

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC129091.3 + phase: 0
(170 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
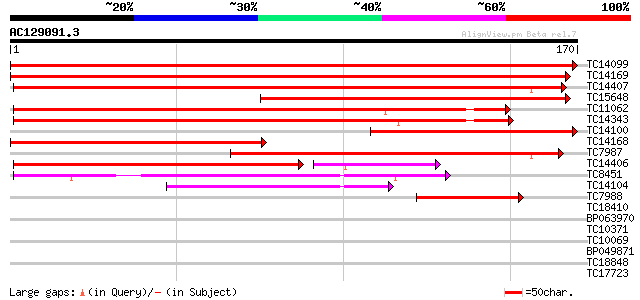
Score E
Sequences producing significant alignments: (bits) Value
TC14099 326 1e-90
TC14169 267 6e-73
TC14407 similar to UP|Q8S2R9 (Q8S2R9) Aluminum-induced protein-l... 199 2e-52
TC15648 150 8e-38
TC11062 similar to UP|O64438 (O64438) ARG10, complete 142 2e-35
TC14343 similar to UP|O64438 (O64438) ARG10, complete 141 7e-35
TC14100 similar to UP|Q9LE80 (Q9LE80) Genomic DNA, chromosome 3,... 130 2e-31
TC14168 120 1e-28
TC7987 similar to UP|Q8S2R9 (Q8S2R9) Aluminum-induced protein-li... 118 5e-28
TC14406 similar to GB|AAM61587.1|21537246|AY085029 aluminum-indu... 101 3e-24
TC8451 GB|CAA61589.1|897771|LJAS1GENE asparagine synthase (gluta... 47 1e-06
TC14104 GB|CAA61590.1|897773|LJAS2GENE asparagine synthase (glut... 44 2e-05
TC7988 42 6e-05
TC18410 weakly similar to UP|O81226 (O81226) Glutamine cyclotran... 26 4.1
BP063970 25 7.0
TC10371 25 7.0
TC10069 weakly similar to UP|O23427 (O23427) DNA chromosome 4, E... 25 7.0
BP049871 25 7.0
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 25 9.2
TC17723 25 9.2
>TC14099
Length = 1075
Score = 326 bits (835), Expect = 1e-90
Identities = 153/170 (90%), Positives = 164/170 (96%)
Frame = +1
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
+G+LHNLSQLNKQYGLSKG NEAMFIIEAYRTLRDRGPYPADQVLK LEGSF FVIYD+K
Sbjct: 334 LGSLHNLSQLNKQYGLSKGTNEAMFIIEAYRTLRDRGPYPADQVLKELEGSFGFVIYDNK 513
Query: 61 DGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLN 120
DGTVF ASGSDG +GL+WGIAADGS+VISENLELVK+SCAKSFAPFPTGCLFHSEHGLL+
Sbjct: 514 DGTVFVASGSDGQVGLFWGIAADGSIVISENLELVKSSCAKSFAPFPTGCLFHSEHGLLS 693
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATWGS 170
F+HPTKKMKAMPRIDSEG+MCGANFNVDSQS+NQMMPRVGSEANWA WGS
Sbjct: 694 FQHPTKKMKAMPRIDSEGIMCGANFNVDSQSKNQMMPRVGSEANWAIWGS 843
>TC14169
Length = 964
Score = 267 bits (683), Expect = 6e-73
Identities = 122/168 (72%), Positives = 146/168 (86%)
Frame = +1
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
+G L+NL LNKQYGL+KG +EAMF+IEAY+TLRDRGPYPADQV+KGL+GSFAFV+YD K
Sbjct: 298 LGTLNNLCSLNKQYGLTKGTDEAMFVIEAYKTLRDRGPYPADQVVKGLDGSFAFVVYDSK 477
Query: 61 DGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLN 120
GTVFAA GSDG + LYWGIAADGSVVIS++L+++K CAKSFAPFPTGC+FHSE GL++
Sbjct: 478 VGTVFAALGSDGGVKLYWGIAADGSVVISDDLDVIKEGCAKSFAPFPTGCMFHSEGGLMS 657
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATW 168
FEHP K+KAMPR+DSEGVMCGANF VD +R +PRVGS+A+W W
Sbjct: 658 FEHPMNKLKAMPRVDSEGVMCGANFKVDKYTRVNSIPRVGSQAHWMEW 801
>TC14407 similar to UP|Q8S2R9 (Q8S2R9) Aluminum-induced protein-like
protein, partial (97%)
Length = 1006
Score = 199 bits (506), Expect = 2e-52
Identities = 88/168 (52%), Positives = 122/168 (72%), Gaps = 2/168 (1%)
