

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC129091.7 + phase: 0
(359 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
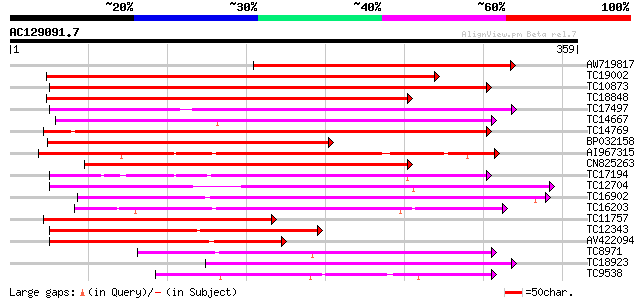
Score E
Sequences producing significant alignments: (bits) Value
AW719817 283 3e-77
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 239 7e-64
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 233 5e-62
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 230 3e-61
TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2,... 222 9e-59
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 218 1e-57
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 207 2e-54
BP032158 204 2e-53
AI967315 197 2e-51
CN825263 197 2e-51
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 193 3e-50
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 189 5e-49
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 188 1e-48
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 182 8e-47
TC11757 UP|Q91W27 (Q91W27) Tuberoinfundibular 39 residue protein... 173 5e-44
TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine prote... 170 4e-43
AV422094 163 4e-41
TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P... 162 6e-41
TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1,... 155 1e-38
TC9538 similar to UP|RLK5_ARATH (P47735) Receptor-like protein k... 146 5e-36
>AW719817
Length = 499
Score = 283 bits (724), Expect = 3e-77
Identities = 135/166 (81%), Positives = 154/166 (92%)
Frame = +1
Query: 155 ELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIAGTTGYLAPEYALG 214
EL HIVHRDIKASN+LLD+DFNPKIGDFG+AKLFPDDITHISTRIAGTTGYLAPEYA+G
Sbjct: 1 ELVPHIVHRDIKASNILLDRDFNPKIGDFGLAKLFPDDITHISTRIAGTTGYLAPEYAMG 180
Query: 215 GQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKWLALVDPEMEE 274
GQLT KADVYSFGVLILE++SGK S+RTNW GS+K LLEWAWQLHEE K L LVDP+M++
Sbjct: 181 GQLTMKADVYSFGVLILEVVSGKGSARTNWGGSNKYLLEWAWQLHEEGKLLELVDPDMDD 360
Query: 275 FPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEIQLNDKQLT 320
FPE+ VI+Y+KVA FCTQAAA RRPLM+QVVDML+K+I+LN+KQLT
Sbjct: 361 FPEE*VIRYMKVAFFCTQAAASRRPLMSQVVDMLTKKIRLNEKQLT 498
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 239 bits (609), Expect = 7e-64
Identities = 118/249 (47%), Positives = 162/249 (64%)
Frame = +1
Query: 24 RHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEI 83
R FS KEL AT+N++ NK+G GGFG+VY G L DG +IAVK L V S + EF E+
Sbjct: 22 RVFSLKELHSATNNFNYDNKLGEGGFGSVYWGQLWDGSQIAVKRLKVWSNKADMEFAVEV 201
Query: 84 KTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICI 143
+ L+ V+H NL+ L G+C +G R +VY+Y+ N +L + L G+ S + W R I I
Sbjct: 202 EILARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLLDWNRRMNIAI 381
Query: 144 GTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIAGT 203
G+A+G+ YLH + T HI+HRDIKASNVLLD DF ++ DFG AKL PD TH++TR+ GT
