

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
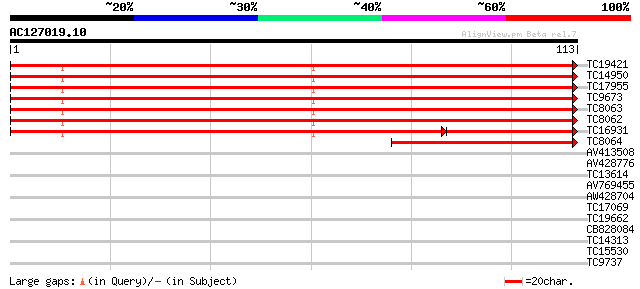
Query= AC127019.10 + phase: 0
(113 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC19421 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {M... 136 6e-34
TC14950 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {M... 136 6e-34
TC17955 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {M... 136 6e-34
TC9673 UP|BAA31218 (BAA31218) Histone H3, complete 129 7e-32
TC8063 UP|BAA31218 (BAA31218) Histone H3, complete 129 7e-32
TC8062 UP|BAA31218 (BAA31218) Histone H3, complete 129 7e-32
TC16931 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {M... 99 5e-29
TC8064 homologue to GB|AAB36496.1|488573|MSU09463 histone H3.2 p... 52 1e-08
AV413508 29 0.098
AV428776 28 0.17
TC13614 27 0.64
AV769455 27 0.64
AW428704 26 1.1
TC17069 similar to UP|Q9XGN6 (Q9XGN6) ESTs C99174(E10437), parti... 26 1.1
TC19662 homologue to GB|CAA65987.2|6015604|PSRPL9PRT ribosomal p... 25 1.4
CB828084 25 1.4
TC14313 similar to UP|O80407 (O80407) Chalcone synthase , parti... 25 1.4
TC15530 similar to UP|AAQ84318 (AAQ84318) Fiber protein Fb27, pa... 24 3.2
TC9737 similar to UP|Q9LRT1 (Q9LRT1) Receptor protein kinase, pa... 24 3.2
>TC19421 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {Medicago
sativa;}, complete
Length = 672
Score = 136 bits (342), Expect = 6e-34
Identities = 78/116 (67%), Positives = 84/116 (72%), Gaps = 3/116 (2%)
Frame = +3
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA A G VKKPH F GTVALREIR YQ+ST
Sbjct: 93 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 272
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQSSAVSA+QE EA V LFEDTN+CAIHAK
Sbjct: 273 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVSALQEAAEAYLVGLFEDTNLCAIHAK 440
>TC14950 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {Medicago
sativa;}, complete
Length = 641
Score = 136 bits (342), Expect = 6e-34
Identities = 78/116 (67%), Positives = 84/116 (72%), Gaps = 3/116 (2%)
Frame = +2
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA A G VKKPH F GTVALREIR YQ+ST
Sbjct: 65 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 244
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQSSAVSA+QE EA V LFEDTN+CAIHAK
Sbjct: 245 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVSALQEAAEAYLVGLFEDTNLCAIHAK 412
>TC17955 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {Medicago
sativa;}, complete
Length = 639
Score = 136 bits (342), Expect = 6e-34
Identities = 78/116 (67%), Positives = 84/116 (72%), Gaps = 3/116 (2%)
Frame = +1
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA A G VKKPH F GTVALREIR YQ+ST
Sbjct: 58 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 237
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQSSAVSA+QE EA V LFEDTN+CAIHAK
Sbjct: 238 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVSALQEAAEAYLVGLFEDTNLCAIHAK 405
>TC9673 UP|BAA31218 (BAA31218) Histone H3, complete
Length = 690
Score = 129 bits (324), Expect = 7e-32
Identities = 74/116 (63%), Positives = 81/116 (69%), Gaps = 3/116 (2%)
Frame = +1
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA G VKKPH + GTVALREIR YQ+ST
