 27 5.7
TC8149 27 5.7
AV408538 26 7.5
TC14241 homologue to UP|Q9XQB4 (Q9XQB4) Photosystem I reaction c... 26 7.5
TC20113 weakly similar to UP|Q8GU51 (Q8GU51) MRP-like ABC transp... 26 7.5
TC20078 26 9.8
AU251599 26 9.8
>TC8581 similar to UP|Q94II4 (Q94II4) NAM-like protein, partial (9%)
Length = 1158
Score = 28.5 bits (62), Expect = 1.5
Identities = 10/24 (41%), Positives = 16/24 (66%)
Frame = +3
Query: 126 EYQEYSKDSSSTDECHCDDIIDGI 149
EY +YS SSS+ H DD+++ +
Sbjct: 42 EYTQYSNGSSSSSSSHLDDVMESL 113
>AW720310
Length = 612
Score = 27.7 bits (60), Expect = 2.6
Identities = 29/80 (36%), Positives = 43/80 (53%), Gaps = 5/80 (6%)
Frame = +2
Query: 169 ELIRKLKKSSK-SIKLVSIAPTELLKPHY----HKLYWANKDIINWVDYKFYNQTVSSAD 223
E I +LKK SK + L+ + EL K H + N++ IN V N T SS
Sbjct: 155 EDILQLKKMSKPATSLLVSSVQELAKQHMIEVPEQYLHPNQEPINVVP----NTTSSSQV 322
Query: 224 ELVNLYNKLLNEYGTDVKLL 243
+++L +KLL+E GT++K L
Sbjct: 323 PVIDL-DKLLSEDGTELKKL 379
>TC11906 similar to UP|O82058 (O82058) Glucose-6-phosphate isomerase
precursor (GPI) (Phosphoglucose isomerase) (PGI)
(Phosphohexose isomerase) (PHI) , partial (9%)
Length = 432
Score = 26.9 bits (58), Expect = 4.4
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = -3
Query: 49 GTLREVYSGDGKGKGEF 65
G +R ++GDG GKGEF
Sbjct: 286 GEVRRSFTGDGAGKGEF 236
>TC16223 weakly similar to PIR|T04762|T04762 chitinase homolog T16H5.170 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(35%)
Length = 570
Score = 26.9 bits (58), Expect = 4.4
Identities = 33/132 (25%), Positives = 56/132 (42%), Gaps = 14/132 (10%)
Frame = +2
Query: 70 NFNNFSPAKVAK-------LKKDHKNVKVIISIGGFGAEN-PFNPKEIESWSTKAKQSIK 121
N SPA A+ L+ + +VK ++SIGG G + N ++ S ++ K I
Sbjct: 200 NTLTISPANAAQFSTFTQTLQAKNPSVKTLLSIGGGGGHDLAANFSKMASQASSRKTFID 379
Query: 122 KLINEYQEYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDE--FSSCIGE----LIRKLK 175
I ++ + G+D+++EY + N D+ F S I E + R
Sbjct: 380 SSIQLARKNN--------------FHGLDLDWEYPSTNTDKTNFGSLIKEWRAAVARDSS 517
Query: 176 KSSKSIKLVSIA 187
+ K L+S A
Sbjct: 518 NTGKPALLLSAA 553
>TC7850 similar to GB|AAG24884.1|10946379|AF303941
ribulose-1,5-bisphosphate carboxylase small subunit
rbcS3 {Glycine max;} , partial (98%)
Length = 796
Score = 26.6 bits (57), Expect = 5.7
Identities = 20/71 (28%), Positives = 34/71 (47%), Gaps = 11/71 (15%)
Frame = -2
Query: 98 FGAENPFNPKEI----------ESWSTKAKQSIKKLINEYQEYSKD-SSSTDECHCDDII 146
F A NP P ++ E W K ++ +K L+ ++ + SST C+ D+I
Sbjct: 321 FQARNPSLPDKVFYFLCQLLHGEWW--KVRKGLKLLLANWRPHLHAVDSSTIACNGGDVI 148
Query: 147 DGIDINYEYSN 157
G+ N+E S+
Sbjct: 147 VGLSCNWETSH 115
>TC9951 homologue to UP|TBB3_SOYBN (P28551) Tubulin beta chain (Fragment),
partial (44%)
Length = 600
Score = 26.6 bits (57), Expect = 5.7
