

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127021.2 - phase: 0
(377 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
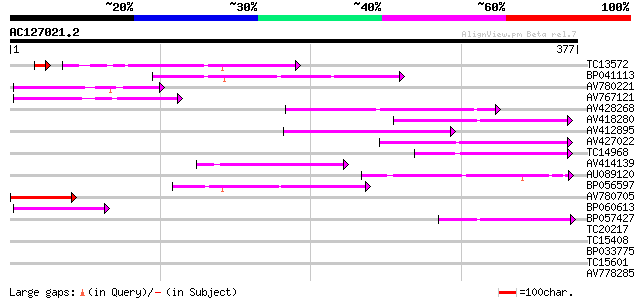
Score E
Sequences producing significant alignments: (bits) Value
TC13572 86 4e-19
BP041113 87 5e-18
AV780221 81 2e-16
AV767121 79 2e-15
AV428268 67 4e-12
AV418280 66 8e-12
AV412895 55 2e-08
AV427022 51 3e-07
TC14968 similar to UP|Q8TTE6 (Q8TTE6) Bacterial extracellular so... 51 4e-07
AV414139 49 1e-06
AU089120 48 2e-06
BP056597 46 9e-06
AV780705 43 1e-04
BP060613 42 2e-04
BP057427 41 4e-04
TC20217 39 0.001
TC15408 similar to UP|Q9LIC2 (Q9LIC2) Multispanning membrane pro... 38 0.003
BP033775 37 0.004
TC15601 35 0.016
AV778285 35 0.020
>TC13572
Length = 534
Score = 86.3 bits (212), Expect(2) = 4e-19
Identities = 55/161 (34%), Positives = 81/161 (50%), Gaps = 3/161 (1%)
Frame = -3
Query: 36 WFKVLSSRYGEQRGRLREGGVTG-SAWWREIVKIQNGIGVEGGSWFEESISKRLGNGFNT 94
W +V+ ++YG+ GG T S+WWR++ Q G E G WFEE + K L +G +T
Sbjct: 478 WCRVVLAKYGD-------GGCTNISSWWRDV---QTAGGGENG-WFEEGVHKVLRSGNHT 332
Query: 95 FFWSDCWVGTVSFMERFRRLYDLSIHKDLSVGEMHALGWGEDGEAW--RWRRRLLAWEEE 152
FW + W G E++ RLY+LS K S+ E GW + W RWRR L E +
Sbjct: 331 RFWLENWTGVGILREKYYRLYNLSKLKWASIDECG--GWSQGVWNWDFRWRRPLAGRELD 158
Query: 153 LVVEIRNLLTNVTLQDTQPDVWLWQPNIGDGYTVRGVYQML 193
+ + + V L PD W+W+P+ Y+V + L
Sbjct: 157 WLQALSLDIVRVPLLAGVPDKWVWKPSEDGSYSVNSAFIFL 35
Score = 24.6 bits (52), Expect(2) = 4e-19
Identities = 9/11 (81%), Positives = 9/11 (81%)
Frame = -1
Query: 17 NIALLGKWCWR 27
N ALLGKW WR
Sbjct: 534 NKALLGKWLWR 502
>BP041113
Length = 503
Score = 87.0 bits (214), Expect = 5e-18
Identities = 54/170 (31%), Positives = 81/170 (46%), Gaps = 3/170 (1%)
Frame = +1
Query: 96 FWSDCWVGTVSFMERFRRLYDLSIHKDLSVGEMHALGWGEDGEAWR--WRRRLLAWEEEL 153
FW++ W+G + +R+ RL++LS K + E GW W+ WRR L+ E
Sbjct: 1 FWTEDWIGMGTLRDRYYRLFNLSKEKRCCILECG--GWNHGVWTWKLSWRRALVGRELGW 174
Query: 154 VVEIRNLLTNVTLQDTQPDVWLWQPNIGDGYTVRGVYQMLMRQEVHNHDVVSDAPWHKSV 213
+ + L+ V L + D WLW P G YTV Y L + + D + W
Sbjct: 175 LELMMKDLSGVCLSEGVEDKWLWLP--GGTYTVNSAYSFLQAPTLADTDPIFATIWRTVA 348
Query: 214 PLKVSICAWRLLRNRWPTKDNLVRRGVITNDTQLCVRGC-GKNETIDHLI 262
P V AWR L R PT DNL++R V+ + + C G+ E++ HL+
Sbjct: 349 P-SVKAFAWRCLLGRLPTYDNLIKRQVVVDPAMTVCKFCQGEVESVTHLL 495
>AV780221
Length = 461
