

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127429.3 - phase: 0 /pseudo
(423 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
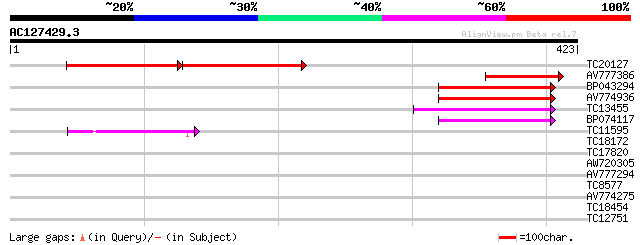
TC20127 102 2e-42
AV777386 102 1e-22
BP043294 94 3e-20
AV774936 82 2e-16
TC13455 weakly similar to UP|Q9LG41 (Q9LG41) Dimethylaniline mon... 78 3e-15
BP074117 75 2e-14
TC11595 weakly similar to UP|Q9FKE7 (Q9FKE7) Dimethylaniline mon... 47 5e-06
TC18172 similar to UP|O81815 (O81815) Monooxygenase, partial (15%) 33 0.12
TC17820 32 0.26
AW720305 30 0.75
AV777294 29 1.3
TC8577 similar to UP|Q9XE94 (Q9XE94) Geranylgeranyl hydrogenase,... 29 1.7
AV774275 27 4.9
TC18454 similar to UP|Q9XEY7 (Q9XEY7) Trehalase 1 GMTRE1, partia... 27 4.9
TC12751 similar to UP|LEU3_BRANA (P29102) 3-isopropylmalate dehy... 27 8.3
>TC20127
Length = 582
Score = 102 bits (254), Expect(2) = 2e-42
Identities = 47/87 (54%), Positives = 63/87 (72%)
Frame = +3
Query: 43 LIVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVCELPLMEF 102
+IVG G SG+A A+ L +K + ++LER +C ASLW+ TYDRL LHL KQ CELP F
Sbjct: 39 IIVGGGTSGIATASCLTKKSISYIMLEREDCFASLWQKYTYDRLHLHLRKQSCELPHFPF 218
Query: 103 PSGFPTYPTKQQFIEYLESYSKNFDIR 129
P +P Y K+QFIEYL++Y K+F+I+
Sbjct: 219 PPSYPHYVPKKQFIEYLDNYVKHFNIK 299
Score = 87.4 bits (215), Expect(2) = 2e-42
Identities = 42/93 (45%), Positives = 66/93 (70%), Gaps = 1/93 (1%)
Frame = +1
Query: 130 PWFNETVMHAEFDATLGFWRVRSEGK-AGMVTEFVCRWLIVATGENAEAVVPEIEGVDEF 188
P ++ V AE D + WRV+++ + +G V E+ ++L+VATGE AE +PE+EG++ F
Sbjct: 301 PLYHRAVELAEHDNSHQNWRVKAKNRTSGHVEEYAGKFLVVATGETAEPRIPEVEGLEGF 480
Query: 189 VGSIRHTSLYKSGEEFRGKKVLVVGCGNSGMEV 221
G + H++ YK+G+EF+ + VLVVG GNSGME+
Sbjct: 481 KGKVIHSTGYKNGKEFKNQNVLVVGSGNSGMEI 579
>AV777386
Length = 582
Score = 102 bits (254), Expect = 1e-22
Identities = 45/58 (77%), Positives = 51/58 (87%)
Frame = -3
Query: 356 LKDKGMFSKEDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAKNIAEDIERCWKA 413
++D MFS++DG PRKPFPNGWKG NGLYA+GFTKRGLLGAS+DAK IAEDIE WKA
Sbjct: 463 IQDNEMFSEKDGLPRKPFPNGWKGSNGLYAIGFTKRGLLGASIDAKRIAEDIEHSWKA 290
>BP043294
Length = 527
Score = 94.4 bits (233), Expect = 3e-20
Identities = 43/87 (49%), Positives = 57/87 (65%)
Frame = -3
Query: 321 IKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWKGE 380
I+ ++ V F DG ++ FD+II TG+K + WLK F EDG+P+ PN WKG
Sbjct: 423 IESIRGNQVLFRDGKSQPFDSIIFCTGFKRSTKKWLKGGDDFLNEDGFPKPGLPNHWKGN 244
Query: 381 NGLYAVGFTKRGLLGASMDAKNIAEDI 407
NGLY VG ++RG GA+MDA+NIA DI
Sbjct: 243 NGLYCVGLSRRGFFGANMDAQNIANDI 163
>AV774936