Sbjct: 382 GSAEGIVYLHHQATPHIIHRDIKASNVLLDSDFQARVADFGFAKLIPDGATHVTTRVKGT 561
Query: 204 TGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEK 263
GYLAPEYA+ G+ + DV+SFG+L+LE+ SGK +S+ +WA L +K
Sbjct: 562 LGYLAPEYAMLGKANECCDVFSFGILLLELASGKKPLEKLSSTVKRSINDWALPLACAKK 741
Query: 264 WLALVDPEM 272
+ DP +
Sbjct: 742 FTEFADPRL 768
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 233 bits (593), Expect = 5e-62
Identities = 118/283 (41%), Positives = 177/283 (61%), Gaps = 3/283 (1%)
Frame = +1
Query: 26 FSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKT 85
F +EL+ AT N+ N IG GGFG VY+G L G +AVK LS +QG +EF+ E+
Sbjct: 415 FGFRELADATRNFKEANLIGEGGFGKVYKGRLTTGEAVAVKQLSHDGRQGFQEFVMEVLM 594
Query: 86 LSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGT 145
LS + H+NLV L+G+C G R +VYEY+ G+L L + W R + +G
Sbjct: 595 LSLLHHTNLVRLIGYCTDGDQRLLVYEYMPMGSLEDHLFELSHDKEPLNWSTRMKVAVGA 774
Query: 146 AKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFP-DDITHISTRIAGTT 204
A+GL YLH +++RD+K++N+LLD +FNPK+ DFG+AKL P D TH+STR+ GT
Sbjct: 775 ARGLEYLHCTADPPVIYRDLKSANILLDNEFNPKLSDFGLAKLGPVGDNTHVSTRVMGTY 954
Query: 205 GYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKW 264
GY APEYA+ G+LT K+D+YSFGV++LE+++G+ + T+ ++L+ WA + +
Sbjct: 955 GYCAPEYAMSGKLTLKSDIYSFGVVLLELLTGRRAIDTSRRPGEQNLVSWARPYFSDRRR 1134
Query: 265 LA-LVDPEME-EFPEKEVIKYIKVALFCTQAAARRRPLMTQVV 305
+VDP ++ FP + + + I + C Q + RPL+T +V
Sbjct: 1135FGHMVDPLLQGRFPSRCLHQAIAITAMCLQEQPKFRPLITDIV 1263
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 230 bits (586), Expect = 3e-61
Identities = 107/232 (46%), Positives = 159/232 (68%)
Frame = +2
Query: 24 RHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEI 83
R F+ KEL AT + NK+G GGFG+VY G DG +IAVK L + + EF E+
Sbjct: 260 RIFTYKELHAATGGFSDDNKLGEGGFGSVYWGRTSDGLQIAVKKLKAMNSKAEMEFAVEV 439
Query: 84 KTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICI 143
+ L V+H NL+ L G+C+ R +VY+Y+ N +L + L G+ ++ V++ W++R I I
Sbjct: 440 EVLGRVRHKNLLGLRGYCVGDDQRLIVYDYMPNLSLLSHLHGQFAVEVQLNWQKRMKIAI 619
Query: 144 GTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIAGT 203
G+A+G+ YLH E+T HI+HRDIKASNVLL+ DF P + DFG AKL P+ ++H++TR+ GT
Sbjct: 620 GSAEGILYLHHEVTPHIIHRDIKASNVLLNSDFEPLVADFGFAKLIPEGVSHMTTRVKGT 799
Query: 204 TGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWA 255
GYLAPEYA+ G++++ DVYSFG+L+LE+++G+ G +++ EWA
Sbjct: 800 LGYLAPEYAMWGKVSESCDVYSFGILLLELVTGRKPIEKLPGGVKRTITEWA 955
>TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2, complete
Length = 2946
Score = 222 bits (565), Expect = 9e-59
Identities = 126/298 (42%), Positives = 179/298 (59%), Gaps = 2/298 (0%)
Frame = +1
Query: 26 FSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKT 85
F +S AT+++ L NK+G GGFG VY+G L +G++IAVK LS S QG+ EF EIK
