

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
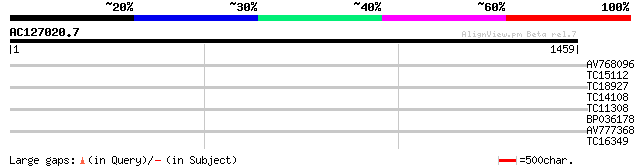
Query= AC127020.7 + phase: 0 /pseudo
(1459 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV768096 40 0.003
TC15112 similar to UP|Q8VYI1 (Q8VYI1) At1g69640/F24J1.22, partia... 32 0.72
TC18927 similar to PIR|AI2934|AI2934 chromate transport protein ... 31 1.2
TC14108 similar to UP|CAHC_PEA (P17067) Carbonic anhydrase, chlo... 30 3.6
TC11308 similar to UP|CENB_HUMAN (P07199) Major centromere autoa... 30 3.6
BP036178 30 3.6
AV777368 29 6.1
TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulat... 28 8.0
>AV768096
Length = 358
Score = 40.0 bits (92), Expect = 0.003
Identities = 27/94 (28%), Positives = 39/94 (40%), Gaps = 4/94 (4%)
Frame = +3
Query: 61 RAWKPGKRVTITKSVADRYLFQFHHKVDDARVLEEGPWLYDNFHIVLDRIAPSAVPNLVP 120
RAW + + + + +LF F R+L EGPW + L P V
Sbjct: 51 RAWGNLQDLQVGDLNLNTFLFTFSDANLATRILAEGPWNIMGNLLSLQFWDPQISMAEVH 230
Query: 121 LNHIEFWVQVHGLPFGFVQ----PKVGQGIGSFL 150
N + FW Q HGLP F+ K+ +G+ L
Sbjct: 231 FNLVPFWTQFHGLPMEFMSLTNVSKLAAAVGNLL 332
>TC15112 similar to UP|Q8VYI1 (Q8VYI1) At1g69640/F24J1.22, partial (37%)
Length = 651
Score = 32.0 bits (71), Expect = 0.72
Identities = 13/31 (41%), Positives = 22/31 (70%)
Frame = +1
Query: 636 HLLIICGFRIFPLLMLKLWWLRLLITTQFTS 666
HLL I G+ + + M ++W+LRLL+ +F+S
Sbjct: 406 HLLRIIGYTLSRMRMKRIWFLRLLLLKEFSS 498
>TC18927 similar to PIR|AI2934|AI2934 chromate transport protein chrA
[imported] - Agrobacterium tumefaciens
(strain C58, Dupont) {Agrobacterium tumefaciens;},
partial (6%)
Length = 561
Score = 31.2 bits (69), Expect = 1.2
Identities = 14/35 (40%), Positives = 18/35 (51%)
Frame = -2
Query: 188 SNGNFVTVNFKYEKLGVFCHRCGVLGHTDKVCPEL 222
S GNFV + + + CHRC GH CP+L
Sbjct: 383 SFGNFVPRPTQSDTSEIVCHRCSKKGHFANRCPDL 279
>TC14108 similar to UP|CAHC_PEA (P17067) Carbonic anhydrase, chloroplast
precursor (Carbonate dehydratase) , partial (98%)
Length = 1458
Score = 29.6 bits (65), Expect = 3.6
Identities = 18/69 (26%), Positives = 36/69 (52%), Gaps = 4/69 (5%)
Frame = +1
Query: 332 PSSSTGTSASNLLPDRPIVLG----LPAAPL*SENTDDALEEIGSELKKRKRMLAIQNGE 387
PSSS+ +S +L+ DRP+ +P + +N ++A+EE+ L+++ + A +
Sbjct: 172 PSSSSSSSFPSLIQDRPVFAAPAPIIPTLDM-GKNYEEAIEELQKLLREKNELKATAAEK 348
Query: 388 PSVAEGELG 396
+LG
Sbjct: 349 VEQITAQLG 375
>TC11308 similar to UP|CENB_HUMAN (P07199) Major centromere autoantigen B
(Centromere protein B) (CENP-B), partial (4%)
Length = 756
Score = 29.6 bits (65), Expect = 3.6
Identities = 14/40 (35%), Positives = 24/40 (60%)
Frame = -3
Query: 699 ILGRNILLIRSFRSFPHVRKTCLYGRKAIVIN*KRILKIV 738
ILG L +R + PH R+ +G+ +V+N + I+K+V
Sbjct: 670 ILGGTRLQVRFHQQRPHPREERAWGKNLLVVNRRWIIKVV 551
>BP036178
Length = 561
Score = 29.6 bits (65), Expect = 3.6
Identities = 13/24 (54%), Positives = 15/24 (62%)
Frame = -2
Query: 196 NFKYEKLGVFCHRCGVLGHTDKVC 219
NFK K V C RC +LGH+ K C
Sbjct: 389 NFKRPKRIVQCGRCHMLGHSQKKC 318
>AV777368
Length = 604
Score = 28.9 bits (63), Expect = 6.1
Identities = 30/100 (30%), Positives = 46/100 (46%), Gaps = 2/100 (2%)
Frame = +1
Query: 588 PLDLLGLLTAFVKLFLTLVYLMSRLKVIRSLGSKVSVIPAPSKKG*I--VHLLIICGFRI 645
P D +L FV F++++ L+ +L GSK ++ P+P K I H L +R
Sbjct: 79 PHDPNDILFMFVLSFISMLLLLFKLP----RGSKRNLPPSPPKLPFIGNFHQLGTIPYRS 246
Query: 646 FPLLMLKLWWLRLLITTQFTSILLLHLGPMLLSVIFAMKM 685
F L K + +LLLHLG + L V+ + M
Sbjct: 247 FQTLSQK-----------YGPLLLLHLGHLPLLVVSSADM 333
>TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulatory subunit
6B homolog, partial (65%)
Length = 901
Score = 28.5 bits (62), Expect = 8.0
Identities = 26/89 (29%), Positives = 46/89 (51%), Gaps = 1/89 (1%)
Frame = +1
Query: 573 AILMTSSIQVRKGGTPLDLLGL-LTAFVKLFLTLVYLMSRLKVIRSLGSKVSVIPAPSKK 631
AI M SS+ + P L L + + L +TL ++S+ + L S I + + K
Sbjct: 406 AIPMLSSMFFHRRQIPASLFSASLRSQMLLTMTLEGVISKSRKFVRLWSYPLHIMSYTNK 585
Query: 632 G*IVHLLIICGFRIFPLLMLKLWWLRLLI 660
+V +L++ + + PL ++K WLRLL+
Sbjct: 586 --LVLILLVVYYYMVPLALVKPCWLRLLL 666
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.351 0.158 0.526
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,657,522
Number of Sequences: 28460
Number of extensions: 377547
Number of successful extensions: 3195
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 3146
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3193
length of query: 1459
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1357
effective length of database: 1,994,680
effective search space: 2706780760
effective search space used: 2706780760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC127020.7


