

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127019.14 + phase: 0 /pseudo
(532 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
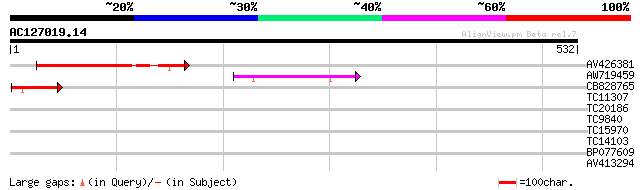
Score E
Sequences producing significant alignments: (bits) Value
AV426381 118 3e-27
AW719459 84 4e-17
CB828765 53 1e-07
TC11307 29 1.7
TC20186 29 2.2
TC9840 UP|R11D_LOTJA (Q40194) Ras-related protein Rab11D, complete 29 2.2
TC15970 28 4.8
TC14103 similar to UP|Q84XC6 (Q84XC6) Plasma intrinsic protein 2... 28 4.8
BP077609 28 4.8
AV413294 27 6.3
>AV426381
Length = 430
Score = 118 bits (295), Expect = 3e-27
Identities = 64/145 (44%), Positives = 91/145 (62%), Gaps = 2/145 (1%)
Frame = +2
Query: 26 SSRTLDQTPTWGVAAVCTVFILISIALEKSLHKLGTWLGKREKKALLEALEKVKAELMIL 85
+ + L++TPTW VA VC V + ISI +E + + WL KR KKAL EA+EK+K ELM++
Sbjct: 23 AQKKLEETPTWAVAVVCFVMLAISIIIEHGIEAIEKWLEKRHKKALHEAVEKIKGELMLM 202
Query: 86 GFISLLLTIGQNYIVRICISEKIANKMLPCPHKYIGGDKKSNSRYLAADTPSFKCSRKGH 145
GFIS LLT+ ++ I ICIS+++A+ PC + KK Y KC++ G
Sbjct: 203 GFISFLLTVFKDPISNICISKQVASTWHPC---HPEEKKKGPEGYYD------KCAKDGK 355
Query: 146 EPL--LSVNGLHQLHILIFFLAVFH 168
+ + +S G+HQLHI IF LA+FH
Sbjct: 356 DKVAFMSQYGIHQLHIFIFVLAIFH 430
>AW719459
Length = 402
Score = 84.3 bits (207), Expect = 4e-17
Identities = 50/132 (37%), Positives = 74/132 (55%), Gaps = 13/132 (9%)
Frame = +1
Query: 211 HETSFVRAHTSFWTRVY-----IGCFFRQFYRSVRKVDYQTLRNGFINVHLAPGSKFNFQ 265
H+ +F++ S + + Y + FF+QFY S K+DY TLR GFI H KFNF
Sbjct: 4 HQHAFIQNRXSGFGKDYTLMGWLKSFFKQFYGSXTKLDYVTLRLGFIMTHCRGNPKFNFH 183
Query: 266 KYIKRSLEDDFKVVVGVSPVLWASAVVFLLLNID--------AYIIFRMAYSNMGILDSN 317
KY+ R+LEDD K VVG+S LW ++ LLLNI+ A+I + + L+
Sbjct: 184 KYMIRALEDDXKKVVGISWYLWVFVIIXLLLNINGWHTYFWIAFIPVILLLAXGTKLEHV 363
Query: 318 YSRMALEISERH 329
++A E++E+H
Sbjct: 364 IMQLAHEVAEKH 399
>CB828765
Length = 343
Score = 53.1 bits (126), Expect = 1e-07
Identities = 31/52 (59%), Positives = 38/52 (72%), Gaps = 4/52 (7%)
Frame = +3
Query: 2 FKRVFSCLC--LFYGG-LSMAASDS-RNSSRTLDQTPTWGVAAVCTVFILIS 49
+K + CL L YGG L+MA+S+ +SS+ LDQTPTW VA VCTVFILIS
Sbjct: 186 YKSLSLCLVSWLCYGGVLAMASSEEGSSSSKNLDQTPTWAVACVCTVFILIS 341
>TC11307
Length = 524
Score = 29.3 bits (64), Expect = 1.7
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = +2
Query: 31 DQTPTWGVAAVCTVFILISIALEKSLHKLGTWL 63
D+TP V A C + + I I LHK+ TW+
Sbjct: 41 DRTPVNKVGAYCWLSLAICIVRTSDLHKVWTWV 139