Sbjct: 1636 FDFSTISSATNHFSLSNKLGEGGFGPVYKGLLANGQEIAVKRLSNTSGQGMEEFKNEIKL 1815
Query: 86 LSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGT 145
++ ++H NLV+L G + + + N +L + S + W +R I G
Sbjct: 1816 IARLQHRNLVKLFGCSVHQDENS-------HANKKMKILLDSTRSKLVDWNKRLQIIDGI 1974
Query: 146 AKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHIST-RIAGTT 204
A+GL YLH++ I+HRD+K SN+LLD + NPKI DFG+A++F D T R+ GT
Sbjct: 1975 ARGLLYLHQDSRLRIIHRDLKTSNILLDDEMNPKISDFGLARIFIGDQVEARTKRVMGTY 2154
Query: 205 GYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKW 264
GY+ PEYA+ G + K+DV+SFGV++LEIISGK R H +LL AW+L EE+
Sbjct: 2155 GYMPPEYAVHGSFSIKSDVFSFGVIVLEIISGKKIGRFYDPHHHLNLLSHAWRLWIEERP 2334
Query: 265 LALVDPEMEE-FPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEIQLNDKQLTA 321
L LVD +++ E+++YI VAL C Q RP M +V ML+ E +L +L A
Sbjct: 2335 LELVDELLDDPVIPTEILRYIHVALLCVQRRPENRPDMLSIVLMLNGEKELPKPRLPA 2508
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 218 bits (556), Expect = 1e-57
Identities = 121/287 (42%), Positives = 171/287 (59%), Gaps = 8/287 (2%)
Frame = +2
Query: 30 ELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVR-EFLTEIKTLSN 88
EL TDN+ IG G +G VY TL DG +AVK L V S+ EFLT++ +S
Sbjct: 335 ELKEKTDNFGSKALIGEGSYGRVYYATLNDGNAVAVKKLDVSSEPETNNEFLTQVSMVSR 514
Query: 89 VKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLS-----VKMKWRERSTICI 143
+K+ N VEL G+C++G R + YE+ G+LH L G+K + + W +R I +
Sbjct: 515 LKNDNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWIQRVRIAV 694
Query: 144 GTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHI-STRIAG 202
A+GL YLHE++ I+HRDI++SNVL+ +D+ KI DF ++ PD + STR+ G
Sbjct: 695 DAARGLEYLHEKVQPAIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAARLHSTRVLG 874
Query: 203 TTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEE 262
T GY APEYA+ GQLT+K+DVYSFGV++LE+++G+ +SL+ WA E+
Sbjct: 875 TFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLSED 1054
Query: 263 KWLALVDPEME-EFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
K VDP+++ E+P K V K VA C Q A RP M+ VV L
Sbjct: 1055KVKQCVDPKLKGEYPPKGVAKLAAVAALCVQYEAEFRPNMSIVVKAL 1195
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 207 bits (528), Expect = 2e-54
Identities = 113/286 (39%), Positives = 174/286 (60%), Gaps = 2/286 (0%)
Frame = +3
Query: 22 NIRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLT 81
+I+ F+ + + +AT+ Y IG GGFG+VY+GTL DG+++AVK S S QG REF
Sbjct: 2121 SIQAFTLEYIEVATERYK--TLIGEGGFGSVYRGTLNDGQEVAVKVRSSTSTQGTREFDN 2294
Query: 82 EIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTI 141
E+ LS ++H NLV L+G+C + + +VY ++ NG+L L G+ + + W R +I
Sbjct: 2295 ELNLLSAIQHENLVPLLGYCNESDQQILVYPFMSNGSLQDRLYGEPAKRKILDWPTRLSI 2474
Query: 142 CIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDD-ITHISTRI 200
+G A+GLAYLH + ++HRDIK+SN+LLD K+ DFG +K P + +++S +