Frame = +3
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G++ N++ L +QYGL+K NE + +IEAYRTLRDRGPYPADQV++ +G FAF+++D
Sbjct: 318 GHIDNVAHLKQQYGLNKTANEVIIVIEAYRTLRDRGPYPADQVVRDFQGKFAFILFDSSS 497
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLNF 121
T F AS SDG + +WG ADG++V+S+ +V SC KSFAPFP GC F + GL NF
Sbjct: 498 KTAFVASDSDGSVPFFWGTDADGNLVLSDETGIVSKSCGKSFAPFPKGCFFTTSGGLRNF 677
Query: 122 EHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQM--MPRVGSEANWAT 167
EHP ++K +PR+DS G +CGA F VD++++ + MPRVGS ANW++
Sbjct: 678 EHPLNELKPVPRVDSSGQVCGATFKVDAETKKETTGMPRVGSAANWSS 821
>TC15648
Length = 536
Score = 150 bits (380), Expect = 8e-38
Identities = 66/93 (70%), Positives = 79/93 (83%)
Frame = +2
Query: 76 LYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLNFEHPTKKMKAMPRID 135
LYWGIAADGSVVIS++L+++K CAKSFAPFPTGC+FHSE GL++FEHP K+KAMPRID
Sbjct: 5 LYWGIAADGSVVISDDLKVIKEGCAKSFAPFPTGCIFHSEGGLVSFEHPMHKLKAMPRID 184
Query: 136 SEGVMCGANFNVDSQSRNQMMPRVGSEANWATW 168
SEG MCGANF VD +R +PRVGS++NW W
Sbjct: 185 SEGAMCGANFKVDKYARVNSIPRVGSQSNWMEW 283
>TC11062 similar to UP|O64438 (O64438) ARG10, complete
Length = 1032
Score = 142 bits (359), Expect = 2e-35
Identities = 70/150 (46%), Positives = 98/150 (64%), Gaps = 1/150 (0%)
Frame = +2
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G L NL L +QYGL+K NE + +IEAY+ LRDR PYPA+ V+ L GSFAF+++D
Sbjct: 332 GALDNLGSLRQQYGLAKSANEVLLVIEAYKALRDRAPYPANHVVGHLSGSFAFIVFDKST 511
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCL-FHSEHGLLN 120
T+F AS G + LYWGI ADG V +++ EL+K++C KS A FP GC F + GL+
Sbjct: 512 STLFVASDQFGKVPLYWGITADGYVAFADDAELLKSACGKSLASFPQGCFYFTAVGGLMC 691
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVDSQ 150
+E+P K+ A+P + E + GA F ++ Q
Sbjct: 692 YENPKSKITAVPCHEEE--IWGATFKLEGQ 775
>TC14343 similar to UP|O64438 (O64438) ARG10, complete
Length = 1242
Score = 141 bits (355), Expect = 7e-35
Identities = 68/151 (45%), Positives = 98/151 (64%), Gaps = 1/151 (0%)
Frame = +3
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G L NL L +QYGL+K NEA+ +IEAYR LRDR PYPA+ V+ L G+FAF+++D
Sbjct: 309 GALDNLGSLRQQYGLAKSANEAVLLIEAYRALRDRAPYPANHVVGHLSGNFAFIVFDKST 488
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSE-HGLLN 120
T+F AS G + LYWGI AD V +++++L+K +C KS A FP GC + ++ GL
Sbjct: 489 STLFVASDQTGKVPLYWGITADACVAFADDVDLLKGACGKSLASFPQGCFYSTDVGGLKC 668
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVDSQS 151
+E+P K+ A+P + E + GA F V+ +
Sbjct: 669 YENPKNKITAVPAEEEE--IWGATFKVEGSA 755
>TC14100 similar to UP|Q9LE80 (Q9LE80) Genomic DNA, chromosome 3, P1 clone:
MJK13 (MJK13.11 protein) (AT3g15450/MJK13_11), partial
(23%)
Length = 652
Score = 130 bits (326), Expect = 2e-31
Identities = 57/62 (91%), Positives = 61/62 (97%)
Frame = +1
Query: 109 GCLFHSEHGLLNFEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWATW 168
