

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC129091.5 - phase: 0 /pseudo
(980 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
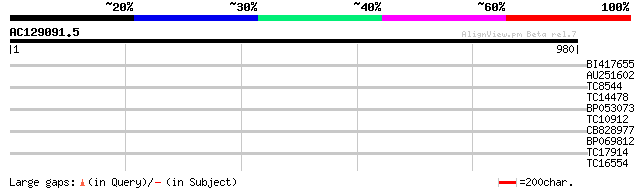
Score E
Sequences producing significant alignments: (bits) Value
BI417655 32 0.37
AU251602 31 0.83
TC8544 31 1.1
TC14478 weakly similar to UP|Q94G86 (Q94G86) Beta-1,3-glucanase-... 31 1.1
BP053073 30 2.4
TC10912 30 2.4
CB828977 28 5.4
BP069812 28 5.4
TC17914 similar to UP|Q39833 (Q39833) ALFA-carboxyltransferase p... 28 9.2
TC16554 homologue to UP|AAN71719 (AAN71719) Glycogen synthase ki... 28 9.2
>BI417655
Length = 509
Score = 32.3 bits (72), Expect = 0.37
Identities = 18/46 (39%), Positives = 27/46 (58%)
Frame = +3
Query: 279 RQVLQLQHLLAHPTGLVLPLYLPFCSVGKINLYHFTAQQFQIFQHM 324
+Q+LQ+QHLL H L L FC VG+ +HF + + + QH+
Sbjct: 351 QQLLQMQHLLGHCI-----LVLMFCFVGQDLCHHFHSHKQRQQQHL 473
>AU251602
Length = 350
Score = 31.2 bits (69), Expect = 0.83
Identities = 16/39 (41%), Positives = 21/39 (53%)
Frame = -2
Query: 275 NFLPRQVLQLQHLLAHPTGLVLPLYLPFCSVGKINLYHF 313
NF P+ LQ H+LA P L +L FC V K + Y +
Sbjct: 277 NFFPKLQLQFNHILAFPCQLA-SFFLFFCFVAKRSCYSY 164
>TC8544
Length = 1278
Score = 30.8 bits (68), Expect = 1.1
Identities = 18/46 (39%), Positives = 25/46 (54%)
Frame = +2
Query: 2 NPMKESEAEAEAAATVNDPMMEEEEVEAGTVFDISLEQSPSTSLHK 47
+P+KE+E A AAA + E EEVE G FD +E ++ K
Sbjct: 401 DPVKETEDSASAAAASS----ESEEVEVGVGFDSKVEGEEVKAIEK 526
>TC14478 weakly similar to UP|Q94G86 (Q94G86) Beta-1,3-glucanase-like
protein, partial (83%)
Length = 1784
Score = 30.8 bits (68), Expect = 1.1
Identities = 19/68 (27%), Positives = 34/68 (49%), Gaps = 3/68 (4%)
Frame = +1
Query: 651 LASHLITRLRHYA---KIFRTLANQAVTVASGSTSGGVPNLTLNPSISGPSSLMLISINT 707
L S I ++R Y +I + LAN + + G+ +G +PNL +P+ + + +N+
Sbjct: 208 LRSTSIGKIRLYGADPEIIKALANSGIGIIIGAANGDIPNLAADPNAA------VQWVNS 369
Query: 708 GTFPGTPA 715
P PA
Sbjct: 370 NVLPYYPA 393
>BP053073
Length = 456
Score = 29.6 bits (65), Expect = 2.4
Identities = 13/35 (37%), Positives = 18/35 (51%)
Frame = -1
Query: 58 FSAVAWCAKLNVIACATETCVNGIPRSSVNPWFWI 92
F V W A L V CAT+ CV+ S++ W+
Sbjct: 411 FXHVVWFAALMVQGCATDACVHRFSAGSLSCSAWV 307
>TC10912
Length = 781
Score = 29.6 bits (65), Expect = 2.4
Identities = 12/28 (42%), Positives = 15/28 (52%)
Frame = +2
Query: 896 RPCTDSNDTPKPFRSNPLDSRSLESNDI 923
RP + P PF NP+DS +L DI
Sbjct: 152 RPIKPNRSEPDPFAQNPVDSSTLSDQDI 235
>CB828977
Length = 528
Score = 28.5 bits (62), Expect = 5.4
Identities = 16/41 (39%), Positives = 26/41 (63%)
Frame = +2
Query: 664 KIFRTLANQAVTVASGSTSGGVPNLTLNPSISGPSSLMLIS 704
K T+A+++ ASG TS +PN +L PS + P+S+ I+
Sbjct: 89 KTLSTVASRSTAQASGRTSSKIPN-SLLPSRAVPTSISRIN 208
>BP069812
Length = 433
Score = 28.5 bits (62), Expect = 5.4
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = +2
Query: 384 DISIPATDDKSKGIHFNPFDLPKYL 408
++ +P + K KG F P+ PKYL
Sbjct: 71 NLKLPPSSSKKKGQSFKPYQTPKYL 145
>TC17914 similar to UP|Q39833 (Q39833) ALFA-carboxyltransferase precursor
(Carboxyl transferase alpha subunit) , partial (23%)
Length = 530
Score = 27.7 bits (60), Expect = 9.2
Identities = 19/54 (35%), Positives = 24/54 (44%), Gaps = 9/54 (16%)
Frame = -3
Query: 775 GSKGGEEPSPGPVRLGNGNAGQGYS---------VEEVKVIFQVLLDLCRRTSG 819
G KG S P LG+G +GQGYS E ++F LDL + G
Sbjct: 288 GLKGDR*LSTPPFTLGSGLSGQGYSCFLTFLSLATTEKSLVFMANLDLPKVLKG 127
>TC16554 homologue to UP|AAN71719 (AAN71719) Glycogen synthase kinase 3 beta
protein kinase DWARF12 , partial (71%)
Length = 927
Score = 27.7 bits (60), Expect = 9.2
Identities = 23/68 (33%), Positives = 27/68 (38%), Gaps = 10/68 (14%)
Frame = +1
Query: 723 HFLHRLC-QLLFFCFFFKRSQL-------ARYKSGLRRTAE--TSLVRSNDGQTGRDRQI 772
HFLH +LFF FFF L S + E TS++ ND TG
Sbjct: 16 HFLHIFSFSILFFFFFFTLP*LLPLLLLSLSLSSSMAEDKEMSTSVINGNDSLTGHIIST 195
Query: 773 VPGSKGGE 780
G K GE
Sbjct: 196 TIGGKNGE 219
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,590,889
Number of Sequences: 28460
Number of extensions: 286287
Number of successful extensions: 2038
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 2030
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2038
length of query: 980
length of database: 4,897,600
effective HSP length: 99
effective length of query: 881
effective length of database: 2,080,060
effective search space: 1832532860
effective search space used: 1832532860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC129091.5