Sbjct: 2475 ALGAARGLAYLHTFPGRSVIHRDIKSSNILLDHSMCAKVADFGFSKYAPQEGDSYVSLEV 2654
Query: 201 AGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHE 260
GT GYL PEY QL++K+DV+SFGV++LEI+SG+ + SL+EWA
Sbjct: 2655 RGTAGYLDPEYYKTQQLSEKSDVFSFGVVLLEIVSGREPLNIKRPRTEWSLVEWATPYIR 2834
Query: 261 EEKWLALVDPEMEEFPEKEVI-KYIKVALFCTQAAARRRPLMTQVV 305
K +VDP ++ E + + ++VAL C + + RP M +V
Sbjct: 2835 GSKVDEIVDPGIKGGYHAEAMWRVVEVALQCLEPFSTYRPSMVAIV 2972
>BP032158
Length = 555
Score = 204 bits (518), Expect = 2e-53
Identities = 103/181 (56%), Positives = 128/181 (69%)
Frame = +2
Query: 25 HFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIK 84
+FS +++ AT+N+ NKIG GGFG VY+G L +G IAVK LS SKQG REF+ EI
Sbjct: 11 YFSLRQIKAATNNFDPANKIGEGGFGPVYKGVLSEGDVIAVKQLSSKSKQGNREFINEIG 190
Query: 85 TLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIG 144
+S ++H NLV+L G CI+G +VYEY+EN +L AL G + + + WR R IC+G
Sbjct: 191 MISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGNEEQKLNLNWRTRMKICVG 370
Query: 145 TAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIAGTT 204
AKGLAYLHEE IVHRDIKA+NVLLDKD KI DFG+AKL ++ THISTRIAGT
Sbjct: 371 IAKGLAYLHEESRLKIVHRDIKATNVLLDKDLKAKISDFGLAKLDEEENTHISTRIAGTI 550
Query: 205 G 205
G
Sbjct: 551 G 553
>AI967315
Length = 1308
Score = 197 bits (501), Expect = 2e-51
Identities = 119/298 (39%), Positives = 184/298 (60%), Gaps = 6/298 (2%)
Frame = +1
Query: 19 PLDNIRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLS--VGSKQGV 76
P + + FS +EL AT+ + N +G+GG+ VY+G L+ G +IAVK L+ ++
Sbjct: 91 PRPSWKCFSYEELFHATNGFSSENMVGKGGYAEVYKGRLESGDEIAVKRLTRTCRDERKE 270
Query: 77 REFLTEIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWR 136
+EFLTEI T+ +V HSN++ L+G CI +V+E G++ + + +K + W+
Sbjct: 271 KEFLTEIGTIGHVCHSNVMPLLGCCIDN-GLYLVFELSTVGSVASLIHDEKM--APLDWK 441
Query: 137 ERSTICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHI 196
R I +GTA+GL YLH+ + I+HRDIKASN+LL +DF P+I DFG+AK P TH
Sbjct: 442 TRYKIVLGTARGLHYLHKGCQRRIIHRDIKASNILLTEDFEPQISDFGLAKWLPSQWTHH 621
Query: 197 S-TRIAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWA 255
S I GT G+LAPEY + G + +K DV++FGV +LE+ISG R DGSH+SL WA
Sbjct: 622 SIAPIEGTFGHLAPEYYMHGVVDEKTDVFAFGVFLLEVISG----RKPVDGSHQSLHTWA 789
Query: 256 WQLHEEEKWLALVDPEMEEFPEKEVIKYIKVAL---FCTQAAARRRPLMTQVVDMLSK 310
+ + + LVDP +E +V ++ +VA C +A++ RP M++V++++ +
Sbjct: 790 KPILSKWEIEKLVDPRLEGC--YDVTQFNRVAFAASLCIRASSTWRPTMSEVLEVMEE 957
>CN825263
Length = 663
Score = 197 bits (501), Expect = 2e-51
Identities = 96/209 (45%), Positives = 138/209 (65%), Gaps = 1/209 (0%)
Frame = +1
Query: 48 GFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKTLSNVKHSNLVELVGFCIQGPNR 107
GFG VY+G L DGR +AVK L ++G REFL E++ LS + H NLV+L+G CI+ R
Sbjct: 1 GFGLVYKGILNDGRDVAVKILKRDDQRGGREFLAEVEMLSRLHHRNLVKLIGICIEKQTR 180
Query: 108 TVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGTAKGLAYLHEELTQHIVHRDIKA 167