Sbjct: 76 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 255
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQS AV A+QE EA V LFEDTN+CAIHAK
Sbjct: 256 LLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAK 423
>TC8063 UP|BAA31218 (BAA31218) Histone H3, complete
Length = 723
Score = 129 bits (324), Expect = 7e-32
Identities = 74/116 (63%), Positives = 81/116 (69%), Gaps = 3/116 (2%)
Frame = +2
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA G VKKPH + GTVALREIR YQ+ST
Sbjct: 71 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 250
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQS AV A+QE EA V LFEDTN+CAIHAK
Sbjct: 251 LLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAK 418
>TC8062 UP|BAA31218 (BAA31218) Histone H3, complete
Length = 768
Score = 129 bits (324), Expect = 7e-32
Identities = 74/116 (63%), Positives = 81/116 (69%), Gaps = 3/116 (2%)
Frame = +3
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA G VKKPH + GTVALREIR YQ+ST
Sbjct: 51 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 230
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQS AV A+QE EA V LFEDTN+CAIHAK
Sbjct: 231 LLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAK 398
>TC16931 PIR|S04520|S04520 histone H3 (clone pH3c-1) - alfalfa {Medicago
sativa;}, partial (96%)
Length = 440
Score = 98.6 bits (244), Expect(2) = 5e-29
Identities = 59/90 (65%), Positives = 63/90 (69%), Gaps = 3/90 (3%)
Frame = +1
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA A G VKKPH F GTVALREIR YQ+ST
Sbjct: 46 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 225
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAV 87
++ FKT LRFQSSAV
Sbjct: 226 LLIRKLPFQRLVREIAQDFKTDLRFQSSAV 315
Score = 42.4 bits (98), Expect(2) = 5e-29
Identities = 19/26 (73%), Positives = 21/26 (80%)
Frame = +2
Query: 88 SAVQEVVEACFVSLFEDTNICAIHAK 113
SA+QE EA V LFEDTN+CAIHAK
Sbjct: 317 SALQEAAEAYLVGLFEDTNLCAIHAK 394
>TC8064 homologue to GB|AAB36496.1|488573|MSU09463 histone H3.2 precursor
{Medicago sativa;} , partial (45%)
Length = 486
Score = 52.0 bits (123), Expect = 1e-08
Identities = 25/37 (67%), Positives = 29/37 (77%)
Frame = +1
Query: 77 KTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
+T LRFQS AV A+QE EA V LFEDT++CAIHAK
Sbjct: 79 QTDLRFQSHAVLALQEAAEAYLVGLFEDTHLCAIHAK 189
>AV413508
Length = 265
Score = 29.3 bits (64), Expect = 0.098
Identities = 14/22 (63%), Positives = 15/22 (67%)
Frame = +3
Query: 28 SALAAGTVKKPHHFCHGTVALR 49
SA G VKKPH + GTVALR
Sbjct: 3 SAPTTGGVKKPHRYRPGTVALR 68
>AV428776
Length = 331
Score = 28.5 bits (62), Expect = 0.17
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = +1
Query: 3 GTKQTARNTGGKAPRKQLATKAARKSALAAGTVK 36
GT QTA NT +P +T+ R AGT++
Sbjct: 178 GTDQTASNTWAHSPASPRSTRPVRSPVTTAGTLQ 279
>TC13614
Length = 514
Score = 26.6 bits (57), Expect = 0.64
Identities = 14/50 (28%), Positives = 27/50 (54%), Gaps = 2/50 (4%)
Frame = -2
Query: 34 TVKKPHHF--CHGTVALREIRNYQRSTSNKEASVSETCEGNCSGFKTGLR 81
T+K H+ +GT + R+ +R+ S+++A + G C GF+ +R
Sbjct: 342 TIKNKHYR*NPNGTSSTATTRDQKRNMSSEDAMIG*IA*GTCQGFQQRVR 193
>AV769455
Length = 556
