

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130200.9 - phase: 0
(393 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
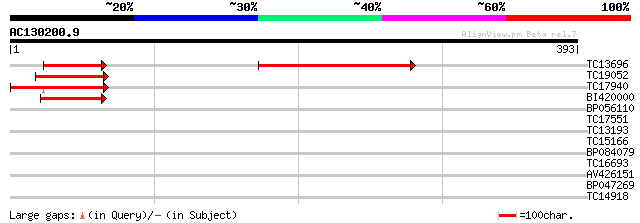
Score E
Sequences producing significant alignments: (bits) Value
TC13696 similar to GB|AAM19869.1|20453261|AY097353 AT4g31410/F8F... 114 3e-26
TC19052 similar to UP|Q9LUA0 (Q9LUA0) Gb|AAF16548.1 (AT3g25910/M... 92 2e-19
TC17940 91 4e-19
BI420000 83 7e-17
BP056110 36 0.013
TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, part... 31 0.41
TC13193 similar to UP|Q9ASQ6 (Q9ASQ6) At1g29720/T3M22_6, partial... 30 0.53
TC15166 similar to GB|AAK28312.1|13506739|AF224702 WRKY DNA-bind... 28 2.0
BP084079 28 2.0
TC16693 weakly similar to PIR|T51842|T51842 RING-H2 finger prote... 28 2.6
AV426151 27 4.5
BP047269 27 5.9
TC14918 similar to UP|Q8VAJ1 (Q8VAJ1) Wsv418 (WSSV477), partial ... 27 7.7
>TC13696 similar to GB|AAM19869.1|20453261|AY097353 AT4g31410/F8F16_230
{Arabidopsis thaliana;}, partial (49%)
Length = 517
Score = 114 bits (285), Expect = 3e-26
Identities = 51/109 (46%), Positives = 70/109 (63%)
Frame = +3
Query: 173 SEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHP 232
S+ + L CPLCRG V GW VV++AR +L+ K R C + C+F G Y EL+ HA HP
Sbjct: 189 SDCRQCKLTCPLCRGDVSGWTVVDKARMHLDEKNRRCDEERCAFVGSYSELQEHAEVNHP 368
Query: 233 TSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENG 281
+ PS +DP R+ W+ F++ E D++S I S IP VV+GDYV+E G
Sbjct: 369 HAHPSKIDPARQLDWENFQQSSEIIDVLSTIHSEIPRGVVLGDYVIEYG 515
Score = 77.8 bits (190), Expect = 3e-15
Identities = 30/44 (68%), Positives = 37/44 (83%)
Frame = +3
Query: 24 VSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFK 67
V CPIC++ PHN+VLL CSS+DKGCR ++CDT+ HSNC DRFK
Sbjct: 6 VKCPICLEFPHNSVLLQCSSYDKGCRPFVCDTNQLHSNCFDRFK 137
>TC19052 similar to UP|Q9LUA0 (Q9LUA0) Gb|AAF16548.1 (AT3g25910/MPE11_6),
partial (14%)
Length = 507
Score = 91.7 bits (226), Expect = 2e-19
Identities = 35/50 (70%), Positives = 45/50 (90%)
Frame = +1
Query: 19 RELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKK 68
+E ++ CP+CM+HPHNAVLL+CSSH+KGCR Y+C+TSYRHSNCLD+F K
Sbjct: 334 KEWEDARCPVCMEHPHNAVLLICSSHEKGCRPYMCNTSYRHSNCLDQFCK 483
>TC17940
Length = 662
Score = 90.5 bits (223), Expect = 4e-19
Identities = 41/73 (56%), Positives = 52/73 (71%), Gaps = 5/73 (6%)
Frame = +1
Query: 1 MAGFKRRLCNDSDMHALHRELD-----EVSCPICMDHPHNAVLLLCSSHDKGCRSYICDT 55
+A KR C+D + LD +V+C +CM+ PHNAVLLLCSSHDKGCR+Y+C T
Sbjct: 298 LAACKRVTCDDMCQKKCTKALDTKEWEDVTCSVCMEFPHNAVLLLCSSHDKGCRAYMCGT 477
Query: 56 SYRHSNCLDRFKK 68
S RHSNCLD++KK
Sbjct: 478 SLRHSNCLDQYKK 516
>BI420000
Length = 502
Score = 83.2 bits (204), Expect = 7e-17
Identities = 33/46 (71%), Positives = 41/46 (88%)
Frame = +2
Query: 22 DEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFK 67
D+V+CPIC+D PHN+VLL CSS DKGCR+++CDT+ HSNCLDRFK
Sbjct: 323 DDVACPICLDFPHNSVLLQCSSCDKGCRAFVCDTNQLHSNCLDRFK 460
>BP056110
Length = 479
Score = 35.8 bits (81), Expect = 0.013
Identities = 17/36 (47%), Positives = 21/36 (58%)
Frame = +3
Query: 41 CSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSKEN 76
CSSHDK C S C+ S+ SNC ++KL S N
Sbjct: 180 CSSHDKDCPSK-CEASFCDSNCFYHYRKLTLKSSNN 284
>TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, partial (22%)
Length = 505
Score = 30.8 bits (68), Expect = 0.41
Identities = 17/40 (42%), Positives = 19/40 (47%)
Frame = -1
Query: 269 GAVVVGDYVLENGDGIGRLPPGGGREGSNGNGNVPWLTTT 308
G V D V E G+G GRL G G G G V W+ T
Sbjct: 169 GEEVENDVVREGGNGGGRLLRWFGDGGGGGGGGVVWIGET 50
>TC13193 similar to UP|Q9ASQ6 (Q9ASQ6) At1g29720/T3M22_6, partial (12%)
Length = 513
Score = 30.4 bits (67), Expect = 0.53
Identities = 15/43 (34%), Positives = 22/43 (50%)
Frame = +1
Query: 65 RFKKLRDNSKENPNLQSSLINTNNSSGNSFDINFTMQSDMHDV 107
RFK +RD + N S T+NS+G + T +DMH +
Sbjct: 196 RFKAMRDIRQHKENHSLSTSQTDNSTGLTHSFPSTSGNDMHQI 324
>TC15166 similar to GB|AAK28312.1|13506739|AF224702 WRKY DNA-binding
protein 6 {Arabidopsis thaliana;} , partial (26%)
Length = 668
Score = 28.5 bits (62), Expect = 2.0
Identities = 14/30 (46%), Positives = 17/30 (56%), Gaps = 1/30 (3%)
Frame = +1
Query: 6 RRLCNDSDMHALHR-ELDEVSCPICMDHPH 34
R LCN D + LHR + +SCP D PH
Sbjct: 46 RTLCNSQDRNILHRSKYPRISCP---DQPH 126
>BP084079
Length = 508
Score = 28.5 bits (62), Expect = 2.0
Identities = 20/54 (37%), Positives = 29/54 (53%)
Frame = +3
Query: 110 LLLYQNEINTLLSVGIPQGSRQGDAQDPSRHLDQHDEGILETADSENLQDRAVL 163
L L QNE T+ + +G A+ S LD +G+LE A+ + L DRAV+
Sbjct: 66 LSLVQNEFATISGI-------KGKAEKLSHDLDLI-KGVLEDAEKKQLTDRAVM 203
>TC16693 weakly similar to PIR|T51842|T51842 RING-H2 finger protein RHA2a
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (35%)
Length = 442
Score = 28.1 bits (61), Expect = 2.6
Identities = 11/25 (44%), Positives = 19/25 (76%)
Frame = +2
Query: 182 CPLCRGTVLGWEVVEEARNYLNNKK 206
CPLCR VL +VV + R++L++++
Sbjct: 347 CPLCRKQVLPCDVVSKHRHFLSHEE 421
>AV426151
Length = 331
Score = 27.3 bits (59), Expect = 4.5
Identities = 15/41 (36%), Positives = 20/41 (48%), Gaps = 2/41 (4%)
Frame = -3
Query: 147 GILETADSENLQDRAVLEE--ELDVDNSSEDSKSSLHCPLC 185
G T +SEN ++R EE E + + S HCPLC
Sbjct: 275 GRFRTRESENEENRENNEEKEESRLSEKPDHSTHCFHCPLC 153
>BP047269
Length = 619
Score = 26.9 bits (58), Expect = 5.9
Identities = 10/17 (58%), Positives = 11/17 (63%)
Frame = -2
Query: 332 GWSRHRRSSDRRRYLWG 348
GW R RR RRR+ WG
Sbjct: 201 GWRRGRRRKPRRRF*WG 151
>TC14918 similar to UP|Q8VAJ1 (Q8VAJ1) Wsv418 (WSSV477), partial (11%)
Length = 629
Score = 26.6 bits (57), Expect = 7.7
Identities = 11/39 (28%), Positives = 23/39 (58%)
Frame = +1
Query: 71 DNSKENPNLQSSLINTNNSSGNSFDINFTMQSDMHDVDE 109
DN +E + +++ N + GN+ + +T QSD+ + +E
Sbjct: 43 DNIEEGSDKFTTVANPSEKKGNNKVVCYTRQSDLDEFEE 159
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.133 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,317,938
Number of Sequences: 28460
Number of extensions: 113700
Number of successful extensions: 715
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 708
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 715
length of query: 393
length of database: 4,897,600
effective HSP length: 92
effective length of query: 301
effective length of database: 2,279,280
effective search space: 686063280
effective search space used: 686063280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC130200.9