++YE V NG++ + L G + + W R I +G A+GLAYLHE+ ++HRD K+
Sbjct: 181 CLIYELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFKS 360
Query: 168 SNVLLDKDFNPKIGDFGMAKLFPDD-ITHISTRIAGTTGYLAPEYALGGQLTKKADVYSF 226
SN+LL+ DF PK+ DFG+A+ D+ HIST + GT GYLAPEYA+ G L K+DVYS+
Sbjct: 361 SNILLECDFTPKVSDFGLARTALDEGNKHISTHVMGTFGYLAPEYAMTGHLLVKSDVYSY 540
Query: 227 GVLILEIISGKSSSRTNWDGSHKSLLEWA 255
GV++LE+++G + ++L+ WA
Sbjct: 541 GVVLLELLTGTKPVDLSQPPGQENLVTWA 627
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 193 bits (491), Expect = 3e-50
Identities = 113/287 (39%), Positives = 170/287 (58%), Gaps = 7/287 (2%)
Frame = +1
Query: 26 FSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKT 85
FS +EL+ AT+N+ L NKIG+GGFG VY L+ G+K A+K + V Q EFL E+K
Sbjct: 1051 FSYQELAKATNNFSLDNKIGQGGFGAVYYAELR-GKKTAIKKMDV---QASTEFLCELKV 1218
Query: 86 LSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGT 145
L++V H NLV L+G+C++G + +VYE+++NGNL L G S + W R I +
Sbjct: 1219 LTHVHHLNLVRLIGYCVEG-SLFLVYEHIDNGNLGQYLHG--SGKEPLPWSSRVQIALDA 1389
Query: 146 AKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHISTRIAGTTG 205
A+GL Y+HE +HRD+K++N+L+DK+ K+ DFG+ KL + + TR+ GT G
Sbjct: 1390 ARGLEYIHEHTVPVYIHRDVKSANILIDKNLRGKVADFGLTKLIEVGNSTLQTRLVGTFG 1569
Query: 206 YLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKS-----LLEWAWQLHE 260
Y+ PEYA G ++ K DVY+FGV++ E+IS K++ + +S L E A +
Sbjct: 1570 YMPPEYAQYGDISPKIDVYAFGVVLFELISAKNAVLKTGELVAESKGLVALFEEALNKSD 1749
Query: 261 E-EKWLALVDPEM-EEFPEKEVIKYIKVALFCTQAAARRRPLMTQVV 305
+ LVDP + E +P V+K ++ CT+ RP M +V
Sbjct: 1750 PCDALRKLVDPRLGENYPIDSVLKIAQLGRACTRDNPLLRPSMRSLV 1890
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 189 bits (481), Expect = 5e-49
Identities = 121/346 (34%), Positives = 174/346 (49%), Gaps = 26/346 (7%)
Frame = +1
Query: 26 FSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKT 85
F +S T+++ NK+G GGFG VY+G L +G++IAVK LS S QG+ EF E+K
Sbjct: 1417 FDFSTISSTTNHFSESNKLGEGGFGPVYKGVLANGQEIAVKRLSNTSGQGMEEFKNEVKL 1596
Query: 86 LSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGT 145
++ ++H NLV+L+G I ++YE++ N
Sbjct: 1597 IARLQHRNLVKLLGCSIHHDEMLLIYEFMHN----------------------------- 1689
Query: 146 AKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHIST-RIAGTT 204
+ L Y + I+HRD+K SN+LLD + NPKI DFG+A++F D T R+ GT
Sbjct: 1690 -RSLDYFIFDSRLRIIHRDLKTSNILLDSEMNPKISDFGLARIFTGDQVEAKTKRVMGTY 1866
Query: 205 GYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEW---------- 254
GY++PEYA+ G + K+DV+SFGV++LEIISGK R H++LL
Sbjct: 1867 GYMSPEYAVHGSFSVKSDVFSFGVIVLEIISGKKIGRFCDPHHHRNLLSHSSNFAVFLIK 2046
Query: 255 --------------AWQLHEEEKWLALVDPEMEEFP-EKEVIKYIKVALFCTQAAARRRP 299
AW+L EE+ L LVD ++ E+++YI +AL C Q RP
Sbjct: 2047 ALRICMFENVKNRKAWRLWIEERPLELVDELLDGLAIPTEILRYIHIALLCVQQRPEYRP 2226