Length = 477
Score = 81.6 bits (200), Expect = 2e-16
Identities = 37/87 (42%), Positives = 53/87 (60%)
Frame = -3
Query: 321 IKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWKGE 380
I+ ++ + F DG ++ FD+II TG++ + WLK F EDG+P+ PN WKG
Sbjct: 352 IESIRGNPMLFRDGQSQPFDSIIFCTGFQRSTKKWLKGGDDFLYEDGFPKPGLPNPWKGN 173
Query: 381 NGLYAVGFTKRGLLGASMDAKNIAEDI 407
GLY VG ++RG GA +DA +A DI
Sbjct: 172 YGLYCVGLSRRGFFGAKLDAPALAHDI 92
>TC13455 weakly similar to UP|Q9LG41 (Q9LG41) Dimethylaniline
monooxygenase-like protein, partial (9%)
Length = 535
Score = 77.8 bits (190), Expect = 3e-15
Identities = 41/106 (38%), Positives = 56/106 (52%)
Frame = +1
Query: 302 DTGPRCGCPCQD*KRRHQGIKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGM 361
DTG D K I+ ++ V +G FD+II TG+K + WLK
Sbjct: 73 DTGTIHKIKSGDLKVLPSEIEYVRDKNVLLKNGELHPFDSIIFCTGFKRSTHKWLKGDDY 252
Query: 362 FSKEDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAKNIAEDI 407
+DG P+ +P WKG+ GLY VG ++RGL GA+ D +NIA DI
Sbjct: 253 LLNDDGLPKPSYPMYWKGKKGLYCVGLSRRGLYGAAADGENIANDI 390
>BP074117
Length = 395
Score = 75.1 bits (183), Expect = 2e-14
Identities = 35/87 (40%), Positives = 49/87 (56%)
Frame = -3
Query: 321 IKRLKRYTVEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKEDGYPRKPFPNGWKGE 380
I+ ++ V F DG + FD+II TG+K + WLK E+G + F + WKGE
Sbjct: 387 IESVRGNQVLFGDGKSHTFDSIIFCTGFKRSTQKWLKGGDDLLNENGLTKTNFQSNWKGE 208
Query: 381 NGLYAVGFTKRGLLGASMDAKNIAEDI 407
NGLY G ++ G G +A+NIA DI
Sbjct: 207 NGLYCAGLSRMGFFGVKDEAQNIANDI 127
>TC11595 weakly similar to UP|Q9FKE7 (Q9FKE7) Dimethylaniline monooxygenase
(N-oxide-forming)-like protein, partial (13%)
Length = 435
Score = 47.4 bits (111), Expect = 5e-06
Identities = 30/101 (29%), Positives = 50/101 (48%), Gaps = 3/101 (2%)
Frame = +2
Query: 44 IVGAGPSGLAVAAYLKQKGVPSLILERSNCIASLWKLKTYDRLRLHLPKQVCELPLMEFP 103
I+GAG SG+A A L ++ E ++ I +W+ Y+ +L E +P
Sbjct: 110 IIGAGVSGIAAAKQLSHHN--PIVFEATDSIGGIWRHCVYNCTKLQSQTWNYEFSDFPWP 283
Query: 104 SGFPT-YPTKQQFIEYLESYSKNFDIRPW--FNETVMHAEF 141
T YP+ + +EYLE+Y++ FD+ FN V+ +F
Sbjct: 284 KRESTDYPSYLEILEYLENYAERFDLNKLVKFNTKVLEVKF 406
>TC18172 similar to UP|O81815 (O81815) Monooxygenase, partial (15%)
Length = 597
Score = 32.7 bits (73), Expect = 0.12
Identities = 16/30 (53%), Positives = 21/30 (69%)
Frame = +2
Query: 43 LIVGAGPSGLAVAAYLKQKGVPSLILERSN 72
+IVG G GLA A L +K + SL+LERS+
Sbjct: 95 VIVGGGICGLATALALHRKSIKSLVLERSD 184
>TC17820
Length = 584
Score = 31.6 bits (70), Expect = 0.26
Identities = 14/25 (56%), Positives = 17/25 (68%)