Score = 81.3 bits (199), Expect = 2e-16
Identities = 40/103 (38%), Positives = 59/103 (56%), Gaps = 2/103 (1%)
Frame = +3
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLREGGVTGSAWW 62
K GGLGV+ L FN ALLGKW WR L +R+GLW+KVL +Y + + S+WW
Sbjct: 180 KDEGGLGVQNLGLFNKALLGKWRWRMLKERDGLWYKVLLIKYKNAIPQ------SASSWW 341
Query: 63 REI--VKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVG 103
++ ++G GG W + + +R+G G FW++ W+G
Sbjct: 342 NDLHSTCFEDG----GGGWMQRGLCRRIGEGTEVKFWNENWLG 458
>AV767121
Length = 525
Score = 78.6 bits (192), Expect = 2e-15
Identities = 41/113 (36%), Positives = 60/113 (52%)
Frame = +2
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLREGGVTGSAWW 62
K +GGLG++ L FN ALLGK WR L + + LW +V+ ++ GS+WW
Sbjct: 218 KELGGLGIKDLFTFNKALLGK*RWRYLTEPDSLWRRVIEAQPDHYS--------CGSSWW 373
Query: 63 REIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRRLY 115
+I+ + + WF + K +G G T FWS+ W+G+ RFRRLY
Sbjct: 374 NDILSL---CPEDADGWFSSGLKKLVGEGDQTKFWSEDWLGSGLLSARFRRLY 523
>AV428268
Length = 429
Score = 67.4 bits (163), Expect = 4e-12
Identities = 44/143 (30%), Positives = 65/143 (44%)
Frame = +3
Query: 184 YTVRGVYQMLMRQEVHNHDVVSDAPWHKSVPLKVSICAWRLLRNRWPTKDNLVRRGVITN 243
Y+V+ Y L D + W VP +W++L NR TK NL RRGV+ +
Sbjct: 6 YSVKSAYDFLHGCANAQWDNIFKLLWSVKVPSNAIALSWKVLINRVQTKVNLNRRGVVIS 185
Query: 244 DTQLCVRGCGKNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQVLDHYYQFVYSSGCY 303
+ +C E+ DHL+ C I +W +I GVFSV P H+ Q + S G
Sbjct: 186 N--VCPLCSLDEESTDHLLFSCPIV*RIWSKISE*FGVFSVFPNDSHGHFLQHLGSCGSL 359
Query: 304 NPRRSFLQLIWLCGIWVLSNERN 326
N R+ +W+ + + RN
Sbjct: 360 NFRKRG-WFVWIAAVVCIWQGRN 425
>AV418280
Length = 398
Score = 66.2 bits (160), Expect = 8e-12
Identities = 37/119 (31%), Positives = 55/119 (46%)
Frame = -2
Query: 256 ETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQVLDHYYQFVYSSGCYNPRRSFLQLIWL 315
E HL C+ +WQ + W+G+ P L ++ QF ++ N R L IW+
Sbjct: 364 ECSGHLFFTCVFSMGVWQALHRWLGISVALPASTLANFAQFSITARNKNQRLGELA-IWI 188
Query: 316 CGIWVLSNERNQRLFTNTARTTVQLLENVKITSLKWLKAKNVCFPFGYHMWWQRPFACL 374
+W L +RN +F N A LL+ ++ S WLKAK F + + W P CL
Sbjct: 187 ATVWSLWIQRNSIIFRNNALDHSYLLDLIQSRSWHWLKAKFCGFTYSLYEWKSCPLECL 11
>AV412895
Length = 405
Score = 54.7 bits (130), Expect = 2e-08
Identities = 34/115 (29%), Positives = 49/115 (42%), Gaps = 1/115 (0%)