Query: 300 LMTQVVDMLSKEIQLNDKQLTAPGLFNYDAGETSQKKSNPESLVYH 345
M VV ML+ E +L L A N D N E ++ H
Sbjct: 2227 DMLSVVLMLNGEKELPKPSLPAFYTGNDDLLWPESTSKNCERVIKH 2364
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 188 bits (477), Expect = 1e-48
Identities = 111/307 (36%), Positives = 170/307 (55%), Gaps = 8/307 (2%)
Frame = +3
Query: 44 IGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKTLSNVKHSNLVELVGFCIQ 103
+G GGFG VY G + G K+A+K + S+QGV EF TEI+ LS ++H +LV L+G+C +
Sbjct: 15 LGVGGFGKVYYGEVDGGTKVAIKRGNPLSEQGVHEFQTEIEMLSKLRHRHLVSLIGYCEE 194
Query: 104 GPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGTAKGLAYLHEELTQHIVHR 163
+VY+++ G L L K+ + W++R ICIG A+GL YLH I+HR
Sbjct: 195 NTEMILVYDHMAYGTLREHLY--KTQKPPLPWKQRLEICIGAARGLHYLHTGAKYTIIHR 368
Query: 164 DIKASNVLLDKDFNPKIGDFGMAKLFPD-DITHISTRIAGTTGYLAPEYALGGQLTKKAD 222
D+K +N+LLD+ + K+ DFG++K P D TH+ST + G+ GYL PEY QLT K+D
Sbjct: 369 DVKTTNILLDEKWVAKVSDFGLSKTGPTLDNTHVSTVVKGSFGYLDPEYFRRQQLTDKSD 548
Query: 223 VYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLHEEEKWLALVDPEMEEFPEKEVI- 281
VYSFGV++ EI+ + + + SL EWA + + ++DP ++ E
Sbjct: 549 VYSFGVVLFEILCARPALNPSLAKEQVSLAEWASHCYNKGILDQILDPYLKGKIAPECFK 728
Query: 282 KYIKVALFCTQAAARRRPLMTQVVDMLSKEIQLNDKQLTAPGLFNYDAGE------TSQK 335
K+ + A+ C RP M V+ L +QL + + F GE S+
Sbjct: 729 KFAETAMKCVSDQGIERPSMGDVLWNLEFALQLQESAEESGNGFGGIHGEDEPLFADSKG 908
Query: 336 KSNPESL 342
K +P+++
Sbjct: 909 KKDPDAM 929
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 182 bits (462), Expect = 8e-47
Identities = 105/283 (37%), Positives = 164/283 (57%), Gaps = 9/283 (3%)
Frame = +2
Query: 42 NKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVRE--FLTEIKTLSNVKHSNLVELVG 99
N IG+GG G VY+G++ +G +A+K L VG G + F EI+TL ++H N++ L+G
Sbjct: 2204 NIIGKGGAGIVYRGSMPNGTDVAIKRL-VGQGSGRNDYGFRAEIETLGKIRHRNIMRLLG 2380
Query: 100 FCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGTAKGLAYLHEELTQH 159
+ ++YEY+ NG+L L G K ++W R I + A+GL Y+H + +
Sbjct: 2381 YVSNKDTNLLLYEYMPNGSLGEWLHGAKG--GHLRWEMRYKIAVEAARGLCYMHHDCSPL 2554
Query: 160 IVHRDIKASNVLLDKDFNPKIGDFGMAK-LFPDDITHISTRIAGTTGYLAPEYALGGQLT 218
I+HRD+K++N+LLD DF + DFG+AK L+ + + IAG+ GY+APEYA ++
Sbjct: 2555 IIHRDVKSNNILLDADFEAHVADFGLAKFLYDPGASQSMSSIAGSYGYIAPEYAYTLKVD 2734
Query: 219 KKADVYSFGVLILEIISGKSSSRTNWDG------SHKSLLEWAWQLHEEEKWLALVDPEM 272
+K+DVYSFGV++LE+I G+ DG +K++ E + Q + LA+VDP +
Sbjct: 2735 EKSDVYSFGVVLLELIIGRKPVGEFGDGVDIVGWVNKTMSELS-QPSDTALVLAVVDPRL 2911
Query: 273 EEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDMLSKEIQLN 315
+P VI +A+ C + RP M +VV ML+ Q N
Sbjct: 2912 SGYPLTSVIHMFNIAMMCVKEMGPARPTMREVVHMLTNPPQSN 3040
>TC11757 UP|Q91W27 (Q91W27) Tuberoinfundibular 39 residue protein, partial
(10%)
Length = 715
Score = 173 bits (438), Expect = 5e-44
Identities = 83/148 (56%), Positives = 108/148 (72%)