Frame = +3
Query: 392 GLLGASMDAKNIAEDIERCWKAEAK 416
GL GAS DA I++DI + WK E K
Sbjct: 3 GLSGASSDAMKISQDIGQVWKEETK 77
>AW720305
Length = 577
Score = 30.0 bits (66), Expect = 0.75
Identities = 14/34 (41%), Positives = 21/34 (61%)
Frame = +3
Query: 44 IVGAGPSGLAVAAYLKQKGVPSLILERSNCIASL 77
+VG+GP+GLA A L + G + ER++ I L
Sbjct: 312 VVGSGPAGLAAADQLNKMGHTVTVFERADRIGGL 413
>AV777294
Length = 628
Score = 29.3 bits (64), Expect = 1.3
Identities = 15/38 (39%), Positives = 23/38 (60%)
Frame = +3
Query: 329 VEFADGSTENFDAIILATGYKSNVPYWLKDKGMFSKED 366
V F DGS D I+ TGYK + P+ L+ +G+ + +D
Sbjct: 132 VVFQDGSAIAVDCILHCTGYKYDFPF-LETEGLVTVDD 242
>TC8577 similar to UP|Q9XE94 (Q9XE94) Geranylgeranyl hydrogenase, partial
(53%)
Length = 790
Score = 28.9 bits (63), Expect = 1.7
Identities = 13/32 (40%), Positives = 20/32 (61%), Gaps = 2/32 (6%)
Frame = +2
Query: 44 IVGAGPSGLAVAAYLKQKGVPSLILER--SNC 73
+VG GP+G A A L + G+ + ++ER NC
Sbjct: 221 VVGGGPAGGAAAETLSKAGIETFLIERKMDNC 316
>AV774275
Length = 471
Score = 27.3 bits (59), Expect = 4.9
Identities = 14/38 (36%), Positives = 20/38 (51%), Gaps = 3/38 (7%)
Frame = +3
Query: 23 INNNKNSTSSSDRC---LWIPGPLIVGAGPSGLAVAAY 57
I+NNKN + RC WI L++ AG + L V +
Sbjct: 144 IDNNKNKQAQESRCPK*KWIISNLLLSAGLATLRVVVF 257
>TC18454 similar to UP|Q9XEY7 (Q9XEY7) Trehalase 1 GMTRE1, partial (30%)
Length = 679
Score = 27.3 bits (59), Expect = 4.9
Identities = 16/49 (32%), Positives = 22/49 (44%)
Frame = +1
Query: 363 SKEDGYPRKPFPNGWKGENGLYAVGFTKRGLLGASMDAKNIAEDIERCW 411
S D + FPNGW + G K GL +A+ +AE+I W
Sbjct: 181 SLSDSGQQWDFPNGWAPLQHMLVEGLLKSGL----EEARTLAEEIAIRW 315
>TC12751 similar to UP|LEU3_BRANA (P29102) 3-isopropylmalate dehydrogenase,
chloroplast precursor (Beta-IPM dehydrogenase) (IMDH)
(3-IPM-DH) , partial (25%)
Length = 450
Score = 26.6 bits (57), Expect = 8.3
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = +3
Query: 29 STSSSDRCLWIPGPLIVGAGP 49
S SSS+RC W+ PL++ P
Sbjct: 282 SISSSERCFWVELPLMLRESP 344
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,354,481
Number of Sequences: 28460
Number of extensions: 120200
Number of successful extensions: 518
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 514
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 518
length of query: 423
length of database: 4,897,600
effective HSP length: 93
effective length of query: 330
effective length of database: 2,250,820
effective search space: 742770600
effective search space used: 742770600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC127429.3