Frame = +1
Query: 183 GYTVRGVYQMLMRQEVHNHDVVSDAPWHKSVPLKVSICAWRLLRNRWPTKDNLVRRGVIT 242
G++V + L Q + D V + W PL + WR+L R T+DNL++R VI
Sbjct: 7 GFSVNSSFVFL*DQYLEEPDPVFN*IWMVPAPLNIKAFVWRVLLGRIQTRDNLLKRQVIH 186
Query: 243 NDTQLCVRGCG-KNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQVLDHYYQF 296
N CG E+ HL+ C +W W+GV + H QF
Sbjct: 187 NALDAICPLCGLAEESGSHLLFSCAESMLIWYECFAWLGVSTAQVSDPKVHLLQF 351
>AV427022
Length = 429
Score = 51.2 bits (121), Expect = 3e-07
Identities = 32/128 (25%), Positives = 55/128 (42%)
Frame = +1
Query: 247 LCVRGCGKNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQVLDHYYQFVYSSGCYNPR 306
+C C E+ HL+ C + W+GV + H QF S G +
Sbjct: 37 ICPLCCLAEESGSHLLFSCSNSMLI*YECHAWLGVSTAQVSDPKVHLLQFS-SIGWSKAQ 213
Query: 307 RSFLQLIWLCGIWVLSNERNQRLFTNTARTTVQLLENVKITSLKWLKAKNVCFPFGYHMW 366
+ IW+ +W + RN +F QLLE +++T+ +WL+++ F + ++ W
Sbjct: 214 KLGESAIWMSVLWSIWCLRNMVVFNGGELDKDQLLEQIQVTAWRWLRSRLDDF*YSWYEW 393
Query: 367 WQRPFACL 374
P CL
Sbjct: 394 KSNPRICL 417
>TC14968 similar to UP|Q8TTE6 (Q8TTE6) Bacterial extracellular
solute-binding family 3 protein, partial (3%)
Length = 503
Score = 50.8 bits (120), Expect = 4e-07
Identities = 28/106 (26%), Positives = 50/106 (46%), Gaps = 1/106 (0%)
Frame = +1
Query: 270 DLWQRIKTWIGVFSVDPQQVLDHYYQFVYSSGCY-NPRRSFLQLIWLCGIWVLSNERNQR 328
++W R+ W+GV V P +DH F +G + ++ + +WL +W + RN+
Sbjct: 88 EIWCRVFNWLGVCFVCPGTPVDHLLSF---AGLFLLAQKDGILSVWLAVVWHVWVGRNEA 258
Query: 329 LFTNTARTTVQLLENVKITSLKWLKAKNVCFPFGYHMWWQRPFACL 374
+F + VK+ +WLKA++ F + W+ P CL
Sbjct: 259 VFREGVFLPECIFYLVKLKVWEWLKARSCHFTHATYEWFNEPILCL 396
>AV414139
Length = 309
Score = 49.3 bits (116), Expect = 1e-06
Identities = 30/101 (29%), Positives = 43/101 (41%)
Frame = -2
Query: 125 VGEMHALGWGEDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQDTQPDVWLWQPNIGDGY 184
+G+ H W + +RWRR L E+ ++ L V LQ D W W + D Y
Sbjct: 299 MGQWHNGVWEWE---FRWRRELSVREQRQEADLIQALAGVCLQQGTEDRWKWTLDTEDIY 129
Query: 185 TVRGVYQMLMRQEVHNHDVVSDAPWHKSVPLKVSICAWRLL 225
TVR Y +L+ + V + W P + WRLL
Sbjct: 128 TVRSAY*LLLDPIADSDSEVYEKVWVGIAPSNANAFVWRLL 6
>AU089120
Length = 586
Score = 48.1 bits (113), Expect = 2e-06
Identities = 40/144 (27%), Positives = 66/144 (45%), Gaps = 3/144 (2%)
Frame = +2
Query: 235 LVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQVLDHYY 294