>TC20186
Length = 538
Score = 28.9 bits (63), Expect = 2.2
Identities = 17/22 (77%), Positives = 19/22 (86%)
Frame = +2
Query: 497 L*ELIMMSKKQKIGIIIVREKP 518
L*+LIMMS KQKI IIVRE+P
Sbjct: 59 L*KLIMMSHKQKIR-IIVRERP 121
>TC9840 UP|R11D_LOTJA (Q40194) Ras-related protein Rab11D, complete
Length = 1302
Score = 28.9 bits (63), Expect = 2.2
Identities = 17/36 (47%), Positives = 18/36 (49%)
Frame = +1
Query: 335 MPLVQGSDRYFWLGQPQLVLHLIHFALFQNSFQITY 370
+P GSDR FW Q QL L IH Q QI Y
Sbjct: 280 LPFQAGSDRRFWCRQVQLALQ-IHQERVQFGVQIHY 384
>TC15970
Length = 553
Score = 27.7 bits (60), Expect = 4.8
Identities = 17/44 (38%), Positives = 23/44 (51%)
Frame = +3
Query: 352 LVLHLIHFALFQNSFQITYILWTWYSFGLENCFRLDYKLAIIKV 395
L L ++HFA+F N +L YSF NCF L KL + +
Sbjct: 396 LPLLVVHFAMFFNVIAFCNVL*C-YSFHHCNCFHLKNKLCNLHI 524
>TC14103 similar to UP|Q84XC6 (Q84XC6) Plasma intrinsic protein 2,2, partial
(94%)
Length = 1199
Score = 27.7 bits (60), Expect = 4.8
Identities = 19/63 (30%), Positives = 28/63 (44%), Gaps = 3/63 (4%)
Frame = -1
Query: 7 SCLCLFYGGLSMAASDSRNSSRTLDQTPTWGVAA---VCTVFILISIALEKSLHKLGTWL 63
S + L GG + + QT + GVA VCT + + +A KSLH+ T
Sbjct: 677 SGVSLGIGGGEDGVDKDKGADNLSAQTSSLGVAISKHVCTAMVPVVVAFLKSLHQPNTAY 498
Query: 64 GKR 66
G +
Sbjct: 497 GSQ 489
>BP077609
Length = 311
Score = 27.7 bits (60), Expect = 4.8
Identities = 14/41 (34%), Positives = 20/41 (48%)
Frame = +2
Query: 350 PQLVLHLIHFALFQNSFQITYILWTWYSFGLENCFRLDYKL 390
P L+LH I +F++ F T W+ EN FR+ L
Sbjct: 17 PNLILHRISMLVFESDFGATNTYPNWHLPLPENTFRITQAL 139
>AV413294
Length = 333
Score = 27.3 bits (59), Expect = 6.3
Identities = 8/20 (40%), Positives = 13/20 (65%)
Frame = +2
Query: 358 HFALFQNSFQITYILWTWYS 377
H F+ +FQI +++W W S
Sbjct: 59 HLNCFEKNFQIDFLMWEWLS 118
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.338 0.147 0.474
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,953,452
Number of Sequences: 28460
Number of extensions: 143778
Number of successful extensions: 1303
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1297
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1302
length of query: 532
length of database: 4,897,600
effective HSP length: 95
effective length of query: 437
effective length of database: 2,193,900
effective search space: 958734300
effective search space used: 958734300
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC127019.14