Frame = +2
Query: 22 NIRHFSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLT 81
++R F+ KEL +ATDN+ NKIG GGFG VY+G LKDG+ A+K LS SKQGV+EF+T
Sbjct: 272 HVRVFTYKELKIATDNFSPVNKIGEGGFGPVYKGVLKDGKVGAIKVLSAESKQGVKEFMT 451
Query: 82 EIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTI 141
EI +S ++H NLV+L G C++G +R +VY Y EN +L LLG ++ WR RS I
Sbjct: 452 EINVISEIEHENLVQLYGCCVEGNHRILVYNYHENNSLAQTLLGGGHSNIHFDWRTRSRI 631
Query: 142 CIGTAKGLAYLHEELTQHIVHRDIKASN 169
CIG A+GL++LHEE+ HIVHRDIKASN
Sbjct: 632 CIGVARGLSFLHEEVRPHIVHRDIKASN 715
>TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine protein kinase
Cdk9 (Cyclin-dependent kinase Cdk9) , partial (7%)
Length = 723
Score = 170 bits (430), Expect = 4e-43
Identities = 84/173 (48%), Positives = 116/173 (66%)
Frame = +3
Query: 26 FSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKT 85
FS + L AT N+H NK+G GGFG V++G L DGR+IAVK LS S QG +F+ E K
Sbjct: 207 FSYETLVAATKNFHAVNKLGEGGFGPVFKGKLNDGREIAVKKLSRRSNQGRTQFINEAKL 386
Query: 86 LSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGT 145
L+ V+H N+V L G+C G + +VYEYV +L LL + ++ W+ R I G
Sbjct: 387 LTRVQHRNVVSLFGYCAHGSEKLLVYEYVPRESL-DKLLFRSQKKEQLDWKRRFDIISGV 563
Query: 146 AKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFPDDITHIST 198
A+GL YLHE+ I+HRDIKA+N+LLD+ + PKI DFG+A++FP+D TH++T
Sbjct: 564 ARGLLYLHEDSHDCIIHRDIKAANILLDEKWVPKIADFGLARIFPEDQTHVNT 722
>AV422094
Length = 462
Score = 163 bits (413), Expect = 4e-41
Identities = 81/150 (54%), Positives = 106/150 (70%)
Frame = +2
Query: 26 FSDKELSLATDNYHLGNKIGRGGFGTVYQGTLKDGRKIAVKPLSVGSKQGVREFLTEIKT 85
FS EL AT+++++ NK+G GGFG VY+G L DG IAVK LS+GS QG +F+ EI T
Sbjct: 14 FSYSELKNATNDFNIDNKLGEGGFGPVYKGILNDGTVIAVKQLSLGSHQGKSQFIAEIAT 193
Query: 86 LSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTICIGT 145
+S V+H NLV+L G CI+G R +VYEY+EN +L AL GK ++ + W R IC+G
Sbjct: 194 ISAVQHRNLVKLYGCCIEGSKRLLVYEYLENKSLDQALFGK---ALSLNWSTRYDICLGV 364
Query: 146 AKGLAYLHEELTQHIVHRDIKASNVLLDKD 175
A+GL YLHEE IVHRD+KASN+LLD +
Sbjct: 365 ARGLTYLHEESRLRIVHRDVKASNILLDHE 454
>TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P14.15
{Arabidopsis thaliana;}, partial (37%)
Length = 1079
Score = 162 bits (411), Expect = 6e-41
Identities = 91/230 (39%), Positives = 132/230 (56%), Gaps = 3/230 (1%)
Frame = +3
Query: 82 EIKTLSNVKHSNLVELVGFCIQGPNRTVVYEYVENGNLHTALLGKKSLSVKMKWRERSTI 141
E++ L+ + H NLV+L+GF +G R ++ EYV NG L L G + + + +R I
Sbjct: 6 EVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKI--LDFNQRLEI 179
Query: 142 CIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLFP--DDITHISTR 199
I A GL YLH + I+HRD+K+SN+LL + K+ DFG A+L P D THIST+
Sbjct: 180 AIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTK 359
Query: 200 IAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQLH 259
+ GT GYL PEY QLT K+DVYSFG+L+LEI++G+ + L WA++ +