Identities = 15/55 (27%), Positives = 25/55 (45%)
Frame = -1
Query: 9 AVVKPIIFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGKGKG 63
AVV + + D+P+ D A + D+E+F LG + E +G+ G
Sbjct: 177 AVVLSGVVDAVLVADDLPEFCADLVAALAALDVEDFSHFLGKIEEKVRENGRENG 13
>TC8149
Length = 564
Score = 26.6 bits (57), Expect = 5.7
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = -2
Query: 60 KGKGEFYRTWNFNNFSPAKVAKLKKDHKNVKV 91
K + Y+ +NN SP K+ K K+DH K+
Sbjct: 422 KSQNPEYKVK*YNNTSPLKLIKPKQDHHMKKI 327
>AV408538
Length = 430
Score = 26.2 bits (56), Expect = 7.5
Identities = 11/33 (33%), Positives = 17/33 (51%)
Frame = +3
Query: 109 IESWSTKAKQSIKKLINEYQEYSKDSSSTDECH 141
+ESW T + +KLI YS + + +E H
Sbjct: 12 VESWMTTKNLNPRKLIPAMMRYSSEPHAKNETH 110
>TC14241 homologue to UP|Q9XQB4 (Q9XQB4) Photosystem I reaction center
subunit III, partial (86%)
Length = 1081
Score = 26.2 bits (56), Expect = 7.5
Identities = 12/29 (41%), Positives = 20/29 (68%)
Frame = +1
Query: 107 KEIESWSTKAKQSIKKLINEYQEYSKDSS 135
KE + ++ + KQSIKKL + + Y+ DS+
Sbjct: 322 KESKQFAKREKQSIKKLESSLKLYAPDSA 408
>TC20113 weakly similar to UP|Q8GU51 (Q8GU51) MRP-like ABC transporter
(Fragment), partial (3%)
Length = 547
Score = 26.2 bits (56), Expect = 7.5
Identities = 17/68 (25%), Positives = 31/68 (45%)
Frame = -1
Query: 104 FNPKEIESWSTKAKQSIKKLINEYQEYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEF 163
F P+E + +++KK + E + + D C C+ +D I Y YS+ +
Sbjct: 451 FLPEESNLLTRGKIENVKKKYKKRIEL*RGENLRDLCDCNCPLDRDVIGYNYSSLH---I 281
Query: 164 SSCIGELI 171
S C+ + I
Sbjct: 280 SKCVSQRI 257
>TC20078
Length = 473
Score = 25.8 bits (55), Expect = 9.8
Identities = 20/59 (33%), Positives = 28/59 (46%), Gaps = 1/59 (1%)
Frame = +2
Query: 30 KDFPAEMINDDIEEFHFI-LGTLREVYSGDGKGKGEFYRTWNFNNFSPAKVAKLKKDHK 87
KD +I D + +F F+ +G R + G EFY +FN AK+AK K K
Sbjct: 26 KDISPMLIRDFVTKFTFVGVGIKRFM------GALEFYGLVDFNYVDVAKLAKAKWPEK 184
>AU251599
Length = 350
Score = 25.8 bits (55), Expect = 9.8
Identities = 16/37 (43%), Positives = 22/37 (59%)
Frame = -2
Query: 179 KSIKLVSIAPTELLKPHYHKLYWANKDIINWVDYKFY 215
K IK V+IA TEL + HK+ K ++ + YKFY
Sbjct: 133 KPIKQVTIAITEL*FHNSHKIRGNKKKVVMF--YKFY 29
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,649,586
Number of Sequences: 28460
Number of extensions: 77537
Number of successful extensions: 255
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 254
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 255
length of query: 309
length of database: 4,897,600
effective HSP length: 90
effective length of query: 219
effective length of database: 2,336,200
effective search space: 511627800
effective search space used: 511627800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC130275.11


