

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
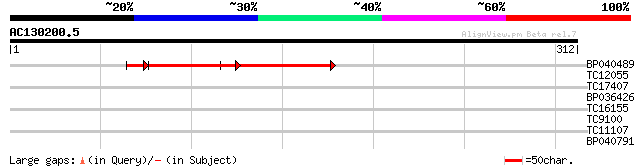
Query= AC130200.5 - phase: 0
(312 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP040489 79 3e-16
TC12055 homologue to UP|Q8L3R8 (Q8L3R8) AT3g08530/T8G24_1, parti... 32 0.18
TC17407 29 0.90
BP036426 28 1.5
TC16155 28 2.6
TC9100 similar to UP|Q8W544 (Q8W544) NADH-ubiquinone oxidoreduct... 27 5.8
TC11107 26 7.6
BP040791 26 7.6
>BP040489
Length = 555
Score = 79.0 bits (193), Expect(2) = 3e-16
Identities = 38/51 (74%), Positives = 45/51 (87%)
Frame = +3
Query: 77 DKVDSALKDMAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYK 127
D+VDSALKDMA+VMKQ +R+EEAIEAIKS R C+ +QESLDN+LLDLYK
Sbjct: 36 DRVDSALKDMAIVMKQQNRSEEAIEAIKSLRSRCSDQAQESLDNILLDLYK 188
Score = 77.8 bits (190), Expect = 2e-15
Identities = 37/63 (58%), Positives = 50/63 (78%)
Frame = +2
Query: 117 SLDNVLLDLYKKCGRVEEQIELLKRKLRLIYQGEAFNGRTTKTARSHGKKFQVSIKQETA 176
++D + + L ++CGR+++QI LL+ KL LI QG AFNG+ TKTARS GKKFQVS++QE
Sbjct: 254 NVDKIEVFLLQRCGRLDDQIALLRHKLYLIQQGLAFNGKRTKTARSQGKKFQVSVEQEAT 433
Query: 177 RLL 179
RLL
Sbjct: 434 RLL 442
Score = 21.9 bits (45), Expect(2) = 3e-16
Identities = 9/12 (75%), Positives = 9/12 (75%)
Frame = +1
Query: 65 AIVYFWKAINAG 76
AI FW AINAG
Sbjct: 1 AIPLFWAAINAG 36
>TC12055 homologue to UP|Q8L3R8 (Q8L3R8) AT3g08530/T8G24_1, partial (22%)
Length = 483
Score = 31.6 bits (70), Expect = 0.18
Identities = 29/96 (30%), Positives = 44/96 (45%), Gaps = 5/96 (5%)
Frame = +1
Query: 51 KAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMKQLDRAEEAIEAIKSFRGL- 109
KA H +LV K VA+ N V+ AL ++ V + DR E+I+ +F +
Sbjct: 49 KAGHLRLV-KPYMVAV-----QSNNVSAVNEALNEIYVEEEDYDRLRESIDLHDNFDQIG 210
Query: 110 ----CNKHSQESLDNVLLDLYKKCGRVEEQIELLKR 141
KH + V +YKK GR ++ I L K+
Sbjct: 211 LAQKIEKHELLEMRRVAAYIYKKAGRWKQSIALSKK 318
>TC17407
Length = 517
Score = 29.3 bits (64), Expect = 0.90
Identities = 15/47 (31%), Positives = 23/47 (48%)
Frame = -1
Query: 24 LKKSSKGKKEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFW 70
LKKS K KK H+ H + YVK ++ K+ ++A + W
Sbjct: 406 LKKSKKKKKIIRRHISHTMDMNGRNYVKKHLVYIIAKNEKMASMAIW 266
>BP036426
Length = 558
Score = 28.5 bits (62), Expect = 1.5
Identities = 19/73 (26%), Positives = 34/73 (46%)
Frame = -1
Query: 133 EEQIELLKRKLRLIYQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNY 192
EE+ L+R LR + +R H + + K ETA ++ ++GW M+K
Sbjct: 222 EEEDNFLRRDLR------------DRRSRGHRRGIERGSKGETAAVVSSMGWDGMEKVVE 79
Query: 193 MMAEVVFKKAQMI 205
++ VV A ++
Sbjct: 78 VIIGVVVVVAVVV 40
>TC16155
Length = 703
Score = 27.7 bits (60), Expect = 2.6
Identities = 20/71 (28%), Positives = 36/71 (50%), Gaps = 11/71 (15%)
Frame = +1
Query: 100 IEAIKSFRGLC-NKHSQESLDNVLLD---------LYKK-CGRVEEQIELLKRKLRLIYQ 148
++ IK R L N + + V+LD L+K CG++ Q+ + + KLR ++
Sbjct: 166 LQVIKIIRKLVLNNYRHQLYQVVILDATHSFRV*ILHKS*CGKISLQVSIQEFKLRWVWM 345
Query: 149 GEAFNGRTTKT 159
+ FNG T++
Sbjct: 346 IQLFNGCGTES 378
>TC9100 similar to UP|Q8W544 (Q8W544) NADH-ubiquinone oxidoreductase
(Fragment), partial (97%)
Length = 712
Score = 26.6 bits (57), Expect = 5.8
Identities = 17/46 (36%), Positives = 26/46 (55%)
Frame = +1
Query: 101 EAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLI 146
+A++SF K QE D ++ CG+VEE IE + +L+LI
Sbjct: 193 KAVESFTRHRLKACQEEEDWEAIENRLGCGQVEELIEEAQDELKLI 330
>TC11107
Length = 841
Score = 26.2 bits (56), Expect = 7.6
Identities = 14/38 (36%), Positives = 20/38 (51%)
Frame = -2
Query: 131 RVEEQIELLKRKLRLIYQGEAFNGRTTKTARSHGKKFQ 168
R Q+E L+ KL QG + + TK++ SH K Q
Sbjct: 429 RTSHQVEKLQIKLSETQQGAYPHPKPTKSSHSHAKSQQ 316
>BP040791
Length = 450
Score = 26.2 bits (56), Expect = 7.6
Identities = 15/41 (36%), Positives = 22/41 (53%)
Frame = -3
Query: 131 RVEEQIELLKRKLRLIYQGEAFNGRTTKTARSHGKKFQVSI 171
R EE+++L R +RL +GE F K + KK+ SI
Sbjct: 295 RPEEEVKLSPRTMRLEVEGERFEHCV*KLYVNRKKKYSGSI 173
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,387,868
Number of Sequences: 28460
Number of extensions: 46687
Number of successful extensions: 209
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 208
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 209
length of query: 312
length of database: 4,897,600
effective HSP length: 90
effective length of query: 222
effective length of database: 2,336,200
effective search space: 518636400
effective search space used: 518636400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC130200.5