GCLFHSEHGLL+F+HPTKKMKAMPRIDSEG+MCGANFNVDSQS+NQMMPRVGSEANWA W
Sbjct: 181 GCLFHSEHGLLSFQHPTKKMKAMPRIDSEGIMCGANFNVDSQSKNQMMPRVGSEANWAIW 360
Query: 169 GS 170
GS
Sbjct: 361 GS 366
>TC14168
Length = 573
Score = 120 bits (301), Expect = 1e-28
Identities = 58/77 (75%), Positives = 68/77 (87%)
Frame = +2
Query: 1 MGNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHK 60
+G+L+NLS LNKQYGL+K +EAMF+IEAYRTLRDRGPYPADQV+K L+GSFAFV+YD K
Sbjct: 341 LGSLNNLSVLNKQYGLTKCTDEAMFVIEAYRTLRDRGPYPADQVVKDLDGSFAFVVYDSK 520
Query: 61 DGTVFAASGSDGHIGLY 77
GTVFAA GSDG + LY
Sbjct: 521 VGTVFAALGSDGGVKLY 571
>TC7987 similar to UP|Q8S2R9 (Q8S2R9) Aluminum-induced protein-like
protein, partial (40%)
Length = 606
Score = 118 bits (296), Expect = 5e-28
Identities = 53/102 (51%), Positives = 71/102 (68%), Gaps = 2/102 (1%)
Frame = +2
Query: 67 ASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLNFEHPTK 126
+S DG + +WG ADG++V+S+ E+V SC KS APFP GC F + GL +FEHP
Sbjct: 2 SSDDDGSVPFFWGTDADGNLVLSDETEIVAKSCGKSSAPFPKGCFFSTSGGLSSFEHPLN 181
Query: 127 KMKAMPRIDSEGVMCGANFNVDSQSRNQM--MPRVGSEANWA 166
+MKA+PR+DS G MCGA F VD+ ++ + MPRVGS ANW+
Sbjct: 182 EMKAVPRVDSSGEMCGATFKVDADAKKETTGMPRVGSAANWS 307
>TC14406 similar to GB|AAM61587.1|21537246|AY085029 aluminum-induced
protein-like {Arabidopsis thaliana;}, partial (66%)
Length = 806
Score = 101 bits (251), Expect(2) = 3e-24
Identities = 43/87 (49%), Positives = 63/87 (71%)
Frame = +3
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G++ N++ L +QYGL+K NE + +IEAYRTLRDRGPYPA QV++ +G F FV++D
Sbjct: 318 GHIQNVAHLKQQYGLNKTANEVIIVIEAYRTLRDRGPYPAAQVVRDFQGKFTFVLFDSGS 497
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVI 88
T F +S +DG + +WG ADG++V+
Sbjct: 498 KTAFISSDADGXVPFFWGTDADGNLVL 578
Score = 25.4 bits (54), Expect(2) = 3e-24
Identities = 15/39 (38%), Positives = 16/39 (40%), Gaps = 1/39 (2%)
Frame = +2
Query: 92 LELVKASC-AKSFAPFPTGCLFHSEHGLLNFEHPTKKMK 129
L L SC APFP + L FEHP K K
Sbjct: 590 LXLXPKSCWGNPSAPFPKRMFLFTSGSLNXFEHPXMKXK 706
>TC8451 GB|CAA61589.1|897771|LJAS1GENE asparagine synthase
(glutamine-hydrolysing) {Lotus corniculatus var.
japonicus;} , complete
Length = 2148
Score = 47.4 bits (111), Expect = 1e-06
Identities = 38/134 (28%), Positives = 61/134 (45%), Gaps = 3/134 (2%)
Frame = +2
Query: 2 GNLHNLSQLNKQYGLS--KGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDH 59
G ++N +L KQ + G++ I Y + + L+G F+FV+ D
Sbjct: 326 GEIYNHEELRKQLPNHQFRTGSDCDVIAHLYEE-------HGENFMDMLDGIFSFVLLDT 484
Query: 60 KDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHS-EHGL 118
+D T A + G LY G DGSV IS ++ + C + F FP G L+ S E
Sbjct: 485 RDNTFIVARDAIGVTSLYIGWGLDGSVWISSEMKGLNDDC-EHFEVFPPGHLYSSRERAF 661
Query: 119 LNFEHPTKKMKAMP 132
+ +PT +++P
Sbjct: 662 RRWYNPTWFSESIP 703
>TC14104 GB|CAA61590.1|897773|LJAS2GENE asparagine synthase
(glutamine-hydrolysing) {Lotus corniculatus var.