L RR +I + C+ E+I HL C +LW + W+ V P++ H+
Sbjct: 8 LKRRRIIQTLSSFCLHS---EESIPHLFFRCFQSWNLWSSVHRWLVFTVVTPEEGKMHFI 178
Query: 295 QF-VYSSGCYNPRRSFLQLIWLCGIWVLSNERNQRLFTNTARTTVQ--LLENVKITSLKW 351
F Y SG ++ L IWL IW + + RN+ +F V+ +++ I ++
Sbjct: 179 MF*DYYSG--RDIKNGLGFIWLSTIWFIWHLRNKVVFNGRCFRLVRGFRFDSISIVAMAK 352
Query: 352 LKAKNVCFPFGYHMWWQRPFACLG 375
K++ V F F + +Q PF LG
Sbjct: 353 RKSERVMF-FAL*VVFQ-PFGLLG 418
>BP056597
Length = 414
Score = 46.2 bits (108), Expect = 9e-06
Identities = 35/135 (25%), Positives = 61/135 (44%), Gaps = 3/135 (2%)
Frame = +3
Query: 109 ERFRRLYDLSIHKDLSVGEMHALGWGEDGEAW--RWRRRLLAWEEELVVEIRNLLTNVTL 166
E F L+ +S ++ + EM W E+ W W+ + + + ++ ++++ +L
Sbjct: 3 ETFDSLFLISNQQNCKIVEMGH--WLENKWVWDFSWKNPISGEDVLKLGVMKQIVSSFSL 176
Query: 167 QDTQPDVWLWQPNIGDG-YTVRGVYQMLMRQEVHNHDVVSDAPWHKSVPLKVSICAWRLL 225
+ + D W+W GDG Y+V+ Y +L + V W P WR+
Sbjct: 177 INGKKDTWVWILE-GDGQYSVKSAYDLLSGLDTTIGISVFSKLWKACAPSNAVALGWRVF 353
Query: 226 RNRWPTKDNLVRRGV 240
+R TKDNL RR V
Sbjct: 354 LDRIQTKDNLSRRHV 398
>AV780705
Length = 524
Score = 42.7 bits (99), Expect = 1e-04
Identities = 19/44 (43%), Positives = 27/44 (61%)
Frame = -2
Query: 1 MSKGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRY 44
+SK GG+G R L+ N+A L + WR L + E LW +V+ S Y
Sbjct: 193 LSKEKGGVGFRDLRTQNLAFLARQAWRVLTNPEALWVRVMKSLY 62
>BP060613
Length = 378
Score = 41.6 bits (96), Expect = 2e-04
Identities = 22/65 (33%), Positives = 34/65 (51%), Gaps = 1/65 (1%)
Frame = +1
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLREGGVTGSAW- 61
K GGLG + + N ALL K WR L + + LW ++L + Y L+ TG++W
Sbjct: 109 KDEGGLGFKDFEIQNQALLAKQAWRILHNPDALWVQILKALYFPHHDFLQTTKRTGASWV 288
Query: 62 WREIV 66
W ++
Sbjct: 289 WSSLL 303
>BP057427
Length = 480
Score = 40.8 bits (94), Expect = 4e-04
Identities = 24/91 (26%), Positives = 40/91 (43%)
Frame = -1
Query: 286 PQQVLDHYYQFVYSSGCYNPRRSFLQLIWLCGIWVLSNERNQRLFTNTARTTVQLLENVK 345
P + ++ QF+ G RR IW+ + + RN +F +L++ ++
Sbjct: 477 PIEASVNFLQFMQGIGNLKRRRGAWS-IWVAALASIWYSRNSLIFKGEELDFQRLVDQIQ 301
Query: 346 ITSLKWLKAKNVCFPFGYHMWWQRPFACLGI 376
S WLKAK F + W +P +CL I
Sbjct: 300 TKSWSWLKAKVTGFSYSLIEWKSQPLSCLDI 208
>TC20217
Length = 561
Score = 39.3 bits (90), Expect = 0.001
Identities = 24/82 (29%), Positives = 37/82 (44%)
Frame = +1
Query: 251 GCGKNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQVLDHYYQFVYSSGCYNPRRSFL 310