Sbjct: 360 VKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKY 539
Query: 260 EEEKWLALVDPEMEEFPEKEVI-KYIKVALFCTQAAARRRPLMTQVVDML 308
E + L+DP MEE +V+ K + ++ C RP M V + L
Sbjct: 540 NEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQL 689
>TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1, complete
Length = 2061
Score = 155 bits (392), Expect = 1e-38
Identities = 87/199 (43%), Positives = 118/199 (58%), Gaps = 2/199 (1%)
Frame = +1
Query: 125 GKKSLSVKMKWRERSTICIGTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFG 184
G + S + W +R I G A+GL YLH++ I+HRD+K SN+LLD + NPKI DFG
Sbjct: 1375 GDSTRSKLLDWNKRLQIIDGIARGLLYLHQDSRLRIIHRDLKTSNILLDNEMNPKISDFG 1554
Query: 185 MAKLFPDDITHIST-RIAGTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTN 243
+A++F D T R+ GT GY+ PEYA+ G + K+DV+SFGV++LEIISGK +
Sbjct: 1555 LARIFIGDQVEARTKRVMGTYGYMPPEYAVHGSFSIKSDVFSFGVIVLEIISGKKIRKFY 1734
Query: 244 WDGSHKSLLEWAWQLHEEEKWLALVDPEMEE-FPEKEVIKYIKVALFCTQAAARRRPLMT 302
H +LL AW+L E L LVD E+ E+++YI VAL C Q RP M
Sbjct: 1735 DPHHHLNLLSHAWRLWIEGSPLELVDKLFEDSIIPTEILRYIHVALLCVQRRPETRPDML 1914
Query: 303 QVVDMLSKEIQLNDKQLTA 321
+V ML+ E +L L A
Sbjct: 1915 SIVLMLNGEKELPKPSLPA 1971
>TC9538 similar to UP|RLK5_ARATH (P47735) Receptor-like protein kinase 5
precursor , partial (10%)
Length = 1067
Score = 146 bits (369), Expect = 5e-36
Identities = 89/229 (38%), Positives = 130/229 (55%), Gaps = 13/229 (5%)
Frame = +2
Query: 93 NLVELVGFCIQGPNRTVVYEYVENGNLHTAL-LGKKSLSVK--------MKWRERSTICI 143
N+V L+ + +VYEY+EN +L L L KS SV + W +R I I
Sbjct: 5 NIVRLLCCISNEASMLLVYEYLENHSLDKWLHLKPKSSSVSGVVQQYTVLDWPKRLKIAI 184
Query: 144 GTAKGLAYLHEELTQHIVHRDIKASNVLLDKDFNPKIGDFGMAKLF--PDDITHISTRIA 201
G A+GL+Y+H + + IVHRD+K SN+LLDK FN K+ DFG+A++ P ++ +ST +
Sbjct: 185 GAAQGLSYMHHDCSPPIVHRDVKTSNILLDKQFNAKVADFGLARMLIKPGELNIMST-VI 361
Query: 202 GTTGYLAPEYALGGQLTKKADVYSFGVLILEIISGKSSSRTNWDGSHKSLLEWAWQ--LH 259
GT GY+APEY ++++K DVYSFGV++LE+ +GK + N+ H SL EWAW+ L
Sbjct: 362 GTFGYIAPEYVQTTRISEKVDVYSFGVVLLELTTGKEA---NYGDQHSSLAEWAWRHILI 532
Query: 260 EEEKWLALVDPEMEEFPEKEVIKYIKVALFCTQAAARRRPLMTQVVDML 308
L ME E+ K+ + CT RP M +V+ +L
Sbjct: 533 GSNVXDLLXKDVMEASYIDEMCSVFKLGVMCTATLPATRPSMKEVLPIL 679
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,146,414
Number of Sequences: 28460
Number of extensions: 82618
Number of successful extensions: 863
Number of sequences better than 10.0: 324
Number of HSP's better than 10.0 without gapping: 688
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 692
length of query: 359
length of database: 4,897,600
effective HSP length: 91
effective length of query: 268
effective length of database: 2,307,740
effective search space: 618474320
effective search space used: 618474320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC129091.7