japonicus;} , complete
Length = 2259
Score = 43.5 bits (101), Expect = 2e-05
Identities = 25/68 (36%), Positives = 37/68 (53%)
Frame = +3
Query: 48 LEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFP 107
L+G F+FV+ D +D + A + G LY G DGSV I+ L+ + C + F FP
Sbjct: 501 LDGIFSFVLLDTRDNSFLVARDAIGVTSLYIGYGLDGSVWIASELKGLNDDC-EHFELFP 677
Query: 108 TGCLFHSE 115
G L+ S+
Sbjct: 678 PGHLYSSK 701
>TC7988
Length = 420
Score = 42.0 bits (97), Expect = 6e-05
Identities = 16/32 (50%), Positives = 23/32 (71%)
Frame = +2
Query: 123 HPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQ 154
HP +MKA+PR+D G MCGA F VD+ ++ +
Sbjct: 2 HPLHEMKAVPRVDRSGEMCGAPFQVDADAKKE 97
>TC18410 weakly similar to UP|O81226 (O81226) Glutamine cyclotransferase
precursor , partial (62%)
Length = 725
Score = 25.8 bits (55), Expect = 4.1
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = +1
Query: 56 IYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISEN 91
IYDHK+ + D G WG+A+DG V+ +
Sbjct: 415 IYDHKNLSQLGTFHHDMKDG--WGLASDGKVLFGSD 516
>BP063970
Length = 397
Score = 25.0 bits (53), Expect = 7.0
Identities = 10/20 (50%), Positives = 13/20 (65%)
Frame = -1
Query: 5 HNLSQLNKQYGLSKGGNEAM 24
H S L + YGL+ GGN A+
Sbjct: 229 HQKSNLERAYGLNTGGNLAL 170
>TC10371
Length = 823
Score = 25.0 bits (53), Expect = 7.0
Identities = 15/31 (48%), Positives = 19/31 (60%), Gaps = 3/31 (9%)
Frame = -1
Query: 88 ISENLEL--VKASC-AKSFAPFPTGCLFHSE 115
+ E L+L V SC AKSF F + C FHS+
Sbjct: 244 VPEYLKLTHVPTSCHAKSFLDFASNCHFHSK 152
>TC10069 weakly similar to UP|O23427 (O23427) DNA chromosome 4, ESSA I
CONTIG fragment NO. 4, partial (9%)
Length = 565
Score = 25.0 bits (53), Expect = 7.0
Identities = 12/35 (34%), Positives = 19/35 (54%)
Frame = +2
Query: 91 NLELVKASCAKSFAPFPTGCLFHSEHGLLNFEHPT 125
N L++ S +S FPT + ++ +LNF H T
Sbjct: 2 NTSLIRQSLMESMKGFPTHAMDMIQNCMLNFLHLT 106
>BP049871
Length = 472
Score = 25.0 bits (53), Expect = 7.0
Identities = 11/34 (32%), Positives = 19/34 (55%)
Frame = -1
Query: 94 LVKASCAKSFAPFPTGCLFHSEHGLLNFEHPTKK 127
L+ A+C ++ P G + +G+LN PT+K
Sbjct: 403 LIAAACERTVLPVSNGHAKLTPNGVLNGHSPTQK 302
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 24.6 bits (52), Expect = 9.2
Identities = 10/36 (27%), Positives = 17/36 (46%)
Frame = +2
Query: 69 GSDGHIGLYWGIAADGSVVISENLELVKASCAKSFA 104
G G +YWG +DG + + L+ + + FA
Sbjct: 323 GEGGFGSVYWGRTSDGLQIAVKKLKAMNSKAEMEFA 430
>TC17723
Length = 842
Score = 24.6 bits (52), Expect = 9.2
Identities = 9/23 (39%), Positives = 13/23 (56%)
Frame = -2
Query: 148 DSQSRNQMMPRVGSEANWATWGS 170
DS + +P+V + NW WGS
Sbjct: 739 DSLVNSTFIPKVPNLVNWGYWGS 671
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,833,411
Number of Sequences: 28460
Number of extensions: 33322
Number of successful extensions: 143
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 141
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 141
length of query: 170
length of database: 4,897,600
effective HSP length: 84
effective length of query: 86
effective length of database: 2,506,960
effective search space: 215598560
effective search space used: 215598560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC129091.3