GCG + L+ C + D+W+ W+GV + P+ H F + + R +
Sbjct: 283 GCG---IL*ALLFSCPVSLDIWRHCYRWMGVCTTLPRNPRQHLL*FQFGGNKKHQRGA-- 447
Query: 311 QLIWLCGIWVLSNERNQRLFTN 332
IWL IW L RN+ +F N
Sbjct: 448 DAIWLAVIWTLWLIRNEIIFRN 513
>TC15408 similar to UP|Q9LIC2 (Q9LIC2) Multispanning membrane protein-like,
partial (10%)
Length = 879
Score = 37.7 bits (86), Expect = 0.003
Identities = 23/74 (31%), Positives = 30/74 (40%), Gaps = 4/74 (5%)
Frame = -1
Query: 140 WRW----RRRLLAWEEELVVEIRNLLTNVTLQDTQPDVWLWQPNIGDGYTVRGVYQMLMR 195
W W RR E LV ++ NLL +V L + D W W N YTV+ Y+ L
Sbjct: 768 WTWSFIRRRNPSTRENLLVADLCNLLESVPLSEDTNDSWSWTANPEGQYTVQTAYKFLRA 589
Query: 196 QEVHNHDVVSDAPW 209
D + W
Sbjct: 588 SSNEFTDPIFHFVW 547
>BP033775
Length = 530
Score = 37.4 bits (85), Expect = 0.004
Identities = 17/53 (32%), Positives = 26/53 (48%)
Frame = -2
Query: 325 RNQRLFTNTARTTVQLLENVKITSLKWLKAKNVCFPFGYHMWWQRPFACLGIG 377
RN LF + +++ +K + KWL+ K F + + W P CLGIG
Sbjct: 394 RNDVLFNGKTINSEAVVDLIKFKAWKWLQCKTKDFSYSSYAWNWNPCICLGIG 236
>TC15601
Length = 613
Score = 35.4 bits (80), Expect = 0.016
Identities = 17/57 (29%), Positives = 27/57 (46%)
Frame = +2
Query: 319 WVLSNERNQRLFTNTARTTVQLLENVKITSLKWLKAKNVCFPFGYHMWWQRPFACLG 375
W L N RN +F +++E ++ + WL+AK F F + W P C+G
Sbjct: 101 WTLWNLRNMIIFKEGHLDGERVVEIAQLRAWNWLRAKKKDFKFSTYEWVVNPRYCIG 271
>AV778285
Length = 519
Score = 35.0 bits (79), Expect = 0.020
Identities = 17/47 (36%), Positives = 24/47 (50%)
Frame = -2
Query: 329 LFTNTARTTVQLLENVKITSLKWLKAKNVCFPFGYHMWWQRPFACLG 375
LF ++ +++E VK S KWLK K F ++ W P CLG
Sbjct: 515 LFNARPISSEKVVELVKFRSWKWLKCKQNHFKASFYEWETNPSVCLG 375
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.141 0.492
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,156,587
Number of Sequences: 28460
Number of extensions: 158068
Number of successful extensions: 1547
Number of sequences better than 10.0: 71
Number of HSP's better than 10.0 without gapping: 1514
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1537
length of query: 377
length of database: 4,897,600
effective HSP length: 92
effective length of query: 285
effective length of database: 2,279,280
effective search space: 649594800
effective search space used: 649594800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC127021.2