Score = 26.6 bits (57), Expect = 0.64
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = -2
Query: 53 NYQRSTSNKEASVSETCEGNCSGF 76
N QR K + ++ C+G+CSGF
Sbjct: 450 NIQRIILRKISLINSICQGSCSGF 379
>AW428704
Length = 369
Score = 25.8 bits (55), Expect = 1.1
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = -1
Query: 39 HHFCHGTVALREIRNYQR 56
HHFC ++LRE RN+ +
Sbjct: 249 HHFCKICISLRERRNHSK 196
>TC17069 similar to UP|Q9XGN6 (Q9XGN6) ESTs C99174(E10437), partial (33%)
Length = 577
Score = 25.8 bits (55), Expect = 1.1
Identities = 14/35 (40%), Positives = 16/35 (45%)
Frame = +2
Query: 15 APRKQLATKAARKSALAAGTVKKPHHFCHGTVALR 49
AP L TK A + T + HF HGT LR
Sbjct: 251 APGSNLKTKLCENFAQGSCTFGERCHFAHGTDELR 355
>TC19662 homologue to GB|CAA65987.2|6015604|PSRPL9PRT ribosomal protein L9
{Pisum sativum;} , partial (75%)
Length = 507
Score = 25.4 bits (54), Expect = 1.4
Identities = 16/50 (32%), Positives = 26/50 (52%), Gaps = 10/50 (20%)
Frame = +3
Query: 37 KPHHFCHGTVALR-EIR---------NYQRSTSNKEASVSETCEGNCSGF 76
+PHH CH V L+ E+R ++Q S+++ +S EG SG+
Sbjct: 297 QPHHRCHQGVPLQDEVRLCPFSYQC*HHQ*QQSHRDPKLSRREEGQESGY 446
>CB828084
Length = 528
Score = 25.4 bits (54), Expect = 1.4
Identities = 20/71 (28%), Positives = 33/71 (46%), Gaps = 2/71 (2%)
Frame = +1
Query: 37 KPHHFCHGTVA-LREIRNYQRSTSNKEASVS-ETCEGNCSGFKTGLRFQSSAVSAVQEVV 94
+P H V ++E+ + +TS K +S ET G CSGF+ +V++ V
Sbjct: 118 EPVHITENCVRRMKELEASESTTSRKMLRLSVET--GGCSGFQYAFDLDDDSVNSDDRVF 291
Query: 95 EACFVSLFEDT 105
E + L D+
Sbjct: 292 EKEGIKLVVDS 324
>TC14313 similar to UP|O80407 (O80407) Chalcone synthase , partial (98%)
Length = 1567
Score = 25.4 bits (54), Expect = 1.4
Identities = 12/25 (48%), Positives = 14/25 (56%)
Frame = +2
Query: 34 TVKKPHHFCHGTVALREIRNYQRST 58
T HH H V + EIRN QRS+
Sbjct: 41 TTTTTHHPPHKMVTVEEIRNAQRSS 115
>TC15530 similar to UP|AAQ84318 (AAQ84318) Fiber protein Fb27, partial (59%)
Length = 692
Score = 24.3 bits (51), Expect = 3.2
Identities = 17/62 (27%), Positives = 28/62 (44%)
Frame = -3
Query: 4 TKQTARNTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTSNKEA 63
+KQ NT K+ + + ++ ALA KP +FC + L E + Y+ +
Sbjct: 384 SKQLKYNT--KSNQLVFGSGMGKRPALAPLGGAKPKNFCKLVLILMEHKKYRNAMEQSVG 211
Query: 64 SV 65
SV
Sbjct: 210 SV 205
>TC9737 similar to UP|Q9LRT1 (Q9LRT1) Receptor protein kinase, partial
(11%)
Length = 563
Score = 24.3 bits (51), Expect = 3.2
Identities = 9/18 (50%), Positives = 11/18 (61%)
Frame = -2
Query: 35 VKKPHHFCHGTVALREIR 52
+K HHFCH A R +R
Sbjct: 307 LKNLHHFCHSRPARRYLR 254
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.125 0.355
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,019,949
Number of Sequences: 28460
Number of extensions: 24924
Number of successful extensions: 139
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 125
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 125
length of query: 113
length of database: 4,897,600
effective HSP length: 89
effective length of query: 24
effective length of database: 2,364,660
effective search space: 56751840
effective search space used: 56751840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 47 (22.7 bits)
Medicago: description of AC127019.10


